![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
81 Cards in this Set
- Front
- Back
|
RNAP II is released from TF's to _________ RNA, in eukaryotes this occurs with the aid of ________ factors.
|
synthesize, factors
|
|
|
Elongation factors ___________ the continuation of transcription.
|
promote
|
|
|
Elongation factors promote the continuation of ____________ in RNAP II.
|
elongation, or transcription
|
|
|
RNAP II is _______ by dephosphorylation of CTD.
|
RNAP II is released by dephosphorylation of CTD.
|
|
|
RNAP II is released by ___________ of CTD.
|
RNAP II is released by dephosphorylation of CTD.
|
|
|
RNAP II is released by dephosphorylation of ________.
|
RNAP II is released by dephosphorylation of CTD.
|
|

|

|
|

|

|
|

|

|
|

|

|
|

|

|
|

|

|
|
|
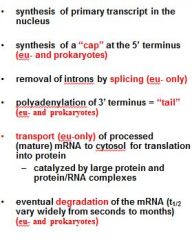
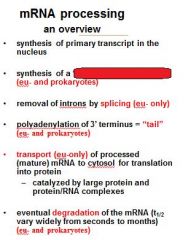
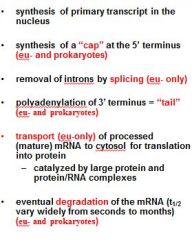
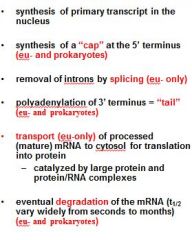
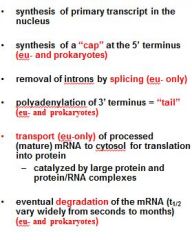
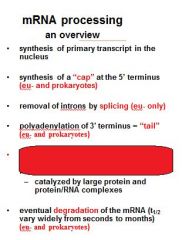
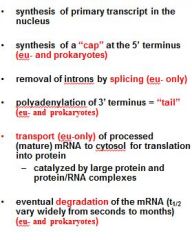
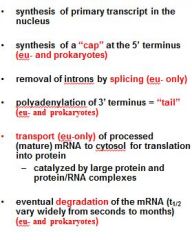
What is a mature transcript?
|
5' cap
no introns 3' PolyA tail |
|
|
Functions of the 5' Cap
1. ????? 2. regulation of export 3. promotion of translation |
Functions of the 5' Cap
1. to protect the mRNA from degradation via exoribonucleases 2. regulation of export 3. promotion of translation |
|
|
Functions of the 5' Cap
1. to protect the mRNA from degradation via exoribonucleases 2. ??????? 3. promotion of translation |
Functions of the 5' Cap
1. to protect the mRNA from degradation via exoribonucleases 2. regulation of export 3. promotion of translation |
|
|
Functions of the 5' Cap
1. to protect the mRNA from degradation via exoribonucleases 2. regulation of export 3. ?????? |
Functions of the 5' Cap
1. to protect the mRNA from degradation via exoribonucleases 2. regulation of export 3. promotion of translation |
|
|
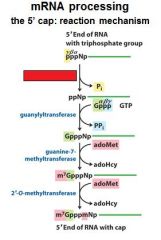
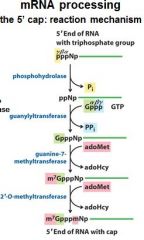
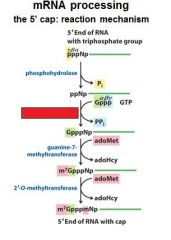
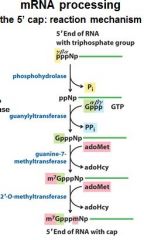
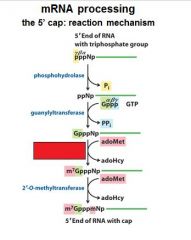
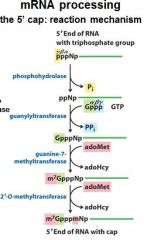
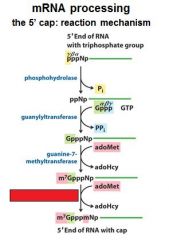
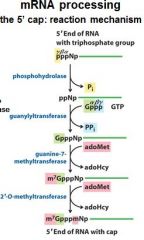
What is the 5' cap of a mature transcript?
|
7-methyl-guanosine linked by a tri-phosphate linkage with possible 2' methylation of the first two ribonucleotides
|
|
|
What is important to remember about the 5' cap?
|
7-methyl-guanosine
tri-phosphate linkage possible 2' methylation of the first two ribonucleotides |
|
|
The enzymes of the __________ ___________ complex are tethered to the CTD of RNAP II.
|
The enzymes of the cap-synthesizing complex are tethered to the CTD of RNAP II.
|
|
|
The enzymes of the cap-synthesizing complex are tethered to the _______________________.
|
The enzymes of the cap-synthesizing complex are tethered to the CTD of RNAP II.
|
|
|
Once complete the _____________ complex is replaced by the nuclear cap-binding complex.
|
Once complete the cap-synthesizing complex is replaced by the nuclear cap-binding complex.
|
|
|
Once complete the cap-synthesizing complex is replaced by the nuclear ______________ complex.
|
Once complete the cap-synthesizing complex is replaced by the nuclear cap-binding complex.
|
|
|
|
|
|
|
|
|
|
|
|
|
|

|

|
|

|

|
|

|

|
|

|

|
|

|

|
|

|

|
|
|
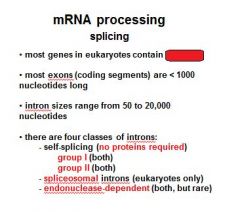
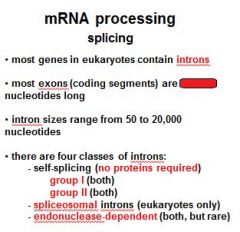
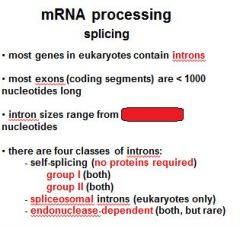
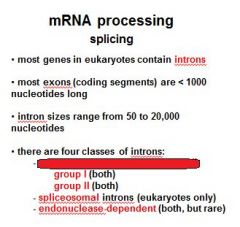
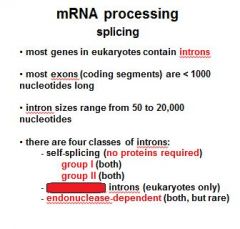
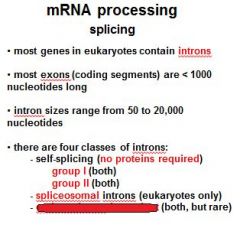
Name the four classes of introns?
|
group I (no proteins required)
group II (no proteins required) spliceosomal introns endonuclease dependent introns |
|
|
What is the cofactor for group I introns?
|
guanine nucleoside or nucleotide
|
|
|
Guanine nucleoside or nucleotide is the cofactor for what intron?`
|
group I
|
|
|
What is the cofactor for group II introns?
|
an adenylate residue within the intron sequence
|
|
|
an adenylate residue within the intron sequence is the cofactor for what intron?`
|
group II
|
|
|
Self splicing in group I and group II introns is two _________________ reactions.
|
transesterification
|
|
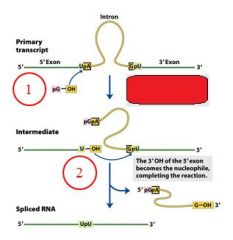
GROUP I INTRONS
|
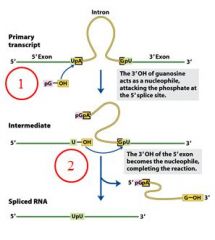
|
|
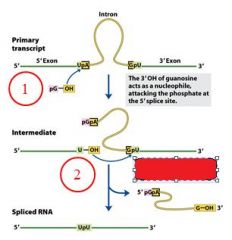
GROUP I INTRONS
|
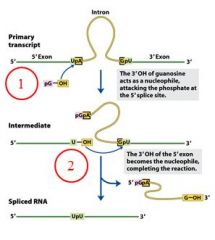
|
|
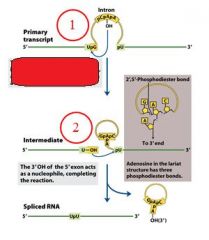
GROUP II INTRONS
|
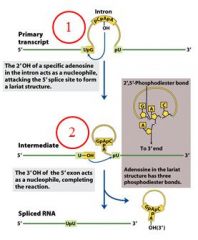
|
|
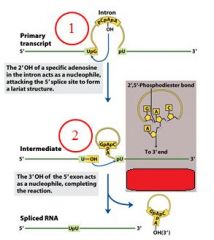
GROUP II INTRONS
|
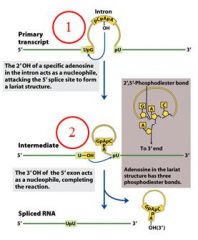
|
|
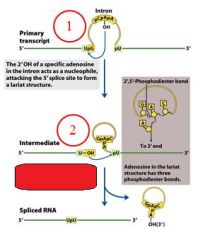
GROUP II INTRONS
|
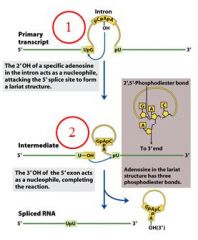
|
|
|
The specific cofactor for group II introns is?
|
the 2'OH of an internal adensoine
|
|

|

|
|

|

|
|

|

|
|

|

|
|

|

|
|
|
Which snRNP catalyzes splicing?
|
U6
|
|
|
Which snRNP masks the catalytic activity of U6?
|
u4
|
|
|
Which snRNP binds the 5' splice site?
|
U1
|
|
|
Which snRNP binds the 5' splice site and then the 3' splice site?
|
U5
|
|
|
Which snRNP binds the branch site and forms part of the catalytic center?
|
U2
|
|
|
Intron sequences have ___________ sequences at both ends.
|
conserved
|
|
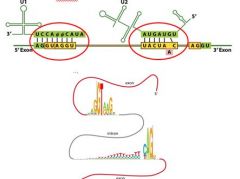
__________________ in snRNA's bind to a consensus 5' splice site and a 3' branch site within an intron.
|
Complementary regions in snRNA's bind to a consensus 5' splice site and a 3' branch site within an intron.
|
|
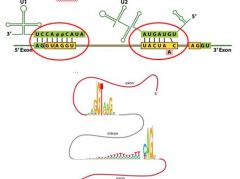
Complementary regions in snRNA's bind to a consensus ________________ and a 3' branch site within an intron.
|
Complementary regions in snRNA's bind to a consensus 5' splice site and a 3' branch site within an intron.
|
|
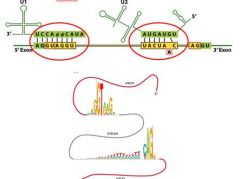
Complementary regions in snRNA's bind to a consensus 5' splice site and a ________________ within an intron.
|
Complementary regions in snRNA's bind to a consensus 5' splice site and a 3' branch site within an intron.
|
|
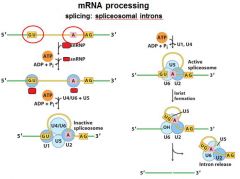
|
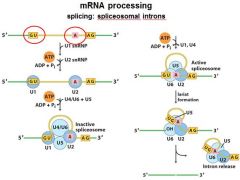
|
|
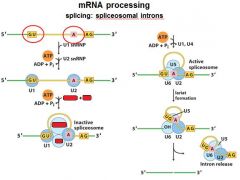
|
|
|
|
|
|
|
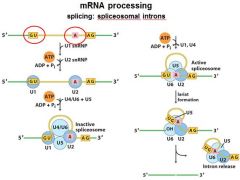
|
|
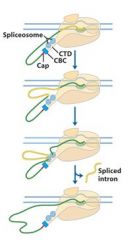
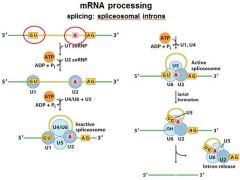
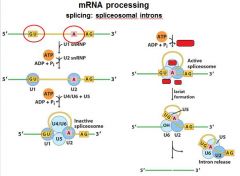
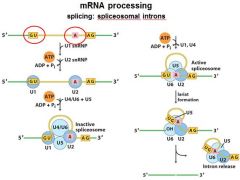
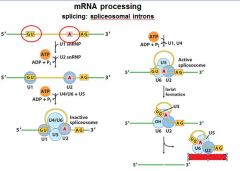
The ______________ is also associated with the CTD of RNAP II. Splicing occurs before synthesis of the primary transcript is complete.
|
The spliceosome is also associated with the CTD of RNAP II. Splicing occurs before synthesis of the primary transcript is complete.
|
|
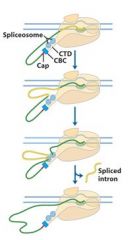
The spliceosome is also associated with the CTD of RNAP II. Splicing occurs _________ synthesis of the primary transcript is ____________.
|
The spliceosome is also associated with the CTD of RNAP II. Splicing occurs before synthesis of the primary transcript is complete.
|
|
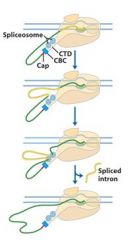
The spliceosome is also associated with the _______ of RNAP II. Splicing occurs before synthesis of the primary transcript is complete.
|
The spliceosome is also associated with the CTD of RNAP II. Splicing occurs before synthesis of the primary transcript is complete.
|
|

|

|
|

|

|
|

|

|
|

|

|
|
|
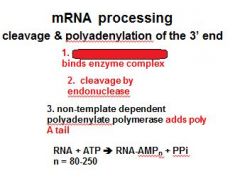
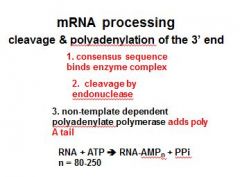
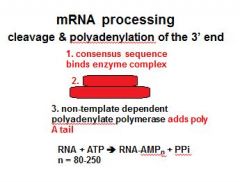
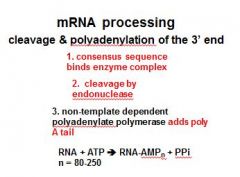
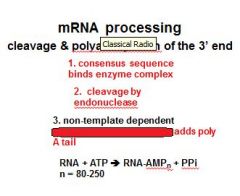
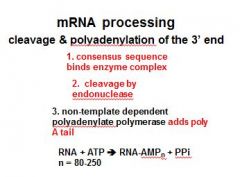
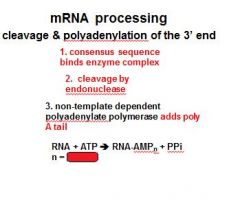
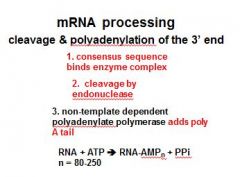
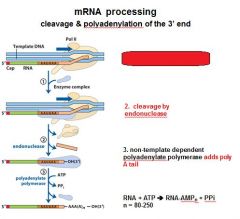
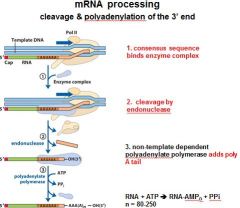
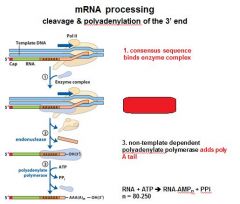
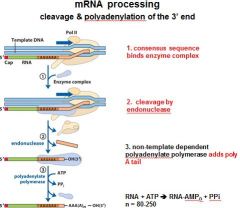
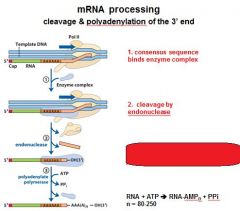
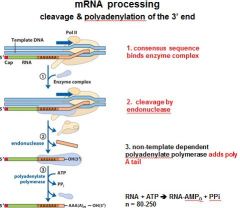
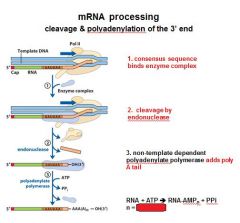
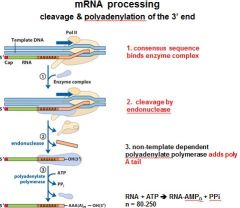
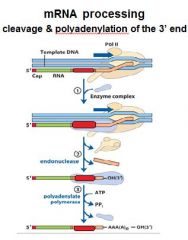
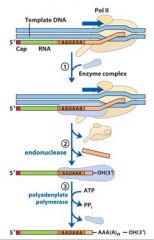
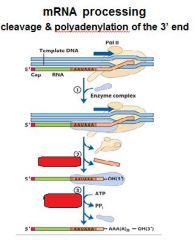
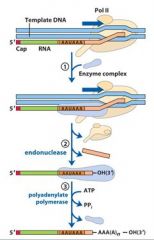
In the cleavage and polyadenylation of the 3' end of a RNA transcript the consensus sequence binds the enzyme complex. What is the consensus sequence?
|
A-A-U-A-A
|
|
|
In the cleavage and polyadenylation of the 3' end of a RNA transcript the consensus sequence binds the enzyme complex. What two proteins are associated with the enzyme complex?
|
endonuclease, polyadenylate polymerase
|
|
|
|
|
|
|
|
|
|
|
|
|
|
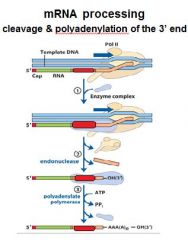
consensus sequence?
|
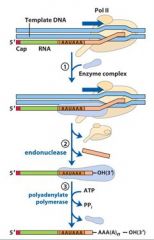
|
|
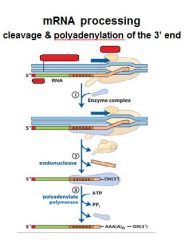
|
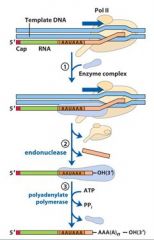
|
|
|
|
|
|
|
|

|

|
|

|

|

