![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
30 Cards in this Set
- Front
- Back
|
Metabolism |
•Living Organisms must be able to take energy & raw material in the environment & convert it to energy/material that the organism can use •Chemical reaction occurs w/in an organism |
|
|
Catabolism |
•Breaks Down large molecules into smaller ones •Release Energy (exergonic) •Energy coupled to other reactions (e.g. ATP production, synthesis of organic material) •Energy released as heat |
|
|
Anabolism |
•Energy (endergonic) to make large molecules from smaller ones •Couple High energy reactions to anabolic reactions to provide the required energy |
|
|
Oxidation |
•Loss Of electrons by a molecule, ion or atom •Gain Of oxygen atoms or loss of hydrogen atoms by a molecule |
|
|
Reduction |
•Gain of electrons by a molecule, ion, or atom •Loss Of oxygen atoms or gain of hydrogen atoms by a molecule |
|
|
Enzymes |
Proteins That catalyze reactions w/out being consumed |
|
|
Enzyme Classification: Hydrolase |
Catabolize Molecules by adding water in a decomposition reaction |
|
|
Enzyme Classification: Isomerase |
Rearrange Atoms within the molecule but do not add or remove anything |
|
|
Enzyme Classification: Ligase |
Join two or more molecules together |
|
|
Enzyme Classification: Lyases |
Split large molecules into smaller ones with using water |
|
|
Enzyme Classification: Oxidoreductase |
Remove Or add electrons to substrates |
|
|
Enzyme Classification: Transferase |
Transfer Functional groups between molecules |
|
|
Enzymes: Cofactors |
•Non-protein molecules that aid in the catalytic activity of enzymes •Inorganic molecules (e.g. Iron, Magnesium, Zinc) |
|
|
Enzymes: Coenzymes |
•Organic molecules (Vitamins) |
|
|
Apoenzyme |
A protein that requires cofactors to function, but is not bound to them |
|
|
Holoenzyme |
A apoenzyme w/ cofactors attached & is active |
|
|
Enzyme Activity |
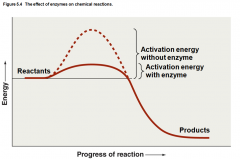
Catalyze Reactions by lowering the activation energy |
|
|
Induced Fit Model |
•Substrate Initially fits loosely to the enzyme •Resulting Compound is the enzyme-substrate complex •Binding Of the substrate induces to change its shape to conform the substrate |
|
|
Enzyme Kinetics |
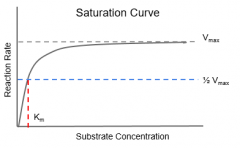
Rate of enzyme reactions |
|
|
Enzyme Kinetics: Vmax |
Maximum Rate achieved by the system |
|
|
Enzyme Kinetics: Km |
Substrate Concentration at ½ Vmax -a measure of the affinity of the substrate to the enzyme -inverselycorrelated to the affinity |
|
|
Enzyme Inhibitors |
•Activity can be blocked by the binding inhibitors Two Types of Enzyme inhibitors •Competitive Inhibitor •Noncompetitive inhibitors |
|
|
Competitive Inhibitor |
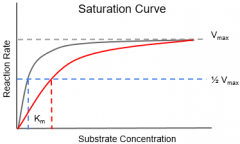
•Binds to the active site of the enzyme •Increasing The concentration of substrate can overcome competitive inhibition |
|
|
Non-Competitive Inhibitor |
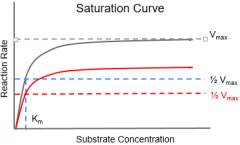
•(Allosteric) binds to enzyme on site other than the active site •Conformational change in enzyme & renders it unable to bind to the substrate |
|
|
Carbohydrate Metabolism |
•Glucose Is the primary source of energy for most organisms •The First step is glycolysis |
|
|
Glycolysis |
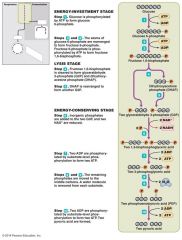
•Breakdown Of glucose •Does Not require oxygen |
|
|
Pyruvate Decarboxylation |
•1 carbon is removed from pyruvate •Molecule of Coenzyme A binds to the 2 carbon molecule •Acetyl CoA 1 |
|
|
Citric Acid Cycle |
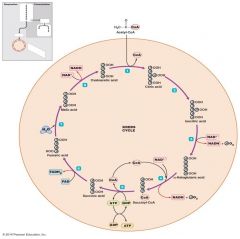
•AKA Tricarboxylic acid cycle/ the Krebs cycle •Acetyl CoA (2C) joins the Citric acid cycle by binding to the 4 carbon oxaloacetate •3 NADH/ FADH2/ 1 GTP |
|
|
1 possible arrangement of Electron Transport Chain |
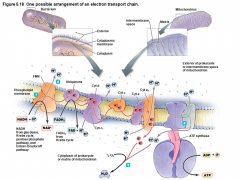
•Mitochondrian Eukaryotes 2 molecules energy •6:2:2 Glycolysis •2 Pyruvate/ 1 Acetyl CoA •1 glucose 2 Acetyl CoA |
|
|
Fermentation |
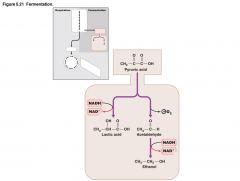
•Cannot Undergo oxidative phosphorylation under anaerobic conditions •Cells Require constant source of NAD+ •Pathways provide cells regenerate NAD+ |

