![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
56 Cards in this Set
- Front
- Back
|
List cytoskeletal systems and functions of bacteria and archaea |
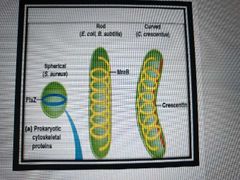
MreB (like microfilaments) for dna segregation and cell shape - crescentin makes rod shaped bacteria curved FtsZ (like microtubules) for regulating division |
|
|
Types of microtubules and their location/funct |
Cytoplasmic - found in cytosol - maintains axons - maintains cell shape - forms mitotic/meiotic spindles (chromosome movement) - moves and places vesicles Axonemal - found in cilia, flagella, and the basal bodies they attach to (think sperm) - forms axoneme, a highly ordered bundle of microtubules that form the central shaft of cilia and flagella - responsible for cell motility |
|
|
What is the structure of microtubules? |
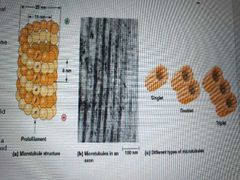

Straight hollow cylinders of tubulin with longitudinal arranged polymers called protofilaments - protofilament = alphabeta-heterodimer noncovalently bonded -alpha and beta tubulins have almost identical 3d structure but only 40% amino sequence match - alpha and beta each have n-terminal GTP binding domain, central colchicine binding domain, and c-terminal MAP domain (microtibule-associated proteins) |
|
|
What are tubulin isoforms? |
alpha and beta tubulin subunits with closely related but not identical Gene's (different isoforms may interact with different proteins) |
|
|
Explain microtubule polarity |
Dimers that make up microtubules are all oriented in the same way (n-terminus to c-terminus) so the entire microtubule has a positive end and negative end. The plus end grows faster |
|
|
What are singlet, doublet, triplet microtubules? |

Singlet - cytoplasmic Doublet and triplet - axonemal - one 13-protofilament A tubule and one or two 10/11 protofilament B and C tubules |
|
|
Describe microtubule assembly |
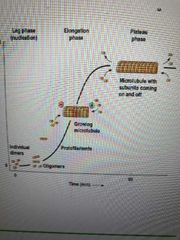
****with gtp and Mg2**** Lag (slow) phase: Nucleation - dimers aggregate into oligomers Elongation (fast) phase: Addition of subunits at either plus or minus end Plateau phase: Disassembly due to diminished amt of free tubulin |
|
|
What is critical concentration? When does growth occur? |
The tubulin concentration at which assembly equals disassembly. Growth occurs when tubulin concentration is above critical concentration |
|
|
How can microtubule plus and minus ends be visualized? |
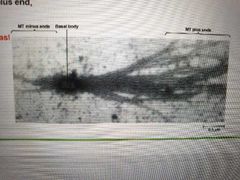
Mix basal bodies (structures at base of cilia) with tubulin heterodimers. Plus end will grow faster |
|
|
What is treadmilling? |

Plus and minus ends have different critical concentrations. Treadmilling - plus end is above critical concentration (growing) while minus end is under (disassembling) |
|
|
List drugs that affect microtubule assembly |
Colchicine - binds to tubulin monomers so they cant assemble into microtubules. Promotes disassembly
Nocodazole - also inhibits assembly but effects are more easily reverse than colchicine |
|
|
Explain dynamic instability (microtubules) |
each heterodimer has 2 GTP's - one on the alpha tubulin and one on the beta.
Once added to microtubule, beta's gtp hydrolyzes to gdp
If there is plenty of GTP-tubulin, it adds to the plus end to create a gtp cap.
If gtp-tubulin is low, new dimers are added at a slower rate than the hydrolysis of beta's GTP, leading to catastrophe (disassembly stops growth in favor of shrinkage). Reversing this effect (going from shrinkage to growth) is called rescue
See youtube Microtubule Assembly & Disassembly Mechanisms by Medical Sciences Animations |
|
|
Where do microtubules originate? |
From a microtubule organizing center (MTOC) that anchors and nucleates microtubules - Usually a centrosome of 2 centrioles surrounded by pericentriolar material. Y-tubulin ring complexes (y-TuRC) nucleate microtubule assembly away from the centrosome with minus ends towards the MTOC (this means growth and shrinkage happens at the plus end) |
|
|
What is the structure of a centriole? Its function? |
9 triplet microtubules formed at right angles Funct - helps form basal bodies, which help form cilia and flagella - no centriole = poorly formed mitotic spindles |
|
|
What are MAPs? List types and funct |
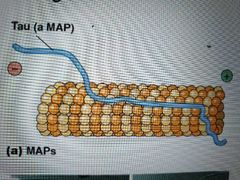
Microtubule-associated proteins. Affects density of microtubule bundles; one region binds at intervals along microtubule wall and the other end extends out to allow interaction with other cell structures and filaments. Length of extension determines spacing/tightness of bundles
Tau Map - causes microtubules to form tight bundles in axons
Map2 - causes loose bundles in dendrites |
|
|
List proteins that encourage/discourage catastrophe |
+-TIPs (Tubulin Interacting Proteins) such as EB1 (End binding protein 1) protect growing plus ends Stathmin/Op18 prevents polymerization by binding to heterodimers Catastrophins promote subunit peeling at microtubule ends Katanin cleaves microtubules |
|
|
Functions of microfilaments |
-Muscle contraction -Cell migration (cell is made and moves to where it performs funct) -Amoeboid protozoan movement -Cytoplasmic streaming (cytoplasm moves) -develop and maintain cell shape (from cell cortex beneath plasma membrane) -forms structural core of microvilli |
|
|
How are microfilaments formed? |
Actin protein enter lag phase and elongation phase to form globular actin/G-actin that binds ATP or ADP (process is reversible)
2 linear G-actin form a helix (F-actin/microfilaments) ***all actin monomers are in the same orientation*** |
|
|
List types of actin |
Alpha actins- muscle specific Beta and Y actins - nonmuscle actins, (B and Y are localized to different regions of the same cell) |
|
|
Describe microfilament polarity |
Can be seen by incubating myosin subfragment 1 (S1) onto microfilaments and observing the resulting arrowhead pattern
Barbed end - plus (faster addition and loss) Pointed end - minus
Like microtubules, atp hydrolyzes as g-actin monomers bind. Growing end has ATP-actin, the rest has adp-actin |
|
|
List drugs that affect microfilament polymerization |
Cytochalasins - fungal metabolites that block addition of new monomers Latrunculin A - toxin that sequesters monomers and prevents their addition Phalloidin - stabilizes microfilament. Prevents depolymerization |
|
|
List structures that microfilaments can form |
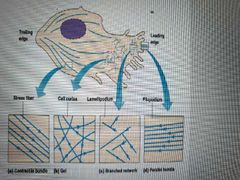
Immobile cells adhered to underlying substratum - thick organized bundles (stress fibers)
Mobile cells in cell cortex beneath membrane - crosslinked gel
Extremely mobile Cells that crawl- parallel straight filopodium and branched lamellipodium |
|
|
How is location and structure of microfilament formation controlled? |
Via actin-binding proteins at nucleation, elongation, and severing stages Microfilaments assemble as long as ATP-g-actin is high, but in the cell, it is sequestered (bound) by thymosin B4. Profilin competes with thymosin to bind g-actin and does not sequester it. More profilin = more microfilament formation |
|
|
List microfilament cap proteins |
CapZ- binds to plus end, prevents addition of subunit Tropomodulins -binds to minus ends, prevents loss of subunits |
|
|
What is filamin? |
Protein that cross links microfilaments by splicing them together where they intersect |
|
|
Funct of intermediate filaments |
Abundant in animal cells as keratin Supports entire cytoskeleton as most stable and least soluble (more detergent, chemical resistant) cytoskeleton component Forms nuclear lamina (fibrous mesh in dna replication and cell division) Involved in cell structure scaffolding |
|
|
List classes of intermediate filaments |
1. Acidic keratin (epithelial surface) 2. Basic/neutral keratin (epithelial surface) 3. Vimentin (connective tissue) Desmin (muscle cells) Glial fibrillary acidic protein (glial cells) 4. Neurofilament proteins (nerve cell) 5. Nuclear lamina A B and C (network along inner surface of nuclear membrane) 6. Nestin (nerve cell neurofilaments of embryos) |
|
|
How are cytoskeletal elements integrated? |
Spectraplakins - linker proteins that connect MT, MF, IF -plectin found where IF connects to MT and MF |
|
|
How is the cytoskeleton mechanically integrated? |
MT resist bending during compression MF contracts and generates tension IF are elastic and resist tension |
|
|
Describe intermediate filament structure/assembly |
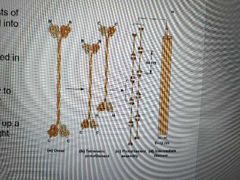
Fibrous (rather than globular) dimers with N and C terminals that differ greatly among IF proteins
1. Two parallel IF polypeptides form a coiled coil dimer 2. Two dimers align to form a tetrameric protofilament 3. Protofilaments overlap to build an intermediate filament |
|
|
List signaling molecule classifications |
Classified by distance: Long - endocrine signals produced far from target tissue; travel via circulatory system
Short - paracrine signals that are diffusable
Physical contact between sending and receiving cells - juxtacrine signals
Signals sent and received on same cell - autocrine signals |
|
|
What are receptors and ligands? |
Receptor - area on cell surface that binds a messenger; has a binding site or binding pocket Ligand - a messenger that binds to the target cell |
|
|
What are 2nd messengers? |
1. Ligand binds to receptor 2. Binding induces production of additional molecules/ions in the cell (2nd messengers) that relay signals from one location in cell to the interior, causing a cascade of changes |
|
|
What is signal transduction? |
Ability of a cell to respond to ligand-recrptor binding by changing behavior or gene expression |
|
|
How are receptors altered after ligand binding? |
They change shape or cluster together. Then, the receptor starts a preprogrammed chain of signal transduction events (cell changes behavior or gene expression), depending on cell's history, that lay dormant until they are triggered |
|
|
What are agonists and antagonists? |
Synthetic ligands that bind more tightly or selectively than real ones. Agonist - receptor activators Antagonist - receptor blockers |
|
|
What gives ligand-receptor binding its specificity? |
Binding site/binding pocket fits the messenger closely, like a puzzle piece Amino acid side chains positioned to form chem bonds with the messenger |
|
|
How do cells shut down signaling? |
- reduce amount of ligands - reduce receptor sensitivity (receptor desensitization) or amount. -Cells sense changes in ligand concentration. Cell will stop responding to a certain ligand when receptors are already preoccupied for a long time (2 ways) so ligand concentration must increase to restimulate the cell 1.lower density of receptors by removing them via receptor-mediated endocytosis 2. Biochem changes that reduce affinity for ligand (ex: add or remove phosphate groups to amino acids) |
|
|
What is signal amplification? Examples? |

A small amt of ligand causes target cell to undergo a cascade of events. At each intermediate, more product is made. Ex: epinephrine causes glycogen breakdown in liver cells. 1 epinephrine releases hundreds of millions of glucose from glycogen |
|
|
List cell-cell signal pathways |
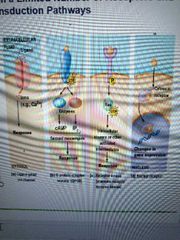
1.Hydrophilic transmembrane receptors - Ligand-gated ion channels, proteins, small peptides, aminos and derivatives, nucleotides or nucleosides
2. G protein-coupled receptors - hydrolysis of GTP
3. cytosolic enzyme receptors - protein kinases
4. Hydrophobic Nuclear receptor - receptor bound to ligand (steroids and retinoids) enters nucleus to regulate gene transcription
|
|
|
What do g protein-coupled receptors cause? Example? |
1. Ligand binding changes conformation change of receptor, which activates guanine nucleotide binding protein 2. Activated g-protein binds a target protein and changes its activity Ex: opioid receptors |
|
|
What is the structure of a g protein-linked receptor? |
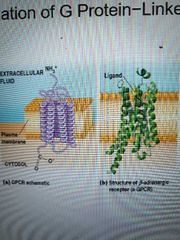
7 transmembrane alpha helices connected with alternating cytosolic or extracellular loops and a unique extracellular messenger binding site. All have similar structure but different amino acid sequences |
|
|
How does phosphorylation regulate g protein-linked receptors? |
G protein-linked receptor kinase phosphorylates cytosolic amino acids on activated receptors.
Ex: amino acid such as B-adrenergic receptor is phosphorylated, then B-arrestin binds to it and inhibits it from interacting with g-protein Ex: protein kinase A is activated by g protein-mediated signalling but then it phosphorylates (inhibits) other amino acids on the receptor; this is negative feedback |
|
|
How are g proteins activated or inactivated? List structure of g proteins |
G proteins turn on and off depending on whether they are bound to GTP or gdp
Structure - small monomeric proteins or large heterotrimeric proteins with Ga, Gb, and Gy subunits -Ga is largest and binds to gdp or GTP |
|
|
Describe g protein activation/inactivation cycle |
1. Ligand binds to g protein-linked receptor 2. Receptor changes conformation and a g protein binds and releases its gdp 3.Ga binds to new Gtp molecule and detaches 4. Depending on g protein, either Ga or Gby initiates signal transduction 5. G protein stays active until Ga hydrolyzes GTP to GDP and rejoins Gyb (some Ga's are not efficient at hydrolysis) 6. If not efficient, regulators of g protein signaling (RGS) proteins are GTPase activating proteins that bind Ga to stimulate hydrolysis |
|
|
What is the most important g protein function? |
Formation of 2nd messengers, identified by the signaling process, such as cyclic AMP |
|
|
Example of Gby subunit signaling |
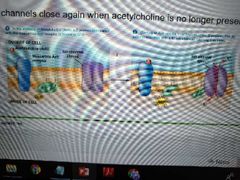
1.Acetylcholine binds muscarinic acetylcholine receptor 2. Gby subunit causes potassium channels to open, and close once acetylcholine is no longer present |
|
|
Describe regulation of cyclic AMP by g proteins |
1. Ligand binds to receptor and g protein attaches, losing gdp 2. Ga attaches to GTP and leaves to bind adenylyl cyclase 3. Adenylyl cyclase makes cAMP (a second messenger) from cytosolic ATP until Ga hydrolyzes and rejoins Gby 4. Remaining cAMP is degraded by phosphodiesterase |
|
|
Main function of cAMP? |
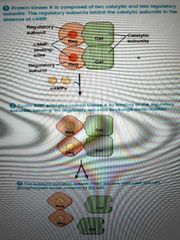
Binds to regulatory subunits of protein kinase A, causing them to dissociate from catalytic subunits, which are then free to phosphorylate target cell proteins on serine or threonine residues using ATP |
|
|
Diseases caused by disruption of g protein? |
Cholera -gs can't hydrolyze whooping cough - pertussis keeps gi from inhibiting adenylyl cyclase |
|
|
How do g proteins act through inositol? |
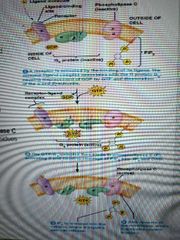
1. Ligand binds receptor and activates Gq protein 2. Gq activates Phospholipase C (Cbeta), which cleaves PIP2 into IP3 second messengers and diacylglycerol (DAG) 3. IP3 diffuses through cytosol and binds ligand-gated calcium channel (IP3 receptor channel) 4. Calcium is released into cytosol 5. Calcium and DAG activate protein kinase C, which phosphorylate serine and threonine residues on various target proteins |
|
|
How do protein kinase-associated receptors function as kinases? |
Ligand binding stimulates tyrosine or serine/threonine kinase activity, and they transmit signals through phosphorylation cascade |
|
|
What are growth factors? List the ones that stimulate receptor tyrosine kinases |
Cells in vitro wont grow without blood serum with growth factor messengers Insulin Insulin-likengrowth factor 1 Fibroblast growth factor Epidermal growth factor Nerve growth factor |
|
|
What is the structure of a tyrosine kinase? |
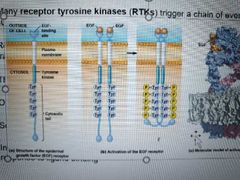
1 polypeptide chain with 1 transmembrane segment. Single - Ligand-binding domain outside, tyrosine kinase domain inside Double - receptor and kinase are separate proteins. Non receptor tyrosine kinase binds to receptor and activates once ligand binds |
|
|
Describe autophosphorylation of tyrosine kinases |
1. Ligand binds 2. Some (fibroblast gf) receptors dimerize and phosphorylate each other 3. Some (epidermal gf) receptors cluster and the tyrosine kinase of each receptor phosphorylates its neighboring receptors |
|
|
Describe Ras signaling |
1. Autophosphorylation 2. Receptor recruits cytosolic proteins with SH2 amino acid domain to phosphotyrosine 3. GRB2 protein from the SH2 domain binds to Sos, a guanine-nucleotide exchange factor 4. Sos stimulates Ras G protein to release gdp and bind gtp ***some receptor tyrosine kinases activate phospholipase Cgamma*** 5. Ras triggers a series of phosphorylations, starting with Raf 6. Raf phosphorylates serinenand threonine residues in MEK protein kinase 7. MEK phosphorylates threonine and tyrosine residues in mitogen-activated protein kinases (MAP kinases), activated to grow and divide by mitogens 8. Map kinases phosphorylate transcription factors 9. GTPase activating protein facilitates Gtp hydrolysis. GDP-bound Ras inactivates
|

