![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
118 Cards in this Set
- Front
- Back
|
adenine
|
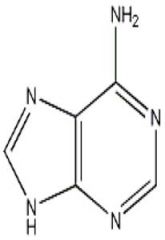
.
|
|
|
cytosine
|
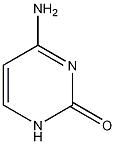
.
|
|
|
guanine
|
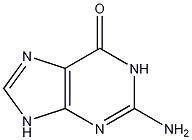
.
|
|
|
thymidine
|
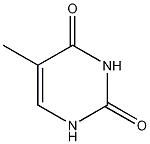
.
|
|
|
What is the linking number?
|
L = T+W
T = twist, or # of helical turns; T = # bp/10 W = writhe, or # of supercoils |
|
|
1. Positive supercoiling _____________ a B-DNA double helix, while negative supercoiling favors ____________ of the helix
|
1: overwinds
2: unwinding |
|
|
True or False?
1. Negative supercoiling compacts the DNA so that it migrates more rapidly on a gel than positive supercoiled DNA. 2. The linking number (L) will not change as long as the two DNA strands remain intact. |
1. FALSE. Both forms of supercoiling compact the DNA so that it would migrate more rapidly on a gel.
2. TRUE |
|
|
Double-stranded DNA molecules assume one of three secondary structures:
What is the major conformation of DNA in solution? |
- A, B, or Z DNA
- B DNA, pitch of 3.4 nm, helix twists every ~10 bp |
|
|
What gives rise to the major and minor grooves of DNA?
|
- the anti-parallel sugar-phosphate backbones of each strand are anti-parallel
- the two glycosidic bonds are not directly across from each other |
|
|
How does A DNA differ from B DNA?
|
- Pitch: B: 3.4 nm, A: 2.46 nm
- Turn: B: 10-10.6 bp; A: 11 bp - Base pairs: B: perpendicular to helical axis; A: 19 deg. twist - Bases: B: one every 0.332 nm; A: one every 0.23 nm - Overall, B is longer and thinner |
|
|
- When does DNA normally form "A" DNA?
- What is typical about the bp sequence in "Z" DNA? |
- When it is a DNA:RNA hybrid
- alternating pyrimidine/purine sequence, results in a left-handed helix |
|
|
- Where do some people say Z DNA is found in vivo?
- Can B and Z DNA interconvert? |
- Some argue that Z DNA is found in front of promoters
- YES: B DNA has to flip from anti to syn to convert to Z DNA |
|
|
What are two types of supercoils?
|
- toroidal, normally found in chromosomes where DNA is wrapped around histones
- interwound; circular DNA wound in on itself |
|
|
Since EtBr unwinds plasmid DNA, what happens to the linking number?
|
- NOTHING - remains the same because the DNA is unbroken
- since the DNA is unwound, the writhing number increases because the twist number decreases |
|
|
What is more common, + or - superhelical density?
|
- (-), everything except bacteria that live near hot springs in very harsh conditions have (-) supercoiling
|
|
|
How can you tell that it is more common to have (-) or underwound DNA?
|
- You know that W<0 because when you add an intercalating agent such as EtBr, you see a slowing down of the sedimentation velocity of the DNA to where it reaches a minimum
- If you keep increasing the EtBr, the DNA will become overwound and W>0, thus making the DNA run faster through a gradient |
|
|
Do restriction enzymes work better on supercoiled or relaxed DNA?
How can you separate the two? |
- supercoiled
- agarose gel electrophoresis; supercoiled DNA will run faster on the gel |
|
|
What is the difference between type I and II topoisomerases? IA and IB?
|
- I: cuts single strand of DNA; relaxes linking number in increments of 1
- II: cuts double strand of DNA - IA: relaxes (-) supercoiled DNA - IB: relaxes - & + supercoiled DNA |
|
|
How does type I topoisomerase change the linking number?
|
- decreases the linking number by 1 every time it is used (comes closer to the 0 or relaxed state)
- with dsDNA, it changes the T-1 - with ssDNA, it can catenate, or interlink, single stranded circles - does it via a "strand passage" mechanism |
|
|
What is similar about Type IA and Type II topoisomerases? Which requires the hydrolyzation of ATP?
|
- they both function via a strand passage mechanism
- Type II |
|
|
How does ciprofloxacin (Cipro) work?
|
- it inhibits the action of DNA gyrase, which inhibits bacterial DNA replication and RNA transcription
|
|
|
How does DNA gyrase differ from other Type II topoisomerases?
|
- it can relax supercoils, catenate and decatenate dsDNA circles as well as tie knots in them
- Other type II can only relax supercoils |
|
|
Why do many chemotherapeutic drugs attack topoisomerases?
|
- these enzymes are active in these rapidly dividing cancer cells
- inhibiting this enzyme will slow down or stop their metastases |
|
|
What are the names of the five common nucleic acid bases?
|
- adenine
- guanine - cytosine - uracil - thymine |
|
|
What is the difference between nucleosides and nucleotides?
What are the names for the five most common nucleotides and nucleosides? |
- sides only contain ribose sugar, tides contain sugar+phosphate
- Nucleosides: - adenosine - guanosine - cytidine - uridine - thymidine - Nucleotide: AMP, GMP, CMP, UMP, dTMP (since it is never attached to a ribose sugar) |
|
|
What is the distance between bases (between each bases' 1' sugar attachment) in DNA?
|
- 10.85 A
|
|
|
How is DNA replicated? How was this proven?
|
- semi-conservative replication (template is incorporated into "new" DNA strand)
- was proven with "heavy" N-labeled DNA and density gradient ultracentrifugation |
|
|
How can you tell if your solution of DNA is in the native form or denatured?
|
- observe the UV absorbance
- denatured DNA will have a greater relative absorbance than native DNA, especially at 260 nm |
|
|
What is Tm? How does it vary within DNA? How can you determine Tm?
|
- the temperature at which 50% of that same DNA molecule species form a stable double helix and the other 50% have been separated to single strand molecules
- Tm increases with increasing G-C content - Tm = 2(A + T) + 4(G + C) |
|
|
How many rings do pyrimidine bases have? Purine?
Why is DNA an acid? |
- Pyrimidine: 1
- Purine: 2 - the phosphate groups on DNA rapidly give up their protons |
|
|
Why is RNA less stable than DNA?
What is Chargaff's Rule? |
- the ribose sugar has an OH on the 2' carbon that readily attacks the phosphodiester bond
- [A] = [T]; [G] = [C] - [pyrimidines] = [purines] |
|
|
If you denature your DNA sample and determine that the [A]= 15% of the total bases, what can you infer about the concentration of the other bases? Would you expect the Tm to be relatively high or low for this sample?
|
[A]=[T], so
[A]=15% [T]=15% [C]=35% [G]=35% Would expect it to be higher b/c the G/C content is high (70%) |
|
|
Nucleosides normally adopt an ________ conformation about the glycosidic bond.
Which are more water soluble: free bases, or nucleosides? |
- anti
- nucleosides due to the hydrophilic sugar moiety |
|
|
What configuration are the glycosidic bonds of nucleosides?
Where is the glycosidic bond in nucleosides? |
- beta
- between the anomeric carbon on the sugar (1' C) and the N9 of purines or the N1 of pyrimidines |
|
|
Why should DNA not be stored in a low-salt buffer?
What does urea do? |
- The presence of salt such as NaCl stabilizes the (-) charge of DNA and prevents the backbone from destabilizing
- urea is a very powerful protein denaturant, and will also denature DNA as well since it competes with the H bonding between bp |
|
|
If you want to anneal two separate pieces of DNA which share some similar sequences, what is a good way to do this?
|
- heat the solution to boiling (~100 C) for 5' and then cool SLOWLY
- if you cool too quickly, the DNA will form random bp and create this huge blob of mismatched DNA |
|
|
Why is it important for DNA to be packed up around histones?
|
- DNA is very long, and would not fit into the nucleus unless it was packaged tightly
- DNA is also subject to shearing forces that will break it down to about 30 kb oligos |
|
|
If the ionic strength of a solution is increased, what happens to the Tm? What about pH? Is it better to denature with acid or base?
|
- if the pH remains constant, the Tm will increase
- if the pH increases above 10, DNA will denature because the bp potential is destroyed - if the pH decreases below 2.3, there will be excessive protonation of the bases that also disrupts base pairing - Alkali is the preferable denaturant because it doesn't destroy the glycosidic bond |
|
|
The DNA template is read in the __________ direction but the new DNA is made in the ___________ direction.
|
- read: 3'-5'
- made: 5'-3' |
|
|
5'-A-G-A-G-G-T-G-C-T-3' (A)
3'-T-C-T-C-C-A-C-G-A-5' (B) results in mRNA of the following sequence: 5'-A-G-A-G-G-U-G-C-U-3' What is the template strand? What would be the corresponding tRNAs for the codon sequence? |
- template: B
- tRNA1: 5'-UCU-3' - tRNA2: 5'-ACC-3' - tRNA3: 5'-AGC-3' |
|
|
RNA synthesis occurs in the _________ direction.
The AA is attached to the ______ end of tRNA. |
- 5'-3', meaning that the incoming nucleotide is appended to the free 3' OH group of the growing RNA chain
-3' |
|
|
Polypeptides grow from their _____ terminus to _______ terminus.
Primary mRNA transcripts contain ______ and ______. After capping, polyadenylation, and splicing, which make up the "mature" mRNA? |
- N to C
- introns and exons - exons |
|
|
How is transcription controlled in the lac operon?
|
- a repressor constantly binds to the lac operon and prevents transcription
- IPTG is an inducer that binds to the repressor and prevents it from binding to the operon, which allows the RNApol to begin with transcription |
|
|
What is Tm? How does it vary within DNA? How can you determine Tm?
|
- the temperature at which 50% of that same DNA molecule species form a stable double helix and the other 50% have been separated to single strand molecules
- Tm increases with increasing G-C content - Tm = 2(A + T) + 4(G + C) |
|
|
How many rings do pyrimidine bases have? Purine?
Why is DNA an acid? |
- Pyrimidine: 1
- Purine: 2 - the phosphate groups on DNA rapidly give up their protons |
|
|
Why is RNA less stable than DNA?
What is Chargaff's Rule? |
- the ribose sugar has an OH on the 2' carbon that readily attacks the phosphodiester bond
- [A] = [T]; [G] = [C] - [pyrimidines] = [purines] |
|
|
If you denature your DNA sample and determine that the [A]= 15% of the total bases, what can you infer about the concentration of the other bases? Would you expect the Tm to be relatively high or low for this sample?
|
[A]=[T], so
[A]=15% [T]=15% [C]=35% [G]=35% Would expect it to be higher b/c the G/C content is high (70%) |
|
|
Nucleosides normally adopt an ________ conformation about the glycosidic bond.
Which are more water soluble: free bases, or nucleosides? |
- anti
- nucleosides due to the hydrophilic sugar moiety |
|
|
What direction does the ribosome move along the mRNA?
What are the two important sites within a ribosome? |
- moves in 5' to 3' direction
- P site: where the tRNA with the growing AA is attached (is more towards 5' end) - A site: where the "new" tRNA attaches with its single AA to be added to the growing chain (towards 3' end) |
|
|
As a class, what are nucleases?
How do nucleases differ in their specificity? |
- they are phosphodiesterases because they hydrolyze the phosphodiester bond
- nucleases as a whole cut either RNA or DNA - DNases are nucleases that only cut DNA - RNases are nucleases that only cut RNA |
|
|
What is the difference between exonucleases and endonucleases?
|
- exonucleases: degrade nucleic acids by sequentially removing nucleotides from the end
- endonucleases: cleaves at certain places iwthin a polynucleotide |
|
|
UV radiation of 200-300 nm promotes the formation of what in dsDNA?
|
- thymidine dimers on intrastrand portions of DNA
|
|
|
DNA polymerases add a nucleotide donated by a ___________ to the free _________ group of a base paired polynucleotide in the _____________ direction.
|
- nucleoside triphosphate
- 3'OH group - 5' to 3' |
|
|
If you have a circular piece of DNA (400 bp) without any writhes, what are L, T, and W?
If you melt out 30 bases, how do the above numbers change? |
- L: 40, T=400/10=40, W=0
- L remains the same (40); T=(400-30)/10=37, W= +3 |
|
|
What is the role of DNA ligase?
|
- repairs nicks in DNA
- joins Okazaki fragments in semidiscontinuous DNA replication - T4 DNA ligase can link together two duplex DNA via blunt end ligation |
|
|
In addition to its polymerase activity, what else can DNA PolI do?
|
- it can edit its mistakes
- can act as a 3'-5' exonuclease - can act as a 5'-3' exonuclease (this is important to excise RNA primers) - detects nicks and repairs them |
|
|
How does DNA Pol III differ from DNA Pol I?
|
- Pol III can polymerize in 5'-3' direction and can act as an exonuclease in the 3'-5' direction, but it CANNOT act as a 5'-3' exonuclease
|
|
|
What are two ways to separate large DNA on an agarose gel?
|
- pulse field gel electrophoresis is used to separate very large DNA strands
- Orthogonal electrophoresis- change direction of the cathode/anode (DNA goes forward, then pulls back, then goes forward, etc.) |
|
|
Describe how the chain termination (dideoxy) method of DNA sequencing works.
|
- Set up 4 DNA polymerase reactions, each reaction will contain the same known primer, dNTPs, and one ddNTP
- Since ddNTP's can't lengthen the DNA chain, extension stops - Run each reaction out in a separate lane on an agarose gel, and read the sequence from the bottom up (5'-3') |
|
|
How does chloramphenicol work?
|
- inhibits the cell from replicating, but the plasmid DNA will continute to replicate
- allows you to get high copies of plasmid DNA from a low-copy plasmid |
|
|
The pUC18 cloning vector has two selectable markers- what are they?
|
- antibiotic resistance, usually to kanamycin or ampicillin
- blue/white selection: a MCS (polylinker) is within the lacz' operon, if a gene inserts here, you will have white colonies, but if a gene does not insert here, the colonies will be blue (since they express B-galactosidase) |
|
|
Why is it best to purify restriction digest reactions on an agarose gel?
|
- to separate the nicked supercoiled, supercoiled, and cut DNA
- You want to utilize only the cut DNA, which will be the band that runs between nicked supercoiled (runs slower) and supercoiled (runs faster-towards anode (+/red)) |
|
|
What is terminal transferase?
|
- if you have blunt-ended DNA, terminal transferase will add dATP to the free 3' ends
- the reaction is not exact, and there is no good way to control the # of A's that get added |
|
|
Sometimes the DNA fragment you want to clone into a vector does not have the restriction sites you desire. How can you get restriction sites flanking this DNA fragment?
|
- use T4 DNA ligase
- if you synthesize synthetic adaptor oligos with the proper restriction enzyme digest sequence in them, you can ligate them to either end of your DNA insert of interest |
|
|
If the item you put into a bacterial vector tends to be toxic, how can you get the bacteria to express it and still recover the product?
|
- clone the foreign DNA into lambda phages
- can insert up to 15 kb of foreign DNA in between the cos sites of lambda DNA - when the phage packages itself and lyses the cell, the resulting plaques will carry the lambda phage with your DNA fragment of interest |
|
|
How many bp can you insert into the following:
- YAC - BAC - cosmid? |
- YAC: 2 million bp; however, yeast love to recombine and you get lots of deletions
- BAC: 100 kb - cosmid: ~36-51 kb in between cos sites |
|
|
When you create a probe for Southern blots using radioactive labels, where must the P^32 be on the dNTP?
|
- must be on the alpha phosphate, since the gamma and beta are lost
|
|
|
What is the difference between Southern, Northern, Southwestern, and Western blotting?
|
- Southern: detect DNA using a DNA or RNA probe
- Northern: detect RNA using a DNA or RNA probe - Southwestern: detects proteins that bind to DNA (DNA is the probe) - Western: detect proteins using Ab |
|
|
What is a degenerate oligonucleotide probe?
|
- if you don't know the DNA sequence of interest, but do know a portion of the peptide sequence
- use multiple oligonucleotides with the corresponding DNA sequences that would encode for that peptide |
|
|
What is the more common method of sequencing DNA?
|
- using fluorescently-labeled ddNTP's and capillary gel electrophoresis
|
|
|
Describe shotgun sequencing.
|
In shotgun sequencing, DNA is broken up randomly into numerous small segments, which are sequenced using the chain termination method to obtain reads.
Multiple overlapping reads for the target DNA are obtained by performing several rounds of this fragmentation and sequencing. Computer programs then use the overlapping ends of different reads to assemble them into a contiguous sequence. |
|
|
How does cosmid walking differ from shotgun sequencing?
|
Cosmid walking is more directed, and requires the continued use of primers:
1. A primer that matches the beginning of the DNA to sequence is used to synthesize a short DNA strand complementary to the unknown sequence, starting with the primer. 2. The new short DNA strand is sequenced using the chain termination method. 3. The end of the sequenced strand is used as a primer for the next part of the long DNA sequence. |
|
|
What is the main method of oligonucleotide synthesis?
|
- Phosphoramidite synthesis
- Takes place in 3'-5' direction |
|
|
Give an overview of the method of oligonucleotide synthesis.
|
- The first base is attached to the solid support and is blocked so it can't react
- the blocking agent is removed from the 5' site, and the next base to be added is activated and added to the mix - the new activated base reacts with the attached base at the 5' site - the new base is attached to the old base and is capped so it can't react - repeat the entire process until oligo is synthesized, then add heated ammonia to the column to release the oligo |
|
|
How do DNA chips get their oligos?
|
- through photolithographic synthesis
|
|
|
What is the primer that reverse transcriptase requires?
What types of viruses contain reverse transcriptase? In what direction does RT transcribe DNA? |
- tRNA molecule that the virus captures from the host
- RNA tumor viruses (retroviruses) - 5' to 3' direction |
|
|
What type of DNA does reverse transcriptase produce?
What drug inhibits RT? |
- transcribes the RNA plate into a complementary DNA (cDNA) strand to form a ds-DNA:RNA hybrid
- AZT (looks like dideoxy-thymidine) |
|
|
What are the three activities of reverse transcriptase?
|
- RNA-directed DNA polymerase
- RNase-H activity: degrades RNA in the RNA:DNA hybrids - DNA-directed DNA polymerase: makes a DNA duplex after RNase H destroys the viral genome |
|
|
Most mutations act through the context of the gene- what is the exception?
|
- polyglutamine repeat disorders
- get CAG repeat, # of CAG repeats equal the process of disease |
|
|
Name three chemical mutagens.
|
- nitrous acid
- hydroxylamine - alkylating agents |
|
|
What is the difference between a transition and a transversion?
|
- Transition: Pu to Pu or Py to Py
- Transversion: Pu to Py or Py to Pu |
|
|
What is the difference between a selective agent and a mutagen?
|
- a selective agent will HIGHLIGHT species that ALREADY have a particular genome that allows them to survive administration of a selective agent (if E.coli is put on a plate of LB+amp, those colonies that grow have ampicillin resistance, which is a selectable marker to the selective agent, amp)
- a mutagen INDUCES changes in a genome |
|
|
Describe the fluctuation experiment by Luria and Delbruk
|
- the Fluctuation Test demonstrates that in bacteria, genetic mutations arise in the absence of selection, rather than being a response to selection
- They inoculated a small number of bacteria into separate culture tubes and allowed them to grow - they plated equal volumes of these separate cultures onto phage (virus) containing agar - If virus resistance in bacteria were caused by a spontaneous activation, then each plate should contain the same number of resistant colonies - Instead, the number of resistant colonies on each plate varied drastically, supporting the random mutation theory |
|
|
What allows A to sometimes bind to C? What allows A to sometimes bind to G? T and C?
|
- A fluctuates between a normal ketone state (H-bond donor) to a rare imino state, which is an H-bond acceptor
- A in the syn orientation (not normal) can bind to G in the normal anti orientation - T and C can pair if they have help from a water molecule |
|
|
What does 5- bromouracil do?
Oxidative deamination of cytosine gives __________. Oxidative deamination of adenine gives ___________. |
- it is a mutagenic agent that enhances tautomerization
- uracil (results in C-G to T-A transition) - hypoxanthine (results in A-T to C-G transition) |
|
|
What are base analogs?
|
- Examples ar 5-bromouracil, hypoxanthine, and 2-aminopurine
- they can cause point mutations - 5-BU caues A-T to G-C transition - 2-AP causes A-T to C-G transition - hypoxanthine causes A-T to G-C transition |
|
|
What is the purpose of the Ames test?
|
- biological assay to assess the mutagenic potential of chemical compounds
- Expose His- mutants to a potential mutagen - Plate these mutants on media that lacks Histidine - Count the number of colonies you get- the more colonies, the more mutagenic the potential compound is |
|
|
Pyrimidine dimers can be repaired by ________.
|
- photolyase
|
|
|
Describe excision repair.
|
- A damaged base is removed by DNA glycosylase, leaving an AP site (DNA backbone still intact, just missing a base)
- AP endonuclease recognizes the site and cleaves the backbone, causing a gap - DNApol and DNA ligase then repair the gap |
|
|
What are the types of glycosidic bonds in pyrimidine and purine ribonucleosides?
What is depurination? |
- pyrimidine: Beta N1
- Purine: Beta N9 - spontaneous breaking of a glycosidic bond of a purine nucleoside - enhanced by alkylation of N7 |
|
|
Describe methyl directed repair.
|
- Methylation of a new DNA strand is slow
- Methylation "protects" the older strand of DNA and enzymes recognize it as being the "correct" strand - any mismatches between the old and new strand are repaired using the old strand as the template |
|
|
Name where the atoms arise from in the purine ring system.
|
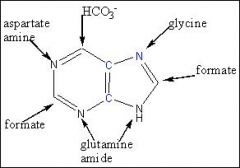
.
|
|
|
What is the immediate precursor to GMP and AMP?
How does de novo synthesis of purines differ from pyrimidines? |
- IMP (inosinic acid)
- purine synthesis occurs as the base is attached to ribose-5-phosphate |
|
|
What is the limiting substance in purine biosynthesis?
|
- PRPP (ribose-5-phosphate pyrophosphokinase)
|
|
|
How are GMP and AMP formed from the precursor?
|
- both are formed in separate two-reaction pathways
- IMP-XMP-GMP; GMP acts as an inhibitor to further GMP biosynthesis - IMP-adenylosuccinate-AMP; AMP inhibits its own production |
|
|
How are pyrimidines synthesized de novo?
|
- cyclization of base to make orotate
- orotate is attached to ribose-5-P to make OMP - OMP is decarboxylated to make UMP - UMP is the precursor for CMP and dTMP |
|
|
What converts NTPs to dNTPs?
|
- Ribonucleotide reductase
- removes OH group from 2' C of NDPs |
|
|
Describe the action of ribonucleotide reductase.
|
- NDP is reduced by ribonucleotide reductase (RR) to dNDP
- oxidized RR is regenerated to reduced RR by thioredoxin - oxidated thioredoxin is reduced by thioredoxin reductase - thioredoxin reductase is reduced by NADPH via FADH2 |
|
|
What is the cycle of dTMP generation?
|
- dUMP gets a methyl group from N5, N10-methylene THF and adds it via the action of thymidylate synthase
- DHF is a byproduct of this reaction, and is reduced by NADPH and DHFR to generate THF - THF gets methyl groups via serine hydroxymethyl transferase, which converts a serine to a glycine - N5, N10 Methylene THF is generated, which can now make more dTMP |
|
|
What is a common 1-C group donor?
|
- tetrahydrofolate (THF)
- THF has many different intermediate states with various levels of oxidation |
|
|
Why are methotrexate, aminopterin, and trimethoprim used as chemotherapeutic agents?
|
- all three of these comounds inhibit DHFR, which is important in the generation of dTMP
- this stops DNA replication, which specifically targets actively dividing cancerous cells |
|
|
Describe the method of HAT selection.
|
- HAT media contains aminopterin, hypoxanthine, and thymidine
- Aminopterin inhibits de novo synthesis of nucleosides, while HPRT and TK supply them from hypoxanthine and thymidine - Only cells which contain HPRT (hypoxanthine-guanine phosphoribosyltransferase) and TK (thymidine kinase) will survive in this media |
|
|
What are the important enzymes in the salvage pathway for purines? Pyrimidines?
|
- HPRT converts guanine and hypoxanthine back to IMP to make more purines
- APRT converts adenine back to AMP - thymidine is rescued by thymidine kinase to make more TMP |
|
|
What is the action of gancyclovir?
|
- viral thymidine kinase from HSV converts ganciclovir to a toxic compound, killing the cell
- this has been used to treat both herpes infections and cancer |
|
|
What is the end product of purine catabolism?
What often builds up in mouse cages and why? |
- uric acid
- ammonia, because bacteria will break down the uric acid to urea and then ammonia |
|
|
What is the major product of pyrimidine catabolism?
|
- malonyl-CoA
|
|
|
When does recombination normally occur?
Is recombination mutagenic? |
- high rate during meiosis
- no, because no new alleles are created by recombination |
|
|
What results in a tetratype?
|
- A single crossover event between A B and a b parents will give a tetratype
- progeny are AB, Ab, aB, and ab |
|
|
Explain the difference between homologous and nonhomologous recombination.
|
- homologous: reaction between two homologous chromosomes that results in a rearrangement of similar DNA
- nonhomologous: incorporation of DNA segment whose seqence differs greatly between the point of insertion (i.e. info from chromosome 12 inserts into Ch. 13) |
|
|
What experiment illustrated that recombination could occur?
|
- Meselson experiment with density ultracentrifugation
- put "heavy" and "light" DNA through cycles of replication and then centrifuged it - found light and heavy parental bands, but with LOTS of intermediates, which resulted from different regions of the DNA recombining |
|
|
Define aberrant segregation.
|
- The aberrant patterns are interpreted as the result of single strand exchanges prior to the first meiotic division
- exchanges result in heteroduplexes (double helices containing mismatches) - The different patterns could result from varying numbers of mismatch repair events (if the crossovers are identified and repaired): No repair ---> aberrant 4:4 One repair ----> 5:3 or 3:5 Two repairs ---> 6:2 or 2:6 |
|
|
Explain how recombination shows polarity.
|
- Unusual segregation patterns suggest that initial strand exchange during recombination may be asymmetric
- One end of the gene (the promoter end) has a high frequency of such events (hotspot), the other a low frequency |
|
|
What is a failure of DNA repair?
|
- post-meiotic segregation
- if DNA always repaired itself, it would recognize recombination events and prevent that DNA from replicating further |
|
|
Define gene conversion.
|
It is a process by which DNA sequence information is transferred from one DNA helix (which remains unchanged) to another DNA helix, whose sequence is altered.
|
|
|
What are some problems with the Holliday model?
|
- you should see events occuring at the same time and be symmetrical
- however, there are few double crossovers, few abberant 4:4m, 5:3m - initiation sites can be heterozygous and still stimulate recombination |
|
|
How does the Meselson model differ from the Holliday model?
|
- with both models, only one DNA strand is nicked
- M: DNApol starts to repair the nick and displaces the strand, does not result in an equal exchange of information between chromosomes. |
|
|
What are problems with the Meselson model?
|
- initiation sites accept information, even though the M model has the initiating chromosome donating information
- crossovers are mapped to both sides of the initation site, where the M model would only have crossovers on one side |
|
|
What theory resolved some of the problems with the Holliday and Meselson models of replication?
|
- instead of a single strand of DNA being cut, the theory was that both DNA strands are cut
- this allows for 2 Holliday junctions, so crossover could occur anywhere |

