![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
541 Cards in this Set
- Front
- Back
|
Microbiology |
Study of microscopic organisms including fungi, protists and viruses |
|
|
What two themes does microbiology revolve around? |
1. Understanding basic life processes. 2. Applying that knowledge to benefit humans |
|
|
Why are microorganisms important? |
They are the oldest form of life, the largest mass of living material on earth, they carry out biogeochemical cycles, can live in Environments unsuitable for others and other life forms require microbes to survive. |
|
|
Cytoplasmic (cell) membrane |
Barrier that separates the inside of the cell from the outside Environment. |
|
|
Cytoplasm |
Aqueous mixture of macromolecules, ions and ribosome |
|
|
Ribosomes |
Protein synthesizing structures |
|
|
Cell wall |
Present in most microbes; confers structural strength |
|
|
What are the three domains? |
Eukarya, Bacteria and Archaea |
|
|
Archaea are more closely related to.. |
Eukarya |
|
|
How are prokaryotes different from eukaryotes? |
There organelles are not membrane enclosed, they have no nucleus, and they are generally smaller than most eukaryotes |
|
|
How are eukaryotes different from prokaryotes? |
They have their DNA enclosed in a membrane bound nucleus and their cells are generally larger and more complex. They contain organelles. |
|
|
Genome |
A cells full complement of genes; includes chromosomes, epigenome, and plasmids |
|
|
Prokaryotic cells dna |
Single, circular DNa (called a chromosome), DNA aggregates to form the nucleoid region, they have small amounts of extra groomsman DNA (plasmids) that confer special properties. |
|
|
Escherichia coli genome vs human cells |
4.63 m BPs, 4300 genes; 1000x more DNA per cell than E. Coli, 7 X more genes than E. Coli |
|
|
What are the six characteristics of living cells? |
Metabolism, reproduction, differentiation, communication, movement (some cells are known with no form of movement) and evolution |
|
|
Age of the earth |
4.6billion |
|
|
First cells appear |
3.8-3.9 bya |
|
|
Anoxic until |
~2 bya |
|
|
Oxygen producing phototrophs that made the earth oxic |
Cyanobacteria; blue-green algae |
|
|
Life was exclusively microbial until |
~1 bya |
|
|
Evolution |
The process of change over time that results in new varieties and species of organisms |
|
|
Phylogeny |
Evolutionary relationships between organisms, relationships can be deduced by comparing genetic info in different specimens. rRNA can help determine phylogeny |
|
|
The three distinct domains were determined by comparison of |
rRNA |
|
|
Microbial communities |
Populations of interacting assemblages. This is how microorganisms exist in nature. |
|
|
Ecosystem |
All living organisms plus physical and chemical constituents in their environment. |
|
|
Microbial ecology |
The study of microbes in their natural environment. |
|
|
Diversity and abundance of microbes are controlled by |
Resources (nutrients) and Environmental conditions (temp, pH, O2,etc) |
|
|
Extremeophiles |
Bacteria and archaea that grow in extremely harsh environments (very hot or cold, acidic or caustic, salty or osmotically stressful, high pressure) |
|
|
What two cycles are microbes involved in? |
Nitrogen and sulfur ki |
|
|
Where in humans can microbes be found? |
Stomach, small intestine, large intestine |
|
|
What (beneficial) function do microbes serve in humans? |
Vitamins and nutrients, and keeping pathogens at bay; mouth has the greatest diversity |
|
|
How can microorganisms be used in food? |
Preservation, spoilage and food-safety |
|
|
What are some forms of glucose breakdown that can be used in foods? |

|
|
|
Robert Hooke (1635-1703) |
The first to describe microbes; illustrated the fruiting structures of molds |
|
|
Antoni von Leewenhoek (1632-1723) |
The first to describe bacteria; further progress required development of more powerful microscopes. |
|
|
Ferdinand Cohn (1828-1898) |
Founded the field of bacterial classification and discovered bacterial endospores |
|
|
Louis Pasteur (1822-1895) |
Discovered that living organisms discriminate between optical isomers. Discovered that alcoholic fermentation was a biologically mediated process. Disproved theory of spontaneous generation; led to the development of methods for controlling the growth of microorganisms (aseptic technique). Developed vaccines for anthrax, fowl cholera, and rabies |
|
|
Theory of spontaneous generation |
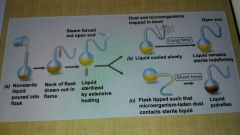
|
|
|
Robert Koch (1843-1910) |
Demonstrated the link between microbes and infectious diseases, identified causative agents of anthrax and tuberculosis. Koch's postulates. Developed techniques for obtaining pure cultures of microbes. Awarded Nobel piece prize for physiology and medicine in 1905. |
|
|
Koch's postulates |
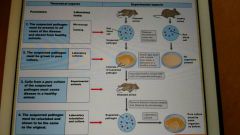
|
|
|
Martinis Beijerinck (1851-1931) |
Developed enrichment culture technique. |
|
|
Enrichment culture technique |
Microbes can be isolated from natural samples in a highly selective fashion by manipulating nutrient and incubation conditions. |
|
|
Sergei Winogradsky (1856-1953) |
Demonstrated that specific bacteria are linked to specific biogeochemical transformations. Proposed concept of chemolithotrophy |
|
|
Chemolithotrophy |
Oxidation of inorganic compounds linked to energy conservation. |
|
|
Genomics |
Study of all of the genetic material in living cells |
|
|
Transcriptomics |
Study of RNA patterns |
|
|
Proteomics |
Study of all the protiens produced by a cell |
|
|
Metabolically |
Study of metabolic expression in cells |
|
|
Coccus |

Spherical or ovoid or nearly spherical |
|
|
Rod |

Cylindrical shape; vaccilus or vacili |
|
|
Spirillum |
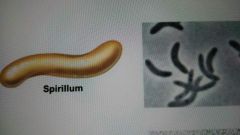
Spiral shape |
|
|
Coccobaccili |
Short rod shaped |
|
|
Vibrio |
Comma or crescent shape |
|
|
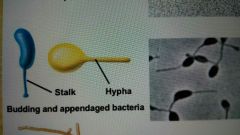
What are cells with unusual shapes |
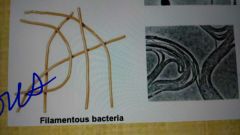
Spirochetes, appendaged bacteria, and filamentous bacteria |
|
|
Spirochete |
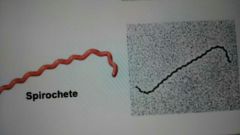
|
|
|
Budding and appendaged bacteria |
|
|
|
Filamentous bacteria |
|
|
|
What does bacterias small size and large surface area allow for? |
Fast replication, support a greater nutrient exchange per unit cell volume and they tend to grow faster than larger cells |
|
|
Cytoplasmic membrane |
Structure that surrounds the cell, vital barrier that separates cytoplasm from environment, highly selective permeable barrier which enables concentration of specific metabolites and excretion of waste products. |
|
|
Mycelles |
Sphere with 1 layer or bilayer membrane. |
|
|
What are the parts of membranes? |
1. Hydrophobic region (includes the fatty acids) 2. Hydrophilic region (includes a phosphate with a functional group) |
|
|
Cytoplasmic membrane |
Semifluid 8-10nm membrane with embedded proteins, that is stabilized with hydrogen bonds and hydrophobic interactions. They use Mg++ and CA++ to help stabilize by forming ionic bonds with neg charges on the phospholipids. |
|
|
What are the two types of membrane proteins? |
Integral and peripheral |
|
|
Integral membrane proteins |
Firmly embedded in the membrane |
|
|
Peripheral membrane proteins |
One portion anchored in the membrane |
|
|
What types of linkages do eukarya and bacteria have linking phospholipids? |
Ester |
|
|
What linkages do archaea have in phospholipids? |
Ether |
|
|
Archaea membranes |
Lack fatty acids, have isoprenes instead. Major lipids are glycerol dieters and tetraethyl. Can exist as lipid monolayers, bilayers or both |
|
|
Permeability barrier |
Polar and charged molecules must be transported which requires energy. Transport proteins accumulate solutes against the concentration gradient. |
|
|
Protein anchor |
Holds transport proteins in place |
|
|
Energy conservation |
Proton motive force |
|
|
Functions of the membrane |
Permeability barrier, protein anchor and energy conservation |
|
|
What are the three major classes of transport systems in prokaryotes? |
Simple transport, group translocation, and the ABC system *All require energy in some form, usually proton motive force |
|
|
Simple transport |
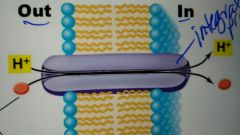

Driven by the energy in the proton motive Force |
|
|
Group translocation |

Chemical modification of the transported substance driven by phosphoenolpyruvate |
|
|
ABC transporter |
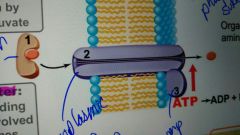
Periplasmic binding proteins are involved in energy comes from ATP; Texans can be delivered this way |
|
|
Peptidoglycan |
In bacteria that have cell walls, this is the main cell wall material |
|
|
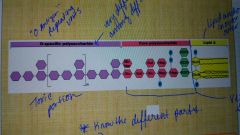
LPS |
Lipidpolysaccharides; they are present in all gram-negatives |
|
|
Gram-negative cell |
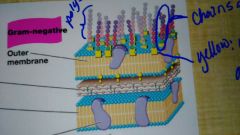
They have one thin cell wall, in two phospholipid bilayers: the cytoplasmic and outer membrane. |
|
|
Gram positive cell |
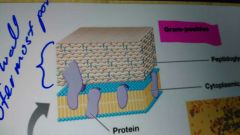
They have one thing cell wall which is about five to 10 times thicker than in a gram-negative cell and they have one phospholipid bilayer, the cytoplasmic membrane |
|
|
When is LPS released? |
Anytime the cell dies which can cause septic shock and lower the blood pressure. |
|
|
Peptidoglycan |
A rigid layer that provides strength to a cell wall. It has a polysaccharide composed of n-acetylglucosamine and n-acetylmuremic acid as well as amino acids and lysine or diaminopimelic acid (DAP). The polysaccharide is Crosslink differently in gram-negative bacteria and gram-positive bacteria |
|
|
What type of amino acids are common in nature? |
L; D forms may be used to stop enzymes from breaking them down but they're still are enzymes like lysozyme that have evolved to be able to break down the D form. |
|
|
Gram positive cell walls |
They can contain up to 90% peptidoglycan, And it is common for them to have teichoic acids embedded in their cell wall including lipoteichoic acids which are teichoic acids that are covalently bound to membrane lipids |
|
|
Give two examples of prokaryotes that lack cell walls |
Mycoplasmas ( a group of pathogenic bacteria including atypical pneumonia and walking pneumonia) and thermalplasma ( species of archaea) |
|
|
LPS |
Present in gram-negative pictorial only. It consists of core polysaccharides and O polysaccharides. It replaces most of the phospholipids in the outer half of the outer membrane and it is an endotoxin. |
|
|
EPS |
Extracellular polysaccharide. It can be present as a slime layer to form a capsule or slime around the cell outside of the LPS |
|

Label the different parts |
|
|
|
Periplasm |
The space located between the cytoplasmic in the outer membranes which is about 50 nanometers wide. The contents have a gel-like consistency and houses many proteins where initial digestion occurs. |
|
|
Porins |
Channels for movement of hydrophilic low-molecular-weight substances which of the involved in facilitated diffusion |
|
|
Archaeal cell walls |
They have no peptidoglycan in typically no outer membrane. Some contain pseudomurein. |
|
|
Pseudomurein |
Polysaccharide similar to peptidoglycan which is composed of N- acetylglucosamine and n-acetylalosaminuonic acid. it is found in the cell walls of certain methanogenic archaea. |
|
|
S-layers |
Most common cell wall type of archaea which consist of protein or glycoprotein and has a paracrystalline structure. |
|
|
Fimbriae/pili |
Short, rigid protein structures that are hairlike and involved in attachment to which can be seen around the outside of a cell |
|
|
Inclusions |
Granular or crystalline structures often involved in energy storage |
|
|
Gas vesicles |
Membrane-bound which can be proteinaceous and can hold gas or very low density proteins and also can be used for buoyancy in aquatic environments which allows the bacteria to dive deeper to block some of the UV light. |
|
|
Endospores |
when nutrients are scarce bacteria sacrifice themselves and form a small hard spore and when conditions are right they germinate and Form 1 cell |
|
|
Flagella |
Long tail like structures about two to three times the length of the cell made up of proteins called flagellins. They are attached to a motor which spans the cytoplasmic membrane cell wall and expands into the cytoplasm. Energy from ATP from the motive Force Powers it |
|
|
What is occurring when bacteria are mating? |
They are transferring genetic material that is not in the chromosome but in a plasmid the donor is f+ and the recipient is f minus |
|
|
Describe the shape and different arrangements of the flagella |
They are helical in shape and can be peritrichous (flagella all over), polar (flagella only at the ends) or lophotrichous (tuft of flagella at one or both ends) |
|
|
Describe the movement of flagella |
Move in runs (straight) and tumbles (turn by rotating flagella opposite direction) |
|
|
Flagellar structure of archaea |
They're half the diameter of bacterial flagella and composed of several different proteins and move by rotation |
|
|
Chemotaxis |
Movement in response to a chemical stimulus usually toward food or away from a chemical toxin |
|
|
Mot protein |
Involved in PH difference and obtains energy from the proton motive Force it works similarly to an electric motor and powered by the positive and negative charges. |
|
|
Describe the swimming motion of peritrichous flagella |
They move slowly and a straight line |
|
|
Describe the swimming and motility of Polarly flagellated cells |
They move more rapidly and typically spin around |
|
|
Gliding motility |
This is for flagella-independant motility which is slower and smoother than swimming. Movement typically occurs along the long axis of a cell and require surface contact the mechanisms used are by the excretion of polysaccharide slime, Type 4 pili or gliding specific proteins. |
|
|
What are the different types of Taxis |
Chemotaxis, phototaxis, Aerotaxis, osmotaxis and Hydrotaxis |
|
|
What is the function of smooth ER |
Lipid production |
|
|
In prokaryote what is the order of transcription and translation? |
They occur simultaneously |
|
|
Nutrients |
Supply of monomers or precursors required by cells for growth |
|
|
Macronutrients |
Nutrients required in large amounts including CNNaPCaFeMg |
|
|
Micronutrients |
Nutrients required in Trace Amounts |
|
|
Growth factors |
Organic compounds required in small amounts by certain organisms such as vitamins amino acids, purines, andpyrimidines |
|
|
Vitamins |
Most commonly required growth factors most function as coenzymes |
|
|
Culture media |
Nutrient Solutions used to grow microbes in the laboratory |
|
|
What are the two broad classes of culture media |
Defined media which has a precise chemical composition and complex media which is composed of digestive chemically undefined substances |
|
|
Enriched media |
Contain complex sometimes defined media plus additional nutrients which enrich for certain types of organisms over others |
|
|
Differential media |
Contain an indicator usually die that detects particular chemical reactions occurring during growth and allows you to differentiate between different microbes |
|
|
Pure culture |
Culture containing only a single kind of microbe |
|
|
Contaminants |
Unwanted organisms in a culture |
|
|
Solid culture media |
Prepared by addition of a gelling agent such as agar or gellatin and when grown and solid media cells form isolated misses called colonies |
|
|
Metabolism |
The sum total of all the chemical reactions that occur in a cell or associated with a cell |
|
|
Catabolism |
Breaking down chemicals |
|
|
Anabolism |
Building a chemical or molecule |
|
|
What energy classes are microorganisms grouped into |
Chemo organotrophs chemolithotrophs phototrophs heterotrophs and autotrophs |
|
|
Energy |
Measured in units of kilojoules which is a measure of heat energy |
|
|
Free energy (G) |
Energy released that is available to do work |
|
|
Change in free energy during reaction is referred to as |

|
|
|
Activation energy |
Energy required to bring all the molecules in a chemical reaction into the reactive state. catalysis is usually required to bridge activation energy barrier |
|
|
Catalyst |
Lowers the activation energy of a reaction increases the reaction rate and does not affect energetics or equilibrium of reaction |
|
|
Enzymes |
They are biological catalyst which are typically proteins (some RNAs), they are highly specific and generally larger than the substrate or product. They typically rely and weak bonds such as hydrogen bonds Van Der waals forces or hydrophobic interactions. They contain an active site and they increase the rate of a chemical reaction. |
|
|
Prosthetic group |
Bind tightly to enzymes usually bind covalently and permanently or at least her tightly associated |
|
|
Coenzymes |
Loosely bound to enzymes most are derivatives of vitamins |
|
|
Co-factors |
They're even smaller than a coenzyme and almost always and inorganic ions such as sodium magnesium calcium. Etc |
|
|
Redox reactions |
Energy from the misuse didn't synthesis of energy rich compounds such as ATP. They occur in pairs of two half reactions. There is an electron donor and an electron acceptor |
|
|
Carriers |
Intermediates usually involved in redox reactions |
|
|
What are the two classes that are electron carriers are divided into |
Prosthetic groups and coenzymes |
|
|
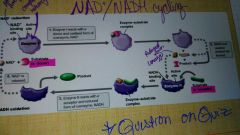
Give an example of a coenzyme |
NAD Plus or nadh |
|
|
NAD+/NADH cycling |
|
|
|
Where is the chemical energy that is released in redox reactions primarily stored? |
In certain phosphorylated compounds such as ATP, phosphoenolpyruvate, glucose-6-phosphate and it can also be stored in coenzyme a |
|
|
How can energy be stored long-term |
An insoluble polymers that can be acted as to generate ATP. In prokaryotes this includes glycogen, poly- beta -hydroxybutyrate and other polyhydroxyalkanoates and elemental sulfur. In eukaryotes this is in the form of starch or lipids |
|
|
What are the two reaction series link to energy conservation in chemoorganotrophs with glycolysis? |
Fermentation and respiration |
|
|
Fermentation |
Substrate level phosphorylation. ATP is directly synthesized from an energy-rich intermediate. Recycles NAD plus produces no ATP |
|
|
Respiration |
Oxidative phosphorylation. ATP is produced from proton motive force from by transport of electrons |
|
|
Glycolysis |
Glucose is consumed it is an anaerobic process where two atp's are produced. Fermentation products are generated and some can be harnessed by humans for consumption. |
|
|
Saccharomyces cerevisiae |
Carry out fermentation or respiration. It uses the one that's most beneficial. Respiration generates ATP fermentation occurs when conditions are anoxic |
|
|
Aerobic respiration |
Oxidation using O2 is the terminal electron acceptor. It's a higher ATP yield then fermentations where ATP is produced at the expense of the proton motive Force which is generated by electron transport |
|
|
Electron transport systems |
It is membrane associated and it mediates the transfer of electrons. It can serve some of the energy released during transfer and used it to synthesize ATP. Many oxidation reduction enzymes are involved in electron transport such as nadh dehydrogenase is Flavorproteins, iron sulfur proteins and cytochromes |
|
|
NADH dehydrogenases |
Proteins found to the inside service of cytoplasmic membrane. Active site binds nadh and accept two electrons into protons that are passed to flavoproteins. |
|
|
Flavoproteins |
Contains Flavin prosthetic group that accept two electrons and two protons but donates electrons only to the next protein and the chain |
|
|
Cytochromes |
Proteins that contain heme prosthetic groups. They accept and donate a single electron via the iron atom in heme |
|
|
Iron-sulfur proteins |
Contain clusters of iron and sulfur. Reduction potentials vary depending on the number and position of iron and sulfur atoms |
|
|
Quinones |
Hydrophobic non protein containing molecules that participate in electron transport. They accept electrons and protons but pass along electrons only |
|
|
The proton motive Force |
The electron transport system is oriented in the cytoplasmic membrane allowing it to separate electrons from protons. Electron carriers are arranged in the membrane and Order of the reduction potential and the final carrier in the chain donates electrons and protons to the terminal acceptor. |
|
|
Where do protons in the electron transfer chain originate from |
Nadh in the dissociation of water |
|
|
What is the electrochemical potential across the membrane called |
The proton motive Force |
|
|
The inside of the three membrane becomes negative and |
Alkaline or basic. The outside is acidic |
|
|
ATP synthase |
Complex that converts proton motive Force into ATP using two components: F1( multi protein extra membrane complex which faces the cytoplasm) and F0 ( proton conducting intramembranous channel). This is reversible and dissipates the proton motive Force |
|
|
Citric acid cycle or Krebs cycle |
Pathway through which pyruvate is completely oxidized to carbon dioxide. The initial steps include glucose to pyruvate the same as glycolysis. Per glucose molecule six carbon dioxide molecules are released and nadh and fadh2 generated this plays a key role in catabolism and biosynthesis. There's an energetix advantage to aerobic respiration |
|
|
How many atp's can potentially be made from glycolysis + the Krebs cycle? |
38 |
|
|
What is the 3 carbon compound that is a product of the Krebs cycle? |
Pyruvate or pyruvic acid |
|
|
What is the input for the Krebs cycle? |
Acetyl CoA |
|
|
alpha-Ketoglutarate and oxaloacetate (OAA) |
Compounds generated from the CAC; precursors of A.A.s; OAA is also converted to phosphoenolpyruvate, a precursor of glucose. |
|
|
Succinyl-CoA |
Generated from the CAC; required for synthesis of cytochromes, chlorophyll, and other tetrapyrrole compounds. |
|
|
Acetyl CoA |
Generated from the CAC; necessary for fatty acid biosynthesis |
|
|
What are some of the mechanisms microbes use to generate energy? |
Fermentation, aerobic respiration, anaerobic respiration, chemolithotrophy, and phototrophy. |
|
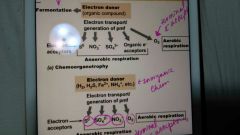
What types of organisms are use these processes? |
Chemolithotrophs |
|

What types of organisms are use these processes? |
Chemolithotrophs |
|
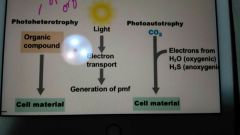
What types of organisms use these processes? |
Phototrophs |
|
|
In fermentation what is the electron donor? |
An organic compound |
|
|
What can beelectron acceptors in fermentation? |
S°, NO3-, SO4 2-, organic electron acceptors and O2 |
|
|
What is the terminal electron acceptor in fermentation? |
O2 |
|
|
In chemoorganotrophy what can be an electron donor? |
H2, H2S, Fe3+, NH4+, etc. |
|
|
In chemoorganotrophy, what can be electron acceptors? |
S°, SO4 2-, NO3-, O2 |
|
|
What is the terminal electron acceptor in chemoorganotrophy? |
O2 |
|
|
What type of compounds are used for a carbon source in photoheterotrophy? |
Organic compounds |
|
|
What is used for a carbon source in photoautotrophy? |
CO2 |
|
|
What are prokaryotic polysaccharides synthesized from? |
Activated glucose |
|
|
Adenosine diphosphoglucose (ADPG) |
Precursor for glycogen biosynthesis |
|
|
Uridine diphosphoglucose (UDPG) |
Precursor of some glucose derivatives needed for biosynthesis of important polysaccharides; examples include N-acetylglucosamine, and N-acetylmuramic acid |
|
|
Gluconeogenesis |
Synthesis of glucose from phosphoenolpyruvate |
|
|
Phosphoenolpyruvate can be synthesized from what? |
Oxaloacetate |
|
|
What is the path from intermediates of the citric acid cycle to the first intermediate in glycolysis? |
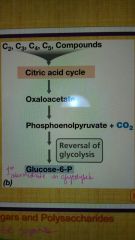
|
|
|
What is required for the synthesis of nucleic acids? |
Pentoses (5 carbon sugars) |
|
|
If pentoses are unavailable in the environment how can organisms synthesize them? |
By the removal of one carbon from a hexose |
|
|
What is the path of using glucose to make DNA and RNA? |

|
|
|
How are carbon skeletons made during biosynthesis of amino acids and nucleotides? |
They come from intermediates of glycolysis or Krebs cycle. |
|
|
How is ammonia used during biosynthesis of AAs and nucleotides? |
It's Incorporated by glutamine dehydrogenase or glutamine synthetase. |
|
|
During biosynthesis of AAs and nucleotides what is the amino group transferred by? |
Transaminase and synthase |
|
|
During biosynthesis of AAs and nucleotides what is the amino group transferred by? |
Transaminase and synthase |
|
|
Oxitroph |
Deficient in 1 or more amino acids; used instead lab setting to manipulate bacteria. |
|
|
Haynes test |
Organisms lacking histodines are grown on media without histodines. If there are no mutagens present the BP can be changed so they no longer lack histodine. |
|
|
How many carbons at a time are fatty acids biosynthesized? |
2 |
|
|
How is Acyl carrier protein (ACP) involved in fatty acid biosynthesis? |
It holds the growing fatty acid as it's being synthesized |
|
|
What is involved in the final assembly of lipids in bacteria and eukarya? |
The addition of fatty acids to glycerol |
|
|
Instead of fatty acids Archaea have lipids that contain what? |
Phytanyl side chains |
|
|
How can nitrogen fixing bacteria be found? |
Free living or symbiotic |
|
|
What is the nitrogen fixing reaction catalyzed by? |
Nitrogenase |
|
|
Nitrogenase is sensitive to the presence of what? |
Oxygen (only functional without it) |
|
|
Rhizobium |
Form nodules on the roots of legumes; symbiots with plants; convert N2 to ammonia |
|
|
Where does nitrogen fixation occur? |
In membrane bound organelles devoid of oxygen |
|
|
Where does nitrogen fixation occur? |
In membrane bound organelles devoid of oxygen |
|
|
Electron flow in nitrogen fixation |

|
|
|
What is involved in a redox reaction during nitrgen fixation? |
Dinitrogenase reductase/flavodioxin |
|
|
In nitrogen fixation for every N2 how many NH3- are made? |
2 |
|
|
What is one way to assay nitrogenase activity? |
Acetylene reduction assay |
|
|
What occurs in a acetylene reduction assay? |
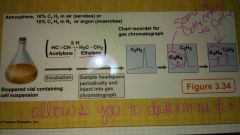
|
|
|
Nitrogen |
Typical bacterial cell is ≈13% nitrogen by dry weight; key element in proteins, nucleic acids, and many more cell constituents |
|
|
Phosphorus |
Synthesis of nucleic acids and phospolipids |
|
|
Sulphur |
Sulphur -chain amino acids (cysteine and methionine); vitamins and coenzyme-A |
|
|
Potassium |
Required by enzymes for activity |
|
|
Potassium |
Required by enzymes for activity |
|
|
Magnesium |
Stabilizes ribosomes, membranes, and nucleic acids; also required for many enzymes |
|
|
Calcium |
Helps stabilize cell walls in microbes; play key role in heat stability of endospores |
|
|
Sodium |
Required by some microbes |
|
|
Functional unit of the generic information |
Gene |
|
|
What are genes composed of? |
DNA |
|
|
What are the three informational macromolecules in a cell? |
DNA, RNA, and protein |
|
|
Three stages of genetic information flow? |
Replication, transcription and translation |
|
|
Replication |
DNA is duplicated |
|
|
Transcription |
Info from DNA is transferred to RNA |
|
|
Transcription |
Info from DNA is transferred to RNA |
|
|
mRNA |
Encodes polypeptides |
|
|
tRNA |
Plays role in protein synthesis |
|
|
rRNA |
Plays role in protein synthesis |
|
|
rRNA |
Plays role in protein synthesis |
|
|
Translation |
Info from RNA is used to build polypeptides |
|
|
Ribosomes |
Protein polymerases |
|
|
Why is DNA in a double helix |
More stable |
|
|
Central dogma |
DNA to RNA to protein |
|
|
Transcription in Eukaryotes |
Each gene is done individually; 1 gene/1 control region(transcriptional unit) which allows for more control but less efficiency. |
|
|
Transcription in prokaryotes |
Multiple genes may be done together in operons; multiple genes/1 control region(transcriptional unit) which allows efficiency and less energy; also don't have the luxury of space for more base pairs |
|
|
Operon |

|
|
|
What are the four nucleotides found in DNA? |
Adenine, Guanine, cytosine and thymine |
|
|
Describe the backbone of DNA |
It consists of alternating phosphates and the pentoses sugar, deoxyribose; phosphates connect 3'-carbon of one sugar to 5'-carbon of the adjacent sugar |
|
|
What bases are pyridimines? |
C,T and U |
|
|
What bases are purine bases? |
A and G |
|
|
What does it mean for DNA to have antiparallel strands? |
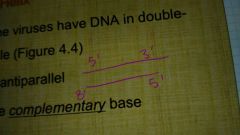
|
|
|
What are the complimentary base sequences in DNa? |
A and T, C and G |
|
|
How does the 3' and 5' ends get their name? |
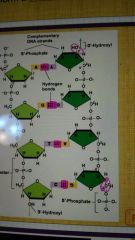
|
|
|
How does the 3' and 5' ends get their name? |

|
|
|
1Mbp |
Mega base pair; 1 million |
|
|
How big is the E. coli genome? |
4.64Mbp |
|
|
How long is each BP? |
.34nm |
|
|
How many BP make up one turn of the helix? |
10 |
|
|
Supercoiled DNA |
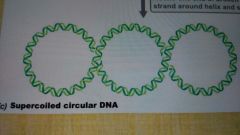
DNA is further twisted to save space |
|
|
Negative supercoiling |
Double helix is underwound |
|
|
Positive supercoiling |
Double helix is overwound |
|
|
Relaxed DNA |

DNA has number of turns predicted by number of bp |
|
|
DNA gyrase |
Introduces supercoils into DNA |
|
|
What are the benefits to DNA being wound? |
Protected and compact |
|
|
When is DNA unwound? |
Only when it needs to be read |
|
|
Genome |
Entire complement of genes in cell or virus |
|
|
Genome |
Entire complement of genes in cell or virus |
|
|
Chromosome |
Main genetic element in prokaryotes (other elements include virus genomes, plasmids, organelles genomes, and transposable elements) |
|
|
Extrachromosomal DNA |
DNA outside the chromosome; includes in plasmids, mitochondria and chloroplasts |
|
|
Describe virus genomes |
They contain either DNA or RNA; can be linear or circular; can be single or double stranded |
|
|
Plasmids |
Replicate separately from chromosomes, thought they may be in sync. The great majority are double stranded. Most are circular and they generally contain genes that are beneficial for the cell. Since it's costly to replicate the plasmids are only kept if they are needed. The smaller they are, the higher the copy number. |
|
|
Chromosome |
A genetic element with "housekeeping genes" (presence of essential genes is necessary to be considered a chromosome) |
|
|
What main way is a plasmid different from a chromosomal gene? |
It is expendable and rarely contains genes for growth under all conditions. |
|
|
Transposable elements |
Segment of DNA that can move from one site to another on the same or a different DNA molecule. |
|
|
What are the three main types of transposable elements? |
1. Insertion sequences 2. Transposons 3. Special Viruses |
|
|
Insertion sequences |
Usually smaller, found at the ends of most transposons |
|
|
Special viruses |
Some are associated with cancer because when they insert in the chromosome they can hit an essential gene. |
|
|
Random mutagenosis |
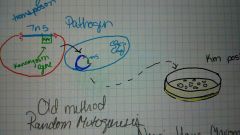
Old method |
|
|
Targeted mutagenosis |
Have the chromosomal DNA sequence. So construct a vector and swap out a targeted gene |
|
|
The Escherichia coli chromosome |
Because E. Coli is a useful model organism (biochemistry, genetics, and bacterial physiology) the genome (from the strain MG1655) has been mapped out using conjugation, transduction, molecular cloning, and sequencing |
|
|
How many minutes is the chromosome of E. Coli usually cut up into? |
100 |
|
|
Conjugation |
Bacterial mating with sex/fertilization pilus; sometimes entire chromosomes can be transferred; process takes 100min. |
|
|
Features of the E. Coli chromosome |
Many genes encoding enzymes of a single biochemical pathway are clustered into operons. Operons are equally distributed on both strands (~5 MBP in size), about 40% of predicted proteins are of unknown function. The Ave protein contains about 300 AAs. Has insertion sequences. |
|
|
Describe plasmids |
Small circular or linear DNA, size: 1kbp to >1Mbp (typically less than 5% size of the chromosome). Carry non-essential but helpful genes. Abundance)copy number is variable. |
|
|
F plasmids |
Conjugated from cell to cell |
|
|
F plasmids |
Conjugated from cell to cell |
|
|
R plasmids |
Resistance plasmids; confer resistance to antibiotics and other growth inhibitors. Widespread and well-studied group of plasmids. Many are conjugative. |
|
|
tra genes |
Transfer; these genes are involved with the transfer of the plasmid |
|
|
Virulence factors |
In several pathogenic bacteria they are encoded by plasmid genes (on chromosome of those that require virulence for survival); enable pathogene to: colonize , cause additional host damage. Can contain hemolysin and/or enterotoxin |
|
|
Virulence |
A range which is determined by a virulence gene. |
|
|
Pathogenicity |
All or nothing (either is pathogenic or not) |
|
|
Hemolysin |
Lysing of cells, especially blood but also others |
|
|
Enterotoxin |
Often enzymes, effects the gut |
|
|
Bacteriocins |
Proteins produced by bacteria that inhibit or kill closely related species or even different strains of the same species; Colicin, nisin (sometimes used as anitmicrobials); genes are often carried on plasmids |
|
|
Semiconservative tendency of DNA replication |
Each of the two progeny double helices have one parental strand and one new strand. |
|
|
What is the precursor to each nucleotide? |
A deoxynucleoside 5'-triphosphate (dNTP) |
|
|
How does replication proceed? |
From the 5' end to the 3' end |
|
|
DNA polymerases catalyze what? |
the addition of dNTPs |
|
|
How many different DNA polymerases are there in E. Coli? |
5 |
|
|
What is the primary enzyme replicating chromosomal DNA in E. Coli? |
DNA polymerase III |
|
|
What do DNA Polymerases require to attach? |
A primer |
|
|
What are primers made from? |
RNA by primase |
|
|
Polymerase |
Anything that takes small monomers and turns them into polymers |
|
|
Where does DNA synthesis begin? |
At the origin of replication in prokaryotes |
|
|
Replication fork |
Zone of unwound DNA where replication occurs; bidirectional |
|
|
How is the leading strand synthesized |
Continuously |
|
|
How is the lagging strand synthesized? |
In fragments (okazaki) |
|
|
DNA helicase |
Unwinds the DNA |
|
|
DNA polymerase I |
Removes the RNA primer and replaced it with DNA |
|
|
DNA ligase |
Seals Nick's in the DNA |
|
|
DNA ligase |
Seals Nick's in the DNA |
|
|
DNA synthesis in prokaryotes |
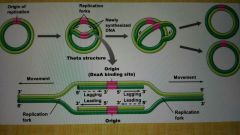
Bidirectional in the chromosome; two replication forks moving in opposite directions |
|
|
Hiw many BP can DNA Polymerase III add per second? |
1000 / 1 kbp |
|
|
Replisome |
Complex of multiple proteins involved in replication; area of synthesis; DNA is pulled through the replisome |
|
|
What helps DNA replication to be extremely accurate? |
Proofreading |
|
|
Mutation rates during replication |
10^-8 – 10^-11 errors per base inserted |
|
|
What can detect mismatch through incorrect hydrogen bonding during DNA synthesis? |
Polymerase |
|
|
What types of organisms have a proofreading mechanism? |
Prokaryotes, eukaryotes and viral DNA replication systems |
|
|
How can you obtain a pure culture easily? |
Using Koch's method of streaking |
|
|
What is almost always involved in metabolic reactions? |
Enzymes |
|
|
Selective medium |
Inhibits some types of organisms while promoting others |
|
|
Selective medium |
Inhibits some types of organisms while promoting others |
|
|
Who was considered the father of infectious disease? |
Koch |
|
|
What type of microscopy are we using in the lab? |
Bright field |
|
|
What is the most simple type of organism? |
Bacteria |
|
|
What type of translocation involves a chemical modification of the delivered material? |
Group translocation |
|
|
What type of bacteria has teichoic acids? |
Gram positive; they are embedded in the cell wall |
|
|
Does aseptic technique include turning the plates upside down? |
No |
|
|
Enriched medium |
Favors the growth of one over all others |
|
|
Who disproved spontaneous generation? |
Louis Pasteur |
|
|
Transcription (DNA to RNA) is carried out by what enzyme? |
RNA polymerase |
|
|
In transcription RNA polymerase uses what as a template? |
DNA |
|
|
In transcription what are the RNA precursors? |
ATP, GTP, CTP, and UTP |
|
|
In transcription how does chain growth take place? |
5'-3' |
|
|
For any gene what is transcribed by RNA polymerase? |
Only one of two strands of DNA |
|
|
How many different subunits does RNA polymerase have? |
Five |
|
|
Promotors |
Sites on DNA recognized by RNA Polymerase |
|
|
Promotors |
Sites on DNA recognized by RNA Polymerase; site of initiation of transcription, recognized by sigma factor of RNA polymerase |
|
|
Transcription terminators |
Specific sites where Transcription stops |
|
|
Transcription terminators |
Specific sites where Transcription stops |
|
|
Why does transcription involve smaller units of DNA (as small as a single gene) unlike with DNA replication? |
It allows the cell to transcribe different genes at different rates |
|
|
How is sigma factor used in transcription? |
It works with DNA polymerase and recognizes two highly conserved regions of the promotor and initiation site. When transcription begins it is released to be used for another RNA polymerase. |
|
|
Simultaneous Transcription-Translation |
Only occurs in prokaryotes |
|
|
Simultaneous Transcription-Translation |
Only occurs in prokaryotes |
|
|
What are the two highly conserved regions of the promotors? |
Pribnow box and -35 region |
|
|
Pribnow box |
Located ten bases before the start of transcription (-10 region); TATA box |
|
|
-35 region |
Located ~35 bases upstream from transcription |
|
|
What is termination of RNA synthesis governed by? |
A specific RNA sequence |
|
|
Intrinsic terminators |
Transcription is terminated without any additional factors |
|
|
Rho-dependent termination |
Rho protein recognizes specific DNA sequences and causes a pause in the RNA polymerase |
|
|
Unit of transcription |
Unit of chromosome bounded by sites where transcription of DNA to RNA is initiated and terminated |
|
|
What are some types of RNA that are not translated and are very stable? |
rRNA, tRNA, etc. |
|
|
What are the three types of rRNA? |
16s, 23s, and 5s |
|
|
What is tRNA cotranscribed with? |
rRNA or other tRNA |
|
|
What type of RNAs have short half lives of a few minutes? |
mRNAs |
|
|
Polycistronic mRNA |
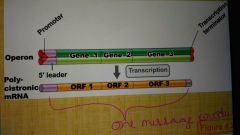
A mRNA encoding group of cotranscribed genes |
|
|
Operon |
A group of related genes cotranscribed on a polycistronic mRNA which allows for expression of multiple genes to be coordinated |
|
|
Describe RNA polymerase in archaea. |
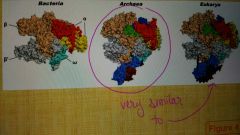
There is only one, which resembles eukaryotic polymerase II. |
|
|
Transcription in Archaea |
It is a simplified version of the eukaryotic transcription apparatus including promoters and RNA polymerase similar to eukaryotes. However, the regulation of transcription has major similarities with Bacteria. |
|
|
Exons |
Coding sequences in Eukaryotic genes |
|
|
Introns |
The intervening sequences in Eukaryotic genes; they are rare in Archaea but found in their tRNA and rRNA genes |
|
|
What are archaeal introns excised by? |
Special endonucleases |
|
|
Eukaryotic RNA processing |
Many RNA molecules are altered before they carry out their role in the cell. |
|
|
Eukaryotic RNA processing |
Many RNA molecules are altered before they carry out their role in the cell. Includes RNA splicing, RNA capping, and a Poly(A) tail. |
|
|
RNA splicing |
Takes place in the nucleus; removes introns from RNA transcripts; performed by the spliceosome |
|
|
Spliceosome |
Bends transcripts so RNA can be spliced out |
|
|
RNA capping |
Addition of methylated Guanine to 5' end of mRNA |
|
|
Poly(A) tail |
Addition of 100-200 adenylate residues; stabilizes mRNA and is required for translation; can isolate mRNA from other RNA in the cell. |
|
|
Proteins |
Play a major role in cell function |
|
|
Proteins |
Play a major role in cell function; polymers of AAs; they are linked by peptide bonds form a polypeptide |
|
|
What are some different types of proteins? |
Catalytic (enzymes), structural and regulatory |
|
|
Primary structure |
linear array of AAs in a polypeptide |
|
|
What are the chemical properties of the amino acid related to? |
Their side chain |
|
|
Secondary structure |
Adjacent nucleotides or nearly adjacent; interactions of the R groups force the molecule to twist and fold in a certain way. |
|
|
Tertiary structure |
3-D shape of polypeptide |
|
|
Quaternary structure |
Number and types of polypeptides that make a protein; >1 polypeptide involved |
|
|
Translation |
The synthesis of proteins from RNA |
|
|
Genetic code |
A triplet of nucleic acid bases (codon) encodes a single amino acid; there are specific codons for starting and stopping translation |
|
|
Degenerate code |
Multiple codons encode a single AA |
|
|
Anticodon |
On tRNA, recognizes codon |
|
|
Anticodon |
On tRNA, recognizes codon |
|
|
Wobble |
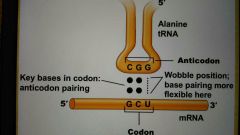
Irregular base pairing allowed at third position of tRNA |
|
|
Stop codons |
Terminate transaction (UAA, UAG, and UGA) |
|
|
Start codon |
Translation begins with AUG (methionine) |
|
|
Reading frame |
Triplet code requires translation to begin at the correct nucleotide |
|
|
Shine-Dalgarno sequence |
Ensures proper reading frame |
|
|
Open reading frame (ORF) |
AUG followed by a number of codons and a stop codon in the same reading frame |
|
|
Codon bias |
Multiple codons for the same AA are not used easily; varies with organism; correlated with tRNA availability; Cloned genes from one organism may not be translated by recipient organism because of codon bias. |
|
|
Transfer RNA |
At least one tRNA per AA; bacterial cells have 60 different tRNAs; Mammalian cells have 100-110 different tRNAs; specific for both a codon and it's cognate AA |
|
|
What are tRNA and AA brought together by? |
aminoacyl-tRNA synthetases; ATP is required to attach AA to tRNA |
|
|
What is the shape of tRNA? |
Cloverleaf |
|
|
Anticodon |
Three bases of tRNA that recognize three complementary bases on mRNA |
|
|
Anticodon |
Three bases of tRNA that recognize three complementary bases on mRNA |
|
|
Why is fidelity of recognition process between tRNA and aminoacyl-tRNA synthetase critical? |
Incorrect AA could result in a faulty or nonfunctioning protein. |
|
|
Ribosomes |
Sites of protein synthesis; thousand per cell; composed of 30s and 50s in prokaryotes to form a 70s (Svedberg units) ribosome in bacteria (40+60=80 in eukaryotes); combination of rRNA and protein |
|
|
How many different distinct ribosomal proteins does E. Coli have? |
52 |
|
|
What are the three main steps that translation is broken down into? |
1. Initiation 2. Elongation 3. Termination |
|
|
Initiation |
Two ribosomal subunits assemble with mRNA; Begins at an AUG start codon |
|
|
Elongation |
AAs are brought to the ribosome and are added to the growing polypeptide. Occurs in the A and P sites of a ribosome. Translocation occurs |
|
|
Translocation |
Movement of the tRNA holding the polypeptide from the A to the P site |
|
|
Termination |
Occurs when ribosome reaches a stop codon. Release factors are involved. Ribosome subunits then dissociate. Subunits free to form new initiation complex and repeat process. |
|
|
Release factors |
Recognize stop codon and cleave polypeptide from tRNA |
|
|
Polysomes |
A complex formed by ribosome simultaneously translating one mRNA |
|
|
What is the advantage to having the ribosomes packed together? |
Process occurs faster |
|
|
Denaturation |
Occurs when proteins are exposed to extremes of heat, pH, or certain chemicals; causes the polypeptide chain to unfold; destroys the 2°, 3°, and/or 4° structure of the protein |
|
|
Renaturation |
Bring back function to protek; may or may not occur when brought back to normal conditions |
|
|
Molecular Chaperones or chaperonins |
Assist in folding, not Incorporated into protein; can also aid in refolding partially denatured proteins. |
|
|
Signal sequences |
Found on proteins requiring transport from cell; 15-20 residues long; found at the beginning of a protein molecule. Signal the cells secretory system; prevent protein from completely folding; needed for exportation |
|
|
SecA |
Helps feed linear protein through cytoplasmic membrane |
|
|
Sec system |
Main secretory system |
|
|
Translational apparatus |
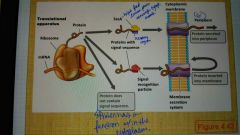
|
|
|
The Tat system |
Secretion of folded proteins; proteins that fold in the cytoplasm are exported by a transport system distinct from SEC, called the Tat protein export system (iron-sulfur proteins and redox proteins) |
|
|
Secretion of proteins types I - VI |
All are a large complex of proteins that form channels through membranes; types 2 and 4 depend on Sec or Tat; types 1, 3, 4 and 6 do not require SEC or Tat |
|
|
Secretion of proteins types I - VI |
All are a large complex of proteins that form channels through membranes; types 2 and 4 depend on Sec or Tat; types 1, 3, 4 and 6 do not require SEC or Tat |
|
|
Injectisome |

|
|
|
Growth |
Increase in the number of cells |
|
|
Binary fission |
Cell division following enlargement of a cell to twice it's minimum size. |
|
|
Generation time |
Time required for microbial cells to double in number (varies by organism and their environment); most prokaryotes have a shorter gen time than Eukaryotes; dependent on growth medium and incubation conditions |
|
|
What does each daughter cell recieves during division in prokaryotes? |
A chromosome and sufficient copies of all other cell constituents to exist as an independent cell. |
|
|
Cell growth for one generation |
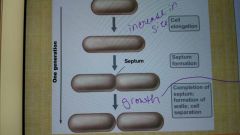
|
|
|
Fts proteins |
Filamentous temperature-sensitive proteins; they are essential for cell division in all prokaryotes. They interact to form the divisome. |
|
|
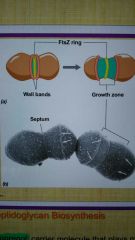
FtsZ |
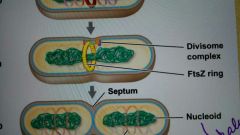
Forms ring around center of the cell; related to tubulin |
|
|
ZipA |

Anchor that connects FtsZ ring to cytoplasmic membrane. |
|
|
FtsA |
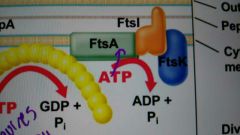
Helps connect FtsZ ring to membrane and also recruits other divisome proteins; related to actin |
|
|
Divisome complex |
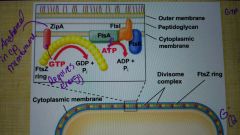
|
|
|
Where is the nucleoid region present in the cell? |
Almost always up against the side |
|
|
Where is the nucleoid region present in the cell? |
Almost always up against the side |
|
|
How are cell walls formed in cocci? |
They grow in opposite directions outward from the FtsZ ring |
|
|
How are cell walls formed in cocci? |
They grow in opposite directions outward from the FtsZ ring |
|
|
How are cell walls formed in rod-shaped cells? |
Growth occurs at several points along the length of the cell. |
|
|
Bactoprenol |

Carrier molecule that plays major role in insertion of peptidoglycan precursors. C55 alcohol. Bonds to N-acetylglucosamine/N-acetylmuramic acid/pentaprism peptidoglycan precursor |
|
|
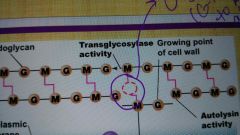
Glycolases |
Enzymes that interact with bactoprenol; they listen the rigid cell wall up for growth; insert wall precursors into growing points of cell wall; catalyze glycosidic bind formation |
|
|
What has to be broke in the growth site of peptidoglycan? |
|
|
|
Transpeptidation |
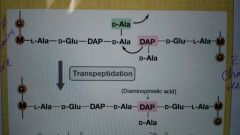
Final step in cell wall synthesis; forms the peptide cross-links between muramic acid residues in adjacent glycan chains; inhibited by the antibiotic penicillin (and others) |
|
|
What form of amino acids are not very common in nature? What one are? |
D; L |
|
|
Asynchronous growth |
Bacteria grow exponentially but daughter cells do not divide at the same exact moment |
|
|
Exponential growth |
Growth of a microbial population in which cell numbers double within a specific time interval |
|
|
Cell replication over time |

|
|
|
Batch culture |
A closed-system microbial culture of fixed volume. |
|
|
What are the four phases of a typical growth curve for a population of cells grown in a closed system? |
Lag, exponential, stationary, and death |
|
|
Lag phase |
No growth, increase in cell size |
|
|
Exponential phase |
Log phase; exponential growth |
|
|
Stationary phase |
Exponential growth stops; horizontal line; still division but cell growth= cell death |
|
|
Death phase |
"decline phase"; exponential decrease, death outnumbers cell division |
|
|
Survival phase |
5th phase, not always mentioned; horizontal line near zero, but above zero then finally complete death |
|
|
What are two ways to count colonies and determine their phase? |
Viable count and turbidity |
|
|
Turbidity |
Measured with a spectrophotometer; it counts both live and dead cells that haven't lysed; these measurements are indirect, rapid, and are useful for measuring microbial growth. Most often measured with a spectrophotometer. Measurement referred to as optical density (OD) |
|
|
Continuos culture |
An open system microbial culture of fixed volume. There is fresh broth and culture brought in at a constant rate; want growth to be locked in late log |
|
|
Batch culture |
Adding new O2 with shaking, but not adding new nutrients |
|
|
Chemostat |

Most common type of continuous culture device which allows control of growth rate and pop density independently and simultaneously. |
|
|
Dilution rate |
Rate at which fresh medium is pumped in and spent medium is pumped out. |
|
|
In a chemostat what is growth rate controlled by? |
Dilution rate |
|
|
In a chemist the growth yield (cell #/ml) is controlled by what? |
The concentration of the limiting nutrient |
|
|
Since in a batch culture growth conditions are constantly changing its impossible to independently control what? |
Both growth parameters |
|
|
What are chemostat cultures sensitive to? |
The dilution rate and limiting nutrient concentration |
|
|
What happens when chemostat cultures are at too high a dilution rate? |
The organism is washed out |
|
|
What happens when cultures in a chemostat are at too low of a dilution rate? |
The cells may die from starvation |
|
|
In a chemist, increasing concentration of a limiting nutrient results in what? |
Greater biomass but the same rate |
|
|
Hemacytometer |
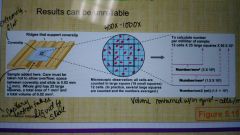
A way to enumerate cells with microscopic observations which can have unreliable results. By picking cells at midlog phase most cells will be viable. |
|
|
Viable cell counts (plate counts) |
Measurement of living, reproducing population. Can be highly unreliable when used to assess total cell numbers of natural samples. By using selective culture media and growth conditions you can target only a particular species. These are viable but non culturable. |
|
|
What are the two main ways to perform plate counts? |
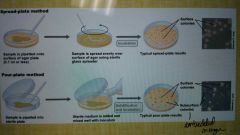
Spread plate method and pour plate method (incorporate bacteria into agar) |
|
|
What are some limitations to performing viable cell counts? |
They have to incubate and need time to grow. They may be hard to grow in a lab setting. |
|
|
Serial dilutions |

|
|
|
To relate a direct cell count to a turbidity value, what must first be established? |
A standard curve |
|
|
When does turbidity work best? |
For low ODs; >5.0 OD may need further dilution |
|
|
Turbidity measurements |
They are quick and easy to perform, they typically do not require destruction or significant disturbance of sample. This is sometimes problematic (clumps or biofilm forms), not accurate at high ODs. |
|
|
qPCR |
Type of real time PCR but for more quantitative studies. Standard PCR but with dye Incorporated into the product; detection for fluorescence over time. |
|
|
Real time PCR |
Amp is in real time; more qualitative analysis. RNA is less stable than DNA so if it's present then there are live samples. |
|
|
PCR |
Contains 2 primers, forward and reverse, (complementary to the targeted DNA sequence), dNTPs (DNA), polymerase Taq (thermistor aquatis), there is DNA/template, fluorescent dye, and a buffer (ions that Pol needs as a cofactor) |
|
|
Cardinal temperatures |
The minimum, optimum, and maximum temperatures at which an organism grows. |
|
|
What occurs at the minimum growth temp? |
Membrane gelling; transport processes are so slow that growth cannot occur |
|
|
What occurs in between the minimum and optimum temp of growth? |
Enzymatic reactions are occurring at increasingly rapid rates |
|
|
What is occurring at the optimum temp of growth? |
Enzymatic reactions are occurring at a maximal possible rate. |
|
|
What is occurring at the maximum temperature of growth? |
Protein denaturation; collapse of the cytoplasmic membrane; thermal lysis |
|
|
Cardinal temp growth curve |

|
|
|
Psychrophile |
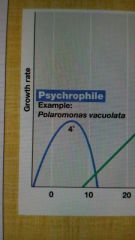
Low temp optima; below freezing point of water |
|
|
Mesophile |
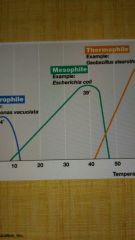
Midrange temp optima; human assoc organisms 25-40°C |
|
|
Thermophile |
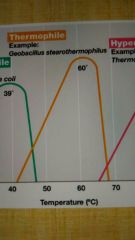
High temp optima; above 45 or 50°C to 80°C |
|
|
Hyperthermophile |

Very high temp optima above 80°C; "extreme thermophiles" can go above 100°C and still grow |
|
|
What adaption support psychrophily? |
1.Production of enzymes that function optimally in the cold and features that may provide more flexibility of proteins so they don't freeze at colder temperatures. 2. Transport processes that function optimally at low temperatures |
|
|
What are some features that allow for more flexibility of proteins seen psychrophily? |
More alpha helices than beta sheets, more polar and less hydrophobic amino acids, fewer weak bonds and decreased interactions between protein domains. |
|
|
What are some modifications in transport processes in psychrophily which allows them to operate optimally at low temps? |
Modified cytoplasmic membranes, and high unsaturated fatty acid content |
|
|
At temps higher than about what can only prokaryotic life forms exist? |
~65°C |
|
|
At temps higher than about what can only prokaryotic life forms exist? |
~65°C |
|
|
Hyperthermophiles in hot springs |
Chemoorganotrophs and Chemolithotrophs present; high prokaryotic diversity of archaea and bacteria are represented |
|
|
Which organisms have the highest temperature optima? |
Archarea |
|
|
Which can grow at higher temps phototrophic or nonphototrophic organisms? |
Nonphototrophic |
|
|
What are some molecular adaptions to thermophily? |
1. Enzyme and proteins function Optimally at high temperatures; features that provide thermal stability. 2. Modifications in cytoplasmic membranes to ensure heat stability |
|
|
What are some features that provide thermal stability in thermophiles? |
Critical amino acid substituents in a few locations provide more heat folds, an increased number of ionic bonds between basic and acidic amino acids resist unfolding in the aqueous cytoplasm, and production of solutes |
|
|
What are the modifications seen in thermophiles cytoplasmic membranes that ensures heat stability? |
Bacteria have lipids rich in saturated fatty acids and archaea have a lipid monolayer rather than a bilayer. |
|
|
Give an example for how enzymes produced by hyperthermophiles can be used in industrial microbiology. |
Taq polymerase (from thermus aquatis. 96-97°C; used for PCR) |
|
|
Neutrophiles |
Organisms that grow best between pH of 6 and 8. |
|
|
Acidophiles |
Organisms that grow best at low pH (<6); some are obligate acidophiles (has to be in acidic ENV. To grow) and there membranes are destroyed at neutral pH; stability of the cytoplasmic membrane is critical |
|
|
Alkaliphiles |
Organisms that grow best at high pH (>9); some have sodium motive force rather than proton motive force |
|
|
Internal pH of a cell |
Has to stay relatively close to neutral even though the external pH is highly acidic or basic; *exceptions has been found to be as low as 4.6in acidophiles and as high as 9.5 in alkaliphiles. |
|
|
Positive water balance |
The tendency of water to move into the cell because of the higher internal conc of solute. |
|
|
Halite |
Mineral-> rock salt |
|
|
Halophiles |

Organisms that grow best at reduced water potential; have a specific requirement for NaCl |
|
|
Extreme halophiles |

Organisms that require high levels (15-30%) of NaCl for growth |
|
|
Halotolerant |

Organisms that can tolerate some reduction in water activity of environment but generally grow best in the absence of the added solute |
|
|
Nonhalophile |

|
|
|
Osmophiles |
Organisms that live in environments high in sugar as solute ( or other non sodium solute) |
|
|
Xerophiles |
Organisms able to grow in very dry environments; they capture and hold onto Environments |
|
|
Aerobes |
Require oxygen to live |
|
|
Anaerobes |
Do not require oxygen and may even be killed with oxygen exposure |
|
|
Facultative orgsnisms |
Can live with or without oxygen, but prefer oxygen if it's available; some can utilize both at the same time |
|
|
Aerotolerant anaerobes |
Can tolerate oxygen and grow in its presence even though they cannot use it |
|
|
Microaerophiles |
Can use oxygen only when it is present at levels reduced from that in air. |
|
|
Reducing agents |
Chemicals that may be added to culture media to reduce oxygen |
|
|
What toxic forms of oxygen. Can be formed in the cell? |

|
|
|
What enzymes are present to reduce most of the toxic species of oxygen? |
Catalase, peroxidase, superoxide dimutase, superoxide reductase |
|
|
Sterilization |
The killing or removal of all viable organisms within a growth medium. |
|
|
Satellite colonies |

|
|
|
Bacteriostatic |
Inhibits growth; does not kill |
|
|
Bacteriocidal |
Kill bacteria |
|
|
Inhibition |
Effectively limiting microbial growth; not necessarily 100%. * Some antibiotics like penicillin and esp. Ampicillin |
|
|
Decontamination |
The treatment of an object to make it safe to handle; removing microbes from an object but not necessarily all of them, just to a safe level. |
|
|
Disinfection |
Directly targets the removal of all pathogens, not necessarily all microorganisms |
|
|
Heat sterilization |
The most widely used method of controlling microbial growth. High temps denature macromolecules. |
|
|
Decimal reduction time |
Amount of time required to reduce viability tenfild |
|
|
Endospores |
Resistant cells produced by bacteria that can survive heat that would rapidly kill vegetative cells. |
|
|
Autoclave |
Sealed device that uses steam at 121°C under pressure of 1.5 ATM (allows temp to go high) which kills microbes including endospores. |
|
|
Pasteurization |
The process of using precisely controlled heat to reduce the microbial load in heat-sensitive liquids; does not kill all organisms so it is different from sterilization |
|
|
What types of light waves can be used for sterilization? |
Microwaves, UV, X-rays, gamma rays, and electrons |
|
|
UV |
Has sufficient energy to cause modifications and breaks in DNA; useful for decontaminating surfaces, but cannot penetrate solid, opaque or light absorbing surfaces. |
|
|
Lamenter file hood |

Uses IV light in nonpathogenic labs |
|
|
Ionizing radiation |
Electromagnetic radiation that produces ions and other reactive molecules (ex. Ozone), generates electrons, hydroxyl radicals, and hydride radicals. Down microrganisms are more resistant to radiation than others |
|
|
Sources of radiation |
Cathode ray tubes, x-rays, and radioactive nuclides |
|
|
Radiation |
Used for sterilization in the medical field and food industry; it is approved by the WHO and is used in the USA for decontaminating foods particularly susceptible to microbial contamination such as hamburger, chicken and spices |
|
|
Filtration |
One of the most widely used forms of control. Avoids the use of heat on sensitive liquids and gases; pores of the filter are too small for organisms to pass through but allow liquids and gases to pass through. |
|
|
What are some times of filters used? |
Nylon, cellulose, and nitrocellulose |
|
|
Depth filters |
HEPA filters |
|
|
Depth filters |
HEPA filters |
|
|
Membrane Filters |
Function more like a sieve |
|
|
What is the standard pore size of filters? |
.45u - .2um |
|
|
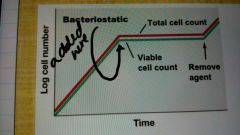
How can antimicrobial agents be classified? |
Bacteriostatic, bacteriocidal, and bacteriolytic |
|
|
Bacteriostatic |
Inhibits cell growth does not kill |
|
|
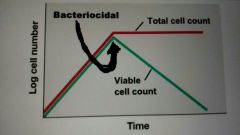
Bacteriocidal |
Death of cells |
|
|
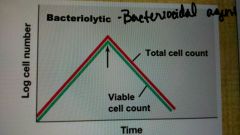
Bacteriolytic |
Cells lyse |
|
|
Minimum inhibitory concentration (MIC) |
The smallest amount of an agent needed to inhibit growth of a microorganism; varies with the organism used, inoculum size, temperature, pH, etc. |
|
|
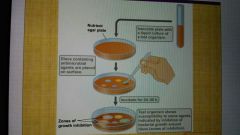
Disc diffusion assay |
Uses solid media; antimicrobial agents is added to the filter paper disc (embedded in or on them) and mic is reached at some distance |
|
|
Zone of inhibition |
Area of no growth around disk |
|
|
What are the two categories that antimicrobial agents can be divided into? |
1. Products used to control microorganisms in commercial and industrial applications (chem in foods, air conditioning cooling towers, textile and paper products, fuel tanks) 2. Products designed to prevent growth of human pathogens in inanimate environments and on external body surfaces (sterilants, disinfectants, sanitizers, and antiseptics) |
|
|
Genome |
Entire complement of genetic information, includes genes, regulatory sequences, and noncoding DNA |
|
|
Genomics |
Study of the genome; discipline of mapping, sequences, analyzing, and comparing genomes |
|
|
What was the first genome sequenced (1976) |
RNA virus MS2; 3569 bp |
|
|
What was the first cellular genome sequenced in 1995? |
Haemophilus influenza; 1,830,137 bp |
|
|
Sequencing |
Determining the precise order of nucleotides in a DNA or RNA molecule |
|
|
Sanger dideoxy method |
Invented by Nobel prize winner Fred Sanger, dideoxy analogs of dNTPs used in conjunction with dNTPs, analog prevents further extension of DNA chain, bases are labeled with radioactivity, gel electrophoresis is then performed on products. |
|
|
Automated DNA sequencing systems |
Based on Sanger method, radioactivity replaced by fluorescent dye. |
|
|
Shotgun sequencing |
Entire genome is cloned, and resultant clones are sequenced, much of the sequencing is redundant, generally 7- to 10-fold coverage; computer algorithms are used to look for replicate sequences and assemble them. |
|
|
Second generation DNA sequencing |
Generated data 100x faster than Sanger method; massively parallel methods (lg # of samples sequenced side by side), uses increased computer power and miniaturization |
|
|
454 sequencing system |
DNA is broken into small segments, then is amplified using PCR, light is released each time a base is added to DNA strand. The instrument measures the release of light. It can handle only short stretches of DNA |

