![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
136 Cards in this Set
- Front
- Back
|
Where does replication begin along a chromosome or plasmid?
|
origin of replication
|
|
|
Whose experiments lead to the finding that the replication fork was bi-directional?
|
Cairns
|
|
|
In Cairns experiment he observed tritium labeled ___________ was incorporated into ______ loops durind DNA replication in bacteria.
|
thymidine, two
|
|
|
In Cairns "pulse-chase" experiments he revealed that ___________ of the replication fork was ____________.
|
movement, bi-directional
|
|
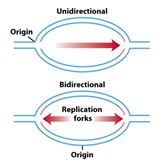
Which image is correct?
|
the bi-directional one
|
|
|
How would you perform a "pulse chase" experiment?
|
In biochemistry and molecular biology, a pulse-chase analysis is a method for examining a cellular process occurring over time by successively exposing the cells to a labeled compound (pulse) and then to the same compound in an unlabeled form (chase). Radioactivity is a commonly used label.
|
|
|
Whose experiments concluded that chromosomes or plasmids have an origin of replication?
|
Inman's experiments
|
|
|
Inman's experiments used denaturation loops as reference marks on plasmids called ____________ ___________.
|
denaturation mapping
|
|
|
Inman concluded that for a given plasmid, replication ____________ at the same place every time. What is this called?
|
originated, point of origin of replication
|
|
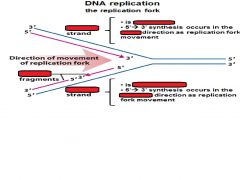
Fill in the blanks.
|
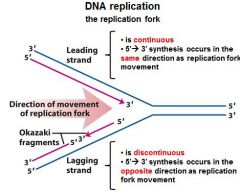
|
|

Fill in the blanks.
|

|
|
|
DNA replication requires an unpaired _________ to copy, or what is called a __________ strand.
|
template, parent
|
|
|
DNA replication requires a primer to provide a free ______.
|
3' OH
|
|
|
DNA replication can add only onto the 3' OH so the strand always grows in a ____________ direction.
|
5'--->3'
|
|
|
Draw the DNA polymerase reaction mechanism. Include the amino acids involved and the metal ions.
|
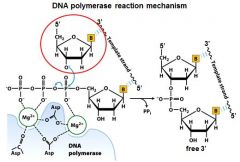
|
|
|
The primers used to start DNA synthesis are created by the enzyme ______________>
|
primase
|
|
|
Primers created from primase are RNA/DNA?
|
RNA
|
|
|
The primers used to start DNA synthesis are small ____ fragments, which are ________ nucleotides in length.
|
RNA, 1-60
|
|
|
Primase primers are ultimately (in the end of DNA synthesis) ____________, and replaced with _______.
|
hydorlyzed, DNA
|
|
|
Which replication strand needs multiple primers?
|
Lagging strand
|
|
|
In E. coli the error rate in DNA synthesis is about 1 per __________ nucleotides incorporated into DNA.
|
10^9-10^10
|
|
|
The level of fidelity of E. coli DNA polymerase is insured by the ability of the enzyme to discriminate between __________ and __________ base pairing.
|
correct, incorrect
|
|

This question is for E. coli DNA polymerase.
|

|
|
|
When a nuclease hydrolyzes DNA/RNA it can yield what kind of ends?
|

|
|
|
Exonucleases hydrolyze from ____________________ of the nucleic strand.
|
one end or the other
|
|
|
Can an exonuclease hydrolyze from either end of the nucleic strand?
|
yes
|
|
|
Exonucleases act on _________ strand and ___________ strand DNA/RNA.
|
single, double
|
|
|
Endonucleases hydrolyze from ___________ the strand.
|
within
|
|
|
Restriction __________ are used in cloning to cut the dsDNA.
|
endonucleases
|
|
|
DNA polymerases have nuclease activities for what functions?
|
proofreading function, repair function
|
|
|
The proofreading function of DNA polymerases does what?
|
corrects immediate errors in replication by acting as a 3'--->5' exonuclease
|
|
|
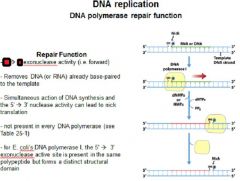
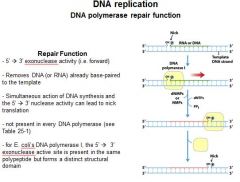
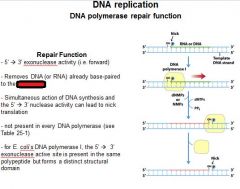
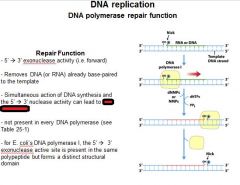
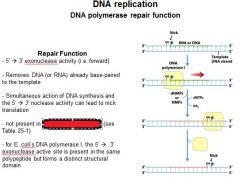
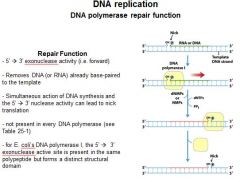
The repair function of DNA polymerases does what?
|
corrects errors that lie ahead of the replication fork acting as a 5'----->3' exonuclease.
|
|
|
The proofreading function and the repair function use what kind of nuclease?
|
exonuclease
|
|
|
What strand direction does the proofreading function work?
|
3'--->5'
|
|
|
What strand direction does the repair function work?
|
5'---->3'
|
|
|
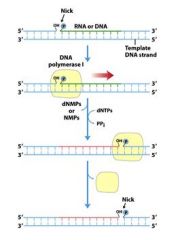
Define nick translation?
|
|
|
|
|
|
|

|
|
|

|
|
|

|
|

|
|
|
|
DNA poly1 is a ___________ polypeptide?
|
single
|
|
|
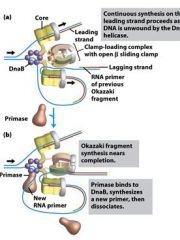
DNA helicase use the energy derived from _____ hydrolysis to _______ and ________ alond dsDNA.
|
ATP, unwind, translocate
|
|
|
The DnaB helicase of E. coli is a __________ structure with each subunit containing its own _______ domain.
|
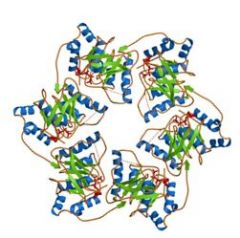
homohexameric, ATPase
|
|
|
DNA helicases have a ________ effect, one subunit _______ when bound to ATP, while the next subunit _________ when bound to ADP.
|
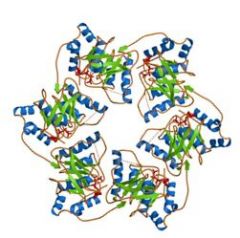
rotary
grabs pulls |
|
|
The rotary effect of a DNA helicase pushes one strand thru the _______, and one strand ________ ________ _______.
|
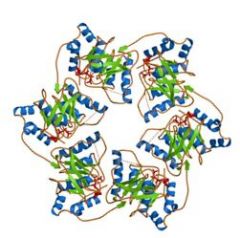
center of the enzyme, around the outside
|
|
|
Coordination between adjacent subunits causes an ATP _________ cycle that pulls ssDNA through the __________ of the enzyme structure.
|
hydrolysis, hole in the center
|
|
|
All enzymes and proteins associated with DNA/RNA replication is called the?
|
Replisome
|
|

This man isolated DNA pol1 from E.coli in 1955.
|
Arthur Kornberg
|
|
|
After complete characterization of DNA pol1 it was realized or suggested that DNApol1 __________ be the only DNA polymerase involved in replication.
|
can not
|
|
|
What were three characteristics of DNApol1 that Arthur Kornberg stated that led to his suggestion of more than one DNA polymerase?
|

|
|
|
What does high and low processivity mean?
|
low processivity means that the polymerase frequently falls off and must rebind to the nucleic acid.
high processivity is a result of not losing this transition time of being off and on the nucleic acid |
|
|
DNA pol3 has how many subunits?
|
10 different types of subunits
|
|
|
DNApol3 is considered fast due to high ____________.
|
processivity
|
|
|
DNApol3 lacks a nuclease activity, name it.
|

It can proofread, but has no nick repair
|
|
|
_____________ is the principle replication DNA polymerase in E.coli.
|
DNApol3
|
|
|
Mutations in DNApol1 are __________, but mutations in DNApol3 are __________.
|
unlethal, lethal
|
|

|
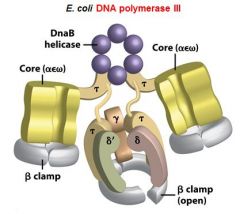
|
|
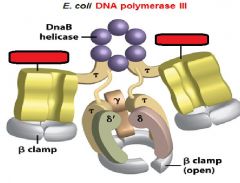
|
|
|

|
|
|
|
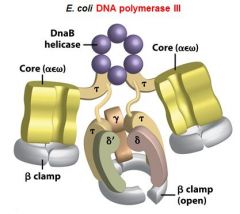
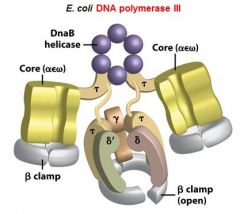
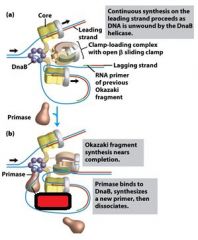
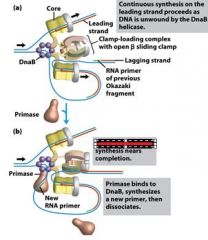
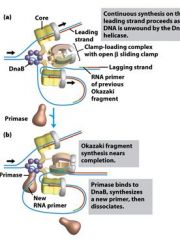
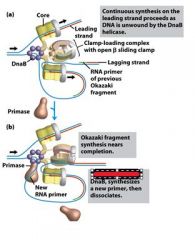
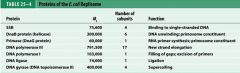
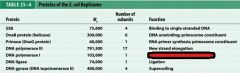
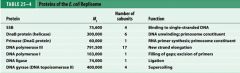
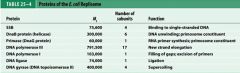
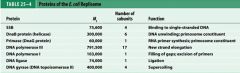
What is the functions of the core subunits of DNApol3?
|
polymerization of the nucleotides, 3'-->5' proofreading exonuclease, stabilization of the proofreading subunit
|
|
|
What is DnaB?
|
helicase for DNApol3
|
|
|
What is the function of the clamp loading complex of DNApol3?
|
Stable template binding, core enzyme dimerization, clamp loader, clamp opener
|
|
|
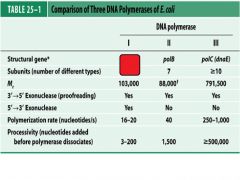
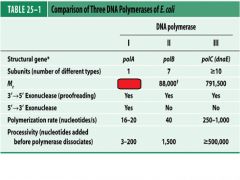
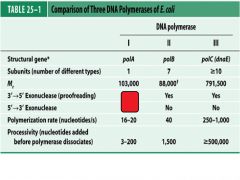
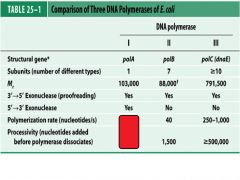
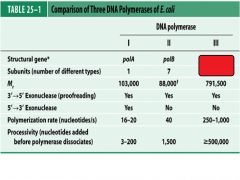
Compare and contrast DNApol1 with DNApol3.
|

|
|
|
What is a Klenow fragment?
|
The Klenow fragment is a large protein fragment produced when DNA polymerase I from E. coli is enzymatically cleaved by the protease subtilisin. First reported in 1970,[1] it retains the 5'-3' polymerase activity and the 3’ → 5’ exonuclease activity for removal of precoding nucleotides and proofreading, but loses its 5' → 3' exonuclease activity.
|
|
|

|
|
|

|
|
|

|
|
|

|
|
|

|
|

|

|
|

|

|
|

|

|
|
|
___________ experiments used denaturation mapping and concluded that for a given plasmid, the replication originated at _________________. He then call this ________________.
|
Inman, the same place every time, point of origin
|
|
|
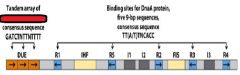
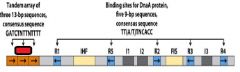
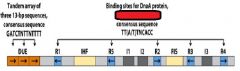
In E.coli the specific site of the origin is _______. This site is a ______bp sequence.
|
oriC, 245
|
|
|
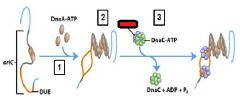
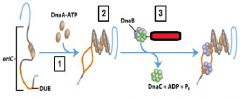
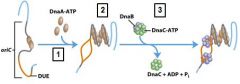
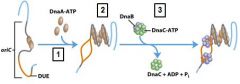
___________ binds to the oriC DNA sequence causing the DUE site to separate.
|
DnaA-ATP
|
|
|
DnaA-ATP bind to the oriC DNA sequence causing an adjacent 45 bp to separate. What is the site where the separation occurs called?
|
DUE DNA unwinding element
|
|
|
DnaA-ATP binding at the oriC site occurs only when both strand of ______ sequences are fully ___________________.
|
GATC, methylated at N6 of adenine.
|
|
|

|
|

|

|
|
|

|
|
|

|
|

|

|
|
|
DnaA-ATP binds to R- and I- sites in the origin and induces ___________________.
|
supercoiling
|
|
|
____________ binds to R- and I- sites in the origin and induces supercoiling.
|
DnaA-ATP
|
|
|
What happens as a result of DnaA-ATP inducing supercoiling in the oriC site?
|
The DUE (DNA unwinding element) separates to relieve the torsional stress.
|
|
|
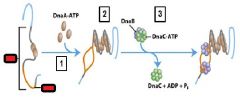
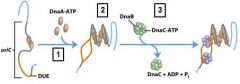
What enzyme binds to and opens the DnaB helicase and loads one enzyme onto each of the separated DUE strands?
|
DnaC-ATP
|
|
|
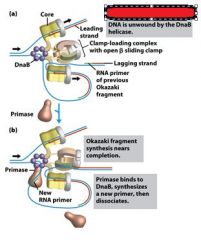
What enzyme generates an RNA primer on each separated strand at the DUE?
|
DnaG primase
|
|
|
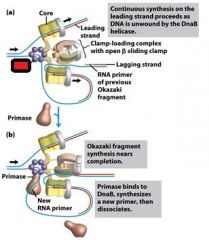
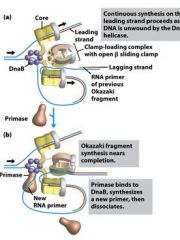
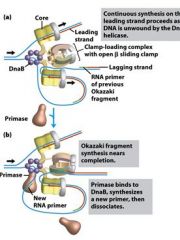
Which enzyme binds to the loaded DnaB helicases?
|
DNA polymeraseIII holoenzyme
|
|
|

|
|

|
|
|
|

|
|
|
|
|
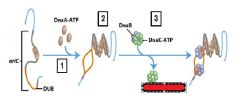
DnaA is active when bound to ______ and inactive when bound to _______.
|
ATP, ADP
|
|
|
The full methylation of the GATC sequences at oriC is called?
|
DAM methylation
|
|
|
Where on the GATC sequences of oriC does methylatio happen?
|
N6 of adenine
|
|
|
___________ is the only phase of DNA replication that is thought to be regulated.
|
Initiation
|
|
|
___________ levels are regulated throughout the cell cycle and peak around the start of replication
|
DnaA-ATP
|
|
|
DnaA-ATP levels are regulated throughout the cellc ycle and peak around the ___________________.
|
start of replication
|
|
|
Loading of the DNA polymerase3 complex stimulates hydrolysis of ___________ to ___________.
|
DnaA-ATP to DnaA-ADP
|
|
|
Hydrolysis of DnaA-ATP results in the ______________________. (long answer)
|
disassembly of the DnaA complex at the origin
|
|
|
What event stimulates the hydrolysis of DnaA-ATP?
|
the loading of the DNApol3 holoenzyme
|
|
|
The conversion of DnaA-ADP back to DnaA-ATP takes a lot of time resulting in?
|
a sufficient lag to allow replication to completeR
|
|
|
Replication creates new sites for ___________ binding that lie outside of the origin, effectively titrating away ____________.
|
DnaA-ATP
|
|
|
Conversion of the __________________ strands that result from replication back to fully methylated strand takes time.
|
hemimethylated
|
|

|

|
|
|
|
|

|
|
|
|

|
|

|
|
|

|

|
|
|
|
|
|

|
|
|
|
|
|
|
|
|

|
|
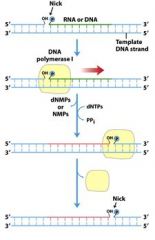
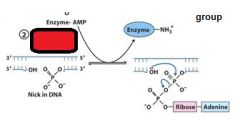
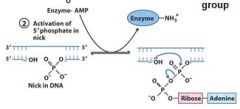
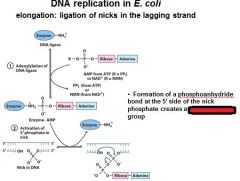
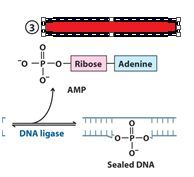
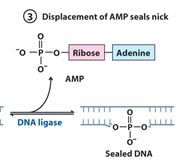
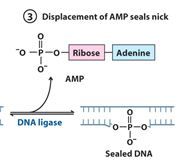
The left over nick is repaired by?
|
DNA ligase
|
|
|
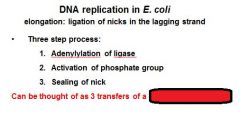
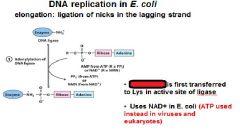
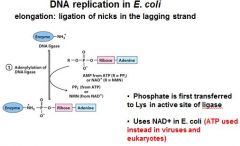
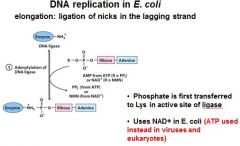
DNA ligase catalyzes formation of __________ bond by using _______. (only in E.coli)
|
phosphodiester, NAD+
|
|

|

|
|

|

|
|

|

|
|

|

|
|
|
|
|

|
|
|
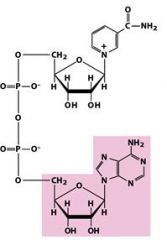
Name this molecule.
|
NAD+
|
|
|
|
|
|

|
|
|
|
|

|
|
|

|

|
|

|
|
|

|
|
|

|
|
|
|
|
|

|
|
|

|
|

