![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
35 Cards in this Set
- Front
- Back
- 3rd side (hint)
|
In an emergency room situation such at the Lawrence Hospital, which of the following would likely be the biggest drawback to identifying a bacterial species using the API-20E procedure we used in this class? |
Long test incubation time/wait for results (API-20E test results took ~ 24 hours) |
|
|
|
From our laboratory manual, regarding the API-20E test: "Suppose, after 24 hours, you notice no growth in the tube containing mineral oil. Assuming that everything is behaving properly under these conditions, what do you know about this organism and what predictions can you safely make about its performance on the fermentation tests and nitrate reduction test? Is it a member of Enterobacteriacae? |
Expect negative for fermentation and nitrate reduction; and no, would NOT be Enterobacteriacae |
|
|
|
On the API-20E test strip, if the first seven test results were: -++++-+ a possible identification code would be: A. 6318902 B. 6305320 C. 7254233 D. 5324322 E. 6313102 |
E. 6313102 |
1-2-4 |
|
|
On the API-20E test strip, if a bacterial species had an identification code 1553277, the last five test results on the API-20E strip would be expected to be? |
+++++ |
+ , - |
|
|
The test we use in this class for differentiation of species during the mixed double unknown project were: |
Qualitative |
|
|
|
In the Kirby-Bauer antibiotic disk sensitivity testing procedure, which of the following processes creates a gradient of antibiotic concentration? |
Passive diffusion (random movement of substances through semi-solid media) |
|
|
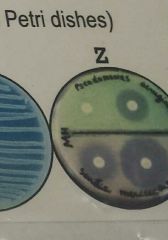
On the antibiotics disc diffusion test result (see image Z), which disc has the smallest zone of inhibition? |
Top left |
|
|
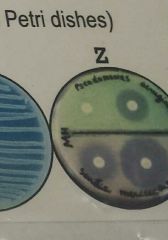
Based on the antibiotic disc diffusion test result (see image Z), the drug on which disc would be the most appropriate drug for treating an infection in humans? |
Not enough information to determine |
|
|
|
In the Kirby-Bauer antibiotic disk sensitivity testing procedure, antibiotic - resistant colonies of bacteria are most likely to be observed near the Blank of the zone of inhibition. |
Periphery (outside edge) |
|
|
|
You have inoculated 10 uL of a sample that had been diluted by a factor of 10^-3 on a nutrient agar plate. After incubation, you count 58 colonies. What was the original cell density. CFU/mL |
5,800,000 |
|
|
|
Suppose you have a culture of 1×10^9 bacteria. An antimicrobial drug is applied which is effective against 99.9% of these bacteria. Calculate the number of bacteria you expect to be resistant to the treatment. |
1,000,000 |
|
|
|
The original concentration in a sample is 2×10^7 CFU/mL. Application of which sample volume would, in theory, yield a plate having between 30 and 300 colonies? |
10 nL |
|
|
|
What is the indication of a positive PYR test result? |
PYR disc turns red in color |
|
|
|
Which of the following are composed of double-stranded DNA? A. Jellyfish proteins B. pGLO plasmids C. Agar D. Antibodies E. mRNA transcrpits |
B. pGLO plasmids |
|
|
|
In a successful pGLO transformation experiment, which gene product enables the transformed cells to grow on LB/amp and LB/amp/ara? |
Beta-lactamase |
|
|
|
What was the pGLO plasmid composed of? A. Virus B. Bacteria C. Protein D. DNA E. Amino Acids |
D. DNA |
|
|

In this image, which of the samples show a positive result for oxidase and catalase activity? |
Both on right side. |
|
|
|
What does "selective toxicity" mean with respect to antibiotic drugs? |
Toxicity varies by species |
|
|
|
Which of the following contained (or "displayed") the epitopes that are recognized in an agglutination reaction? A. The antisense DNA sequence B. The Antigen C. The isotope D. The antagonist E. The antibody |
B. The Antigen |
|
|
|
The locations where chemically-induced DNA replication errors occur are? |
Randomly distributed |
|
|
|
How would you expect chemical MUTAGENICITY to affect growth of Ames-strain Salmonella typhimurium? |
Induce mutations |
|
|
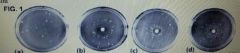
In the figure above, assume different concentrations of a single chemical were applied to plates B, C, and D. Which of the plates had the highest concentration of the test chemical? |
Plate B had the most |
|
|
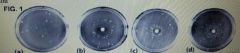
In the figure above, assume a single concentration of different chemicals was applied to plates B, C, and D. Which of the following conclusions (regarding the chemical tested on plate C) would be most supported? |
It is mutagenic |
|
|
|
How would you expect chemical TOXICITY to affect growth of Ames-strain Salmonella typhimurium? |
Prevent growth |
|
|
|
In a typical Ames test procedure, approximately what percent of mutations result in rescue of auxotrophic mutants? |
Very few - less than 1% |
|
|
|
Which plates do you need to examine to determine if your test substance is mutagenic? |
The minimal media plate |
|
|
|
Why can't your test substance [the potential mutagen] contain proteins? |
Bacteria could obtain histidine from the test substance |
|
|
|
List these in order from smallest to largest (in terms of physical size)? Prokaryotic chromosome, eukaryotic chromosome, operon, gene, plasmid |
Gene, operon, plasmid, Prokaryotic chromosome, Eukaryotic chromosome. |
|
|
|
After E. coli is successfully transformed using the pGLO plasmid, where is the gene for GFP? |
Inside the E. coli cell, but not in bacterial genome |
|
|
|
Why does the LB/amp/ara/+DNA plate fluoresce when the LB/amp/+DNA plate does not? |
Because the GFP is only expressed in presence of arabinose. |
|
|
|
Which of the following could cause false species identification when using a dichotomous key? A. A transformation / conjugation event previously occurred B. The test specimen / culture contained multiple species C. Improper dichotomous key used D. A test result was misread E. Any of the answers shown |
E. Any of the answers shown |
|
|

Using the image above, if you had tested an unknown on the right hand side of the mannitol salt plate, the right-hand side of the DNase plate, and the area marked "A" on the blood agar plate, which of the following species would be the most likely identification? A. S. Aureus B. S. Pyogenes C. S. Agalactiae D. None of these E. All of the answers |
A. S. aureus |
|
|
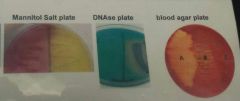
Using the image above, if you had tested an unknown on the left-hand side of the mannitol plate, the left-hand side of the DNase plate, and the area marked "C" on the blood agar plate, which of the following species would be the most likely identification? A. S. Aureus B. S. Pyogenes C. S. Agalactiae D. None of the answers E. All the answers shown |
D. None of the answers |
|
|
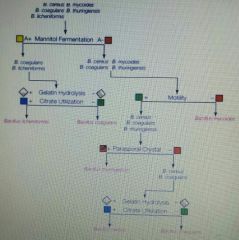
Based on the dichotomies key shown below, a specimen that was non-motile, positive for citrate utilization, and negative for gelatin hydrolysis could be: A. B. myocoides B. B. thuringensis C. B. lichenformis D. B. coagulans E. B. cereus |
A. B. myocoides |
|
|
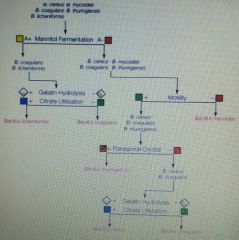
Based on the dichotomous key shown below, a specimen that was citrate utilization positive, mannitol fermentation negative, parasporal crystal negative, and positive for motility could be: A. B. myocoides B. B. thuringensis C. B. lichenformis D. B. coagulans E. B. cereus |
E. B. cereus |
|

