![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
57 Cards in this Set
- Front
- Back
|
What kind of molecule is tRNA and what does it link?
|
1. Adapter molecule
2. Links mRNA codons to it's anti-codons |
|
|
Each amino acid has its own adapter (tRNA). T or F?
|
True
|
|
|
There are two key events to make a protein with tRNA. What are they?
|
1. tRNA must read mRNA codons correctly
2. tRNAs must deliver amino acids corresponding to each codon |
|
|
What are the 3 primary functions of tRNA?
|
1. Must be charged (carrying an amino acid)
2. Associates with mRNA molecules 3. Interacts with ribosomes |
|
|
Where is the anticodon located on the tRNA? The amino attachment site? Hydrogen bonds?
|
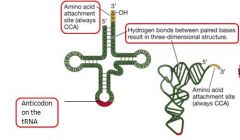
See illustration
|
|
|
The sequence of the attachement site is ALWAYS ACC. T or F?
|
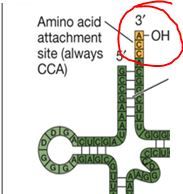
False: It is always CCA
|
|
|
______ different codons encode the ______ amino acids in all protiens
|
61, 20
|
|
|
Explain the wobble phenomenon
|
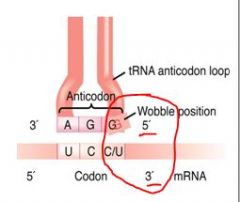
The specificity for the base at the 3' end of the codon and the 5' end of the anti-codon are not always observed.
For example: The codons CGA, GCC and GCU can all be recognized by just one single species of tRNA. |
|
|
Wobble does not allow the genetic code to be redundant. T or F?
|
False: Wobble does not allow genetic code to be ambiguous
|
|
|
What is a charged tRNA?
|
A tRNA that has an amino acid attached to it
|
|
|
Exactly how is a tRNA charged?
|

The charge is "activated" by the enzyme Aminoacyl-tRNA snythase
|
|
|
The small subunit ribosome is made up of what?
|
One molecule of rRNA and 33 protiens
|
|
|
The large subunit has three tRNA binding sites. What are they?
|
A site (amino)
P site (polypeptide) E site (exit) |
|
|
Illustration of how tRNA is charged
|
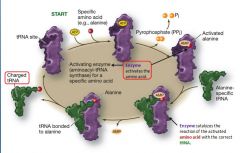
See illustration
|
|
|
What are some characteristics of the enzyme aminoacyl-tRNA synthase?
|
1. Highly specific for each amino acid and it corresponding tRNA
2. Has three active sites that recognize ATP, a specific amino acid and a specific tRNA |
|
|
Explain the purpose of the three sites (EPA) on the larger subunit.
|
A: a charged tRNA anticodon binds to an mRNA codon
P: tRNA adds its amino acid to the growing chain E: tRNA sits here before being released from ribosome |
|
|
The large subunit ribosome is made up of what?
|
Three different molecules of rRNA and 49 protiens
|
|
|
The ribosome has "fidelity function". What does this mean?
|
It ensures that mRNA and tRNA interactions are accurate and that the bindings are appropriate
|
|
|
In elongation, the large subunit catalyzes two reactions. What are they?
|
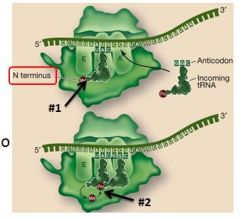
1. Breaks bonds between tRNA in P site and its amino acid
2. Forms a peptide bond between the above acid and the acid on tRNA in the A site |
|
|
How do codons and anticodons bind together?
|
Hydrogen bonds
|
|
|
During elongation, the first tRNA releases its amino acid. It then moves to the ______ site and becomes ______ again.
|
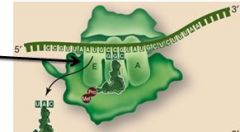
E, charged
|
|
|
Ribosomes are specific and can only make a certain protein. T or F?
|
False: They are NOT specific and can make ANY protien
|
|
|
During termination, stop codons bind to what type of protien?
|
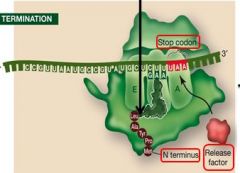
A protien release factor
|
|
|
If no hydrogen bonds form between codons and anticodons, what probably happened?
|
There was an incorrect base pairing - the ribsome will reject the tRNA
|
|
|
In termination, what does the protein release factor do?
|
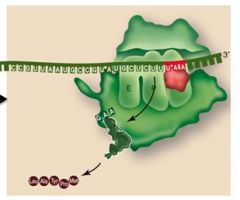
Hydrolysizes the bond between the chain and tRNA on the P site (the chain separates from the ribosome)
|
|
|
Translation has 3 steps. What are they?
|
1. Initiation
2. Elongation 3. Termination |
|
|
A single strand of mRNA can have MANY ribosomes on it creating MANY chains. What is this assemblage called?
|
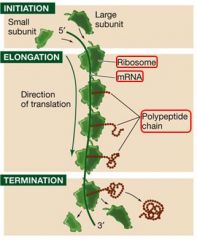
A polyribosome (or polysome)
|
|
|
What does an initiation complex consist of?
|

A charged tRNA, small ribsome unit that are both bound to mRNA
|
|
|
Nucleus' mitochondria and chloroplasts have special receptor protiens that the chains (signal sequence) binds to. What are these protiens called?
|
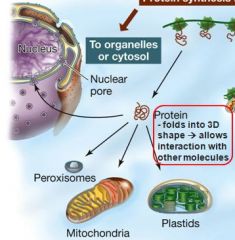
Docking protiens
|
|
|
In prokaryotes, the rRNA fist binds to a special site on the mRNA. What is this special site?
|

The Shine Dalgarno Sequence
|
|
|
If a chain is destined for the ER, the chain's signal sequence binds to a ______ BEFORE translation is complete.
|
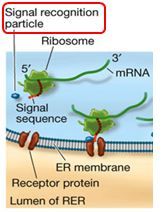
Signal recognition particle
|
|
|
In initiation, the small subunit binds to the ______ cap on the mRNA and moves until it reaches the start codon
|
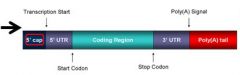
5'
|
|
|
Illustration of translation occuring at the ER membrane.
|
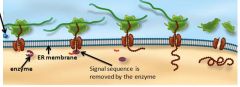
See illustration
|
|
|
During initiation, the start codon is ALWAYS AUG. What amino acid does this code for?
|
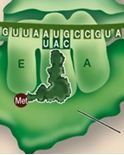
Methionine (MET) (It is sometimes removed after translation)
|
|
|
Once a protien is inside the ER lumen, there are two types of signals that can direct the protein to its final destination. What are they?
|
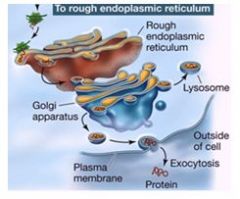
1. A certain sequence directs it to remain inside the ER
2. Sugars in the golgi result in glycoproteins and these end up at the plasma membrane OR vacuole in plants |
|
|
During initiation, after the tRNA, mRNA and small subunit connect, what joins the complex?
|
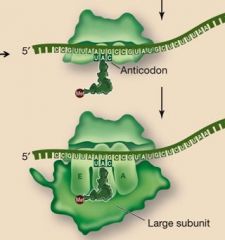
The large subunit (tRNA ends up in the P site of this unit)
|
|
|
What is glycosylation modification?
|

Addition of sugars to form glycoproteins. Important for targeting and recognition
|
|
|
What are the "ingredients" that make up the "initiation factors"?
|
mRNA, two ribosomal units (lrg and sml), and methionine-charged tRNA
|
|
|
How are the ribosomal subunits bound together?
|
Non-covalently (by ionic and hydrophobic forces)
|
|
|
How is it determined which tRNA enters into the A site?
|
Any tRNA whose anticodons is COMPLEMENTARY to the codon that is currently sitting in the A site
|
|
|
In elongation, the incoming second tRNA enter the ______ site
|
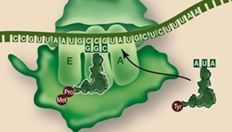
A
|
|
|
Elongation catalyzes two reactions. Because of these two reactions, it is considered to have ______ activity.
|
Peptidyl transferase
|
|
|
During elongation, if rRNA is destroyed, all activity STOPS! Thus, rRNA is the ______ in peptidyl tranferase activity.
|
Catalyst (this supports the idea that RNA evolved before DNA)
|
|
|
All of the steps in elongation are assisted by ribosomal proteins. What are these proteins collectively called?
|
Elongation factors
|
|
|
The process of elongation ultimately produces what?
|
A long polypeptide chain
|
|
|
When is the only time a tRNA can "sit" in site E (the last slot)?
|
If it has lost its amino acid (which occured in site P)
|
|
|
When talking about codons, we are referring to ______ and when talking about anticodons, we are referring to ______.
|
mRNA, tRNA
|
|
|
When a stop codon (UAA for example) enters the A site, what happens?
|
Translation ends
|
|
|
What is the N terminus?
What is the C terminus? |
1. The start (methionine)
2. The end (last amino acid) |
|
|
What happens to the subunits at the very end of translation?
|
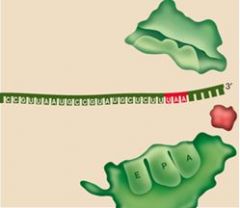
The separate
|
|
|
Newly formed polypeptide chain have a sort of "address label". What is this called?
|
A signal sequence
|
|
|
What exactly do docking receptor proteins do?
|
They form a channel to let the protein through
|
|
|
What directs the polypeptide chain to the nucleus?
|
A nuclear localization signal
|
|
|
If a polypeptide chain is destined to travel to the ER, its signal sequence binds to a ______ BEFORE translation is complete.
|
Signal recognition particle
|
|
|
After translation, three different modifications can occur. What are they?
|
1. Proteolysis
2. Glycosylation 3. Phosphorylation |
|
|
What is proteolysis modification?
|
Cleaving of a long chain with porteases - the fragments fold into different shapes
|
|
|
What is phosphorylation modification?
|
The addition of phosphate groups by protein kinases which alter the protein shape
|

