![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
231 Cards in this Set
- Front
- Back
|
Homeostasis |
The bodies ability to maintain regulated variables at a narrow range which supports life. |
|
|
Law of mass balance |
If the amount of a substance in the body is to remain constant, any gain must be offset by a loss and visa versa. E.g. Water lost to external environment via sweat or urine must be balanced by water intake from the external environment. |
|
|
Local control systems |
Restricted to the tissue or cell involved. In local control an isolated change occurs in a cell or tissue. A nearby cell senses the change and responds. |
|
|
Long distance control/reflex control |
Changes that are widespread throughout the body (systemic). The nervous and endocrine systems rely on reflex control. Physiological reflexes can be: Response loops Feedback loops |
|
|
Response loop |
1. Stimulus 2. Sensor 3. Input signal 4. Integrating center 5. Output signal 6. Target 7. Response |
|
|
Positive feedback |
Not-homeostatic Response reinforces a stimulus An outside factor is required to shut off the cycle. E.g. childbirth |
|
|
Negative feedback |
Homeostatic Keep the system at or near set point
Set point = normal range of a regulated variable |
|
|
Biomolecules |
Carbohydrates Lipids Nucleotides Proteins |
|
|
Lipids |
Mostly carbon and hydrogen with a glycerol backbone. 1-3 fatty acid tails. Non-polar & non-soluble Saturated fats = more hydrogens. The more hydrogens the more saturated. |
|
|
Carbohydrates |
Carbon and water. For every carbon there is 2 hydrogen and 1 oxygen. Monosacharides = 5 or 6 carbon sugars (ribose, glucose, lactose) Disacharides = glucose + a monosacharide Polysacharide = glucose polymer |
|
|
Lipid related molecules |
Eicosanoids - modified 20 carbon fatty acids with a complete carbon ring at one end and 2 carbon tails at the other. E.g. prostoglandins - regulate physiological functions Steroids - cholesterol Phospholipids - cell membrane |
|
|
Functional groups |

Combinations of elements that occur repeatedly and move from molecule to molecule as a single unit. Amino group Carboxyl group Hydroxyl group Phosphate group |
|
|
Proteins |
Amino acid polymers. Amino acids consist of a carboxyl group, an amino group and a hydrogen attached to a carbon. There is also a variable R group. |
|
|
Peptide bonds |

Occur when amino acids join to form a chain. The amino group of one amino acid joins the carboxyl group of another |
|
|
Essential amino acids |
Must be obtained through diet. Glutamine, tyrosine, arginine etc... |
|
|
Primary protein structure |
The sequence of any 20 amino acids in a peptide chain |
|
|
Peptides |
Oligiopeptide 2-9 amino acids Polypeptide 10-100 amino acids Protein 100+ amino acids |
|
|
Secondary protein structure |
Covalent bonds form between amino acid peptide chains to form a double helix |
|
|
Tertiary protein structure |
A proteins 3D shape Globular proteins and fibrous proteins such as collagen fibers |
|
|
Quaternary protein structure |
Many sub-units combine with non-covalent bonds (many 3D protein structures come together) E.g. hemoglobin is made of 4 globular protein sub-units |
|
|
Covalent bonds |
Sharing of electrons between atoms. Will always involve a non-metal. Covalent bonds form molecules. Covalent bonds are strong and require energy to break them apart. |
|
|
Ions |
When an atom or molecule gains or loses an electron it becomes an ION. The ion then has an electrical charge.
|
|
|
Cations |
Metals Lose electrons = overall positive charge |
|
|
Anions |
Non-metals Gain electrons = overall negative charge |
|
|
High energy electrons |
Only occur in certain atoms. They capture energy from their environment and transfer it to other atoms. This energy is used for synthesis, movement and other life processes. |
|
|
Free radicals |
Unstable molecules with an unpaired electron. Contribute to ageing and disease development |
|
|
Ionic bonds |
Metals. Taking or losing electrons via electrostatic attraction. |
|
|
Nucleotides |
Essential for energy and information transfer. Nucleic acids (DNA & RNA) store and transmit genetic information. ATP, AMP and cAMP are single nucleotides. |
|
|
Nucleotide structure |
Nitrogenous bases 5 carbon sugar One or more phosphate groups |
|
|
RNA |
Ribonucleic acid. Single strand nucleic acid. Ribose is the sugar. |
|
|
DNA |
Deoxyribonucleic acid. Double helix - 3D structure. Formed by 2 DNA strands linked by hydrogen bonds between complimentary base pairs. Deoxyribose is the sugar. |
|
|
Nitrogenous bases |
ADENINE -THYMINE = 2 hydrogen bonds ADENINE - URACIL (mRNA) = 2 hydrogen bonds GUANINE - CYTOSINE = 3 hydrogen bonds |
|
|
Proton |
Positive charge Atomic number is calculated by number of protons |
|
|
Neutron |
Neutral |
|
|
Electron |
Negative charge |
|
|
Atomic mass |
Number of protons and electrons |
|
|
Isotope |
An atom that loses or gains a neutron becomes an isotope of the same element |
|
|
Polar molecules |
Overall neutral molecule BUT has regions of partial positive and negative charge. Polar molecules are water soluble. Nitrogen and oxygen have a strong attraction to electrons and are often associated with polar molecules. |
|
|
Hydrogen bonds |
Weak attractive force between a hydrogen and nearby oxygen, nitrogen or fluorine. No electrons are gained or lost, instead oppositely charged regions in polar molecules are attracted to each other. |
|
|
Van der waals forces |
Weak, non-specific attractions between the nucleus of ANY atom and the electrons of nearby atoms. Van der waals allow atoms to pack in closely together to occupy minimum amounts of space. E.g. globular proteins. |
|
|
Solubility |
The degree to which a molecule is able to dissolve in a solvent. Water is the biological solvent. |
|
|
Hydrophillic |
Water loving. Soluble. |
|
|
Hydrophobic |
Water hating Non-soluble lipids and fats. |
|
|
pH |
Power of hydrogen. Free hydrogen ions lower pH which results in an acidic solution. Hydroxyl ions increase pH resulting in a basic solution. Acids = free hydrogen Bases = hydrogen acceptors |
|
|
Buffer |

Any substance that moderates changes in pH. E.g. carbonic acid/bicarbonate buffer. Usually consists of 2 parts. A weak acid and its conjugate base. The role of the buffer is to replace a strong acid or base with a weak acid or base resulting in minimal change to pH. |
|
|
Moles |
An expression of the number of solute molecules without regard to their weight. |
|
|
Electrolyte |
A solution that contains ions and can generate an electrical current. Electrolytes when combined with water dissociate (ionize) into their ions. Strong electrolytes = 100% dissociation (NaCl) Weak electrolytes (HF) = partial dissociation |
|
|
Enzymes |
Biological catalysts Speed up chemical reactions by lowering activation energy. Crucial in metabolism |
|
|
Membrane transporters |
Proteins found imbedded in the cell membrane. Integral and carriers. Move substances between the ICF & ECF |
|
|
Signal molecules |
Proteins and smaller peptides acting as hormones or other signal molecules for cell-to-cell communication. |
|
|
Receptors |
Proteins that bind to signal molecules which then initiate cellular responses. |
|
|
Binding proteins |
Found mostly in ECF. Bind and transport molecules around the body. E.g. hemoglobin transports oxygen. |
|
|
Immunoglobins |
Extracellular immune proteins (antibodies). Protect the body from foreign invaders. |
|
|
Regulatory proteins |
Turn cell process on, off, up & down. |
|
|
Ligand |
Any molecule or ion that binds to another molecule. Ligands that bind to an enzyme or membrane transporter are called SUBSTRATES. |
|
|
Specificity |
The ability of a protein to bind to a specific ligand or a group of ligands. The ligand and binding site must be compatible. When a ligand and binding site combine they interact via hydrogen bonds, ionic bonds and van der waals forces. |
|
|
Induced fit |
When a proteins binding site changes shape to compliment its specific ligand |
|
|
Affinity |
The degree to which a protein is attracted to a ligand. If a protein has a high affinity for a ligand it is more likely to bind. |
|
|
Competition |
Related ligands compete for binding sites. Agonist - mimics a ligand Antagonist - inhibit the effect, block the binding site and overall decrease activity. |
|
|
Saturation |
Proteins reach saturation when all binding sites are occupied. Addition of extra ligand has no effect. |
|
|
Central nervous system |
Brain and spinal cord. The central nervous system acts as the intergrating center for the body |
|
|
Peripheral nervous system |
Located out of the CNS. Consists of afferent (in) neurons and efferent (out) neurons. |
|
|
Afferent division |
Also known as SENSORY division. Afferent neurons send information to the Central nervous system. |
|
|
Efferent division |
Efferent neurons take information from the central nervous system to target cells. There are 2 branches of the efferent division: SOMATIC & AUTONOMIC |
|
|
Somatic divison of PNS |
Controls the skeletal muscles. Voluntary movement. |
|
|
Autonomic division of the PNS |
Controls smooth and cardiac muscle, exocrine glands, some endocrine glands and adipose tissue. Referred to as the visceral nervous system as it controls contraction and secretion in various organs. Autonomic neurons are further divided into the SYMPATHETIC AND PARASYMPATHETIC division. |
|
|
SYMPATHETIC division of the autonomic branch of PNS |
Involuntary. Fight or flight. Increases heart rate etc. |
|
|
PARASYMPATHETIC division of autonomic branch of PNS |
Rest and digest. Slows heart rate. |
|
|
Neuron |

Nerve cell. Functional unit of the nervous system. |
|
|
Interneurons |

Found in the central nervous system. Come in variety of forms and commonly have complex branching. |
|
|
Sensory neurons |

Carry sensory information from the periphery to the central nervous system (afferent neurons). |
|
|
Efferent neurons |

Carry information from the central nervous system to the periphery. |
|
|
Nerve |
The long axons of both afferent and efferent neurons that are bundled together with connective tissue in to cord like fibers. They extend from the central nervous system to targets. |
|
|
Schwann cell |
Glial cell peripheral nervous system. Makes up myelin sheath of the peripheral nervous system |
|
|
Oligodendrocytes |
Glial cell central nervous system. Myelin sheath of central nervous system. |
|
|
Node of ranvier |
Spaces between schwann cells/oligodendrocytes on the axon. The area of the axon where voltage gated sodium ion channels are located. |
|
|
Enteric nervous system |
Located in the digestive system. Can be controlled by the autonomic division of the peripheral nervous system OR autonomously. |
|
|
Axonal transport |
Proteins that are synthesised in the neuron cell body are moved down the axon by axonal transport. This process can be FAST or SLOW. |
|
|
Slow axonal transport |
Moves material cytoplasmically. Used for components that are not consumed rapidly. E.g. enzymes. Move at a rate of 0.2mm per day. |
|
|
Fast axonal transport |
Move at a rate of 400mm per day. The neuron uses microtubules to transport vesicles and mitochondria down the axon with the help of motor proteins. Motor proteins bind and unbind to microtubules with the help of ATP. Transport moves in both directions. Anterograde - moving forward from cell body to axon terminal. Retrograde - moving from axon terminal to cell body - empty vesicles and debris for recycling. |
|
|
Presynaptic cells |
Delivers the signal to a target. |
|
|
Post-synaptic cells |
Recieve the signal from pre-synaptic cells. |
|
|
Synaptic cleft |
Space between two cells where neurotransmitters are secreted. The synaptic cleft is filled with extracellular matrix which holds the pre and post synaptic cells in place. |
|
|
Glial cells |
Supporting cells. Peripheral nervous system has 2 types: Schwann cells and satelite cells. Central nervous system has 4 types: oligodendrocytes, astrocytes, microglia and ependymal cells. |
|
|
Satellite cells |
Glial cells peripheral nervous system. Non-myelinating schwann cell. Forms supportive capsules around cell bodies located in ganglia. |
|
|
Ganglion |
Collection of nerve cell bodies found outside of the central nervous system. They appear as swellings or knots along the nerve. |
|
|
Astrocytes |
Glial cells central nervous system. Make up 50% of all brain cells. Provide neurons with substrates for ATP production. Maintain homestatis by monitoring potassium ion concentration and water. Surround blood vessels to form the blood-brain barrier to ensure regulated movement of materials between the blood and ECF in the brain. |
|
|
Microglia |
Glial cells central nervous system. Not made of neural tissue. Specialised immune cells which when activated protect the brain and spinal cord from damaged cells and foreign invaders.
|
|
|
Ependymal cells |
Glial cells central nervous system. Create the selectively permeable epithelial cells that separate the fluid compartments in the central nervous system. |
|
|
Excitable tissue |
Nerve and muscle cells are known as excitable tissues due to their ability to propegate electrical signals rapidly in response to stimulus. |
|
|
Sodium ion concentration |
145mM extracellular fluid 15 mM intracellular fluid |
|
|
Potassium ion concentration |
5mM extracellular fluid 150mM intracellular fluid |
|
|
Chloride ion concentration |
108mM extracellular fluid 10mM intracellular fluid |
|
|
Calcium ion concentration |
1mM extracellular fluid 0.0001 mM intracellular fluid |
|
|
Membrane potential |
Membrane potential is influenced by an electrochemical gradient of ions across a membrane and the membranes permeability to those ions. A resting membrane has a high permeability to potassium ions and a low permeability to sodium ions. Neurons have a resting membrane potential of -70mV |
|
|
Depolarisation and membrane potential |
If a membrane becomes more permeable to sodium ions, the sodium moves IN to the cell causing the cell membrane to have a more positive charge - therefore the cell is depolarized. |
|
|
Hyperpolarisation membrane potential |
If the membrane becomes more permeable to potassium ions, the potassium moves down its concentration gradient OUT of the cell causing the cell to become more negative. This is referred to as hyperpolarisation. |
|
|
Repolarisation membrane potential |
When the cell is returning to its resting membrane potential |
|
|
Channel activation |
The speed at which a voltage gated ion channel opens/closes to allow the flow of ions. Sodium ion voltage gated channels are RAPID potassium ion voltage gated channels are SLOWER. |
|
|
Graded potential |
Occur in the dendrites and cell body. Travel a short distance. Lose strength as they travel through the cell. If a graded potential is strong enough it will produce an action potential. |
|
|
Sub-threshold graded potentials |
Start above threshold (-55mV) at initiation point but decreases in strength as it moves through the cell. By the time it reaches the trigger zone the graded potential is below threshold and therefore no action potential is generated. |
|
|
Supra-threshold graded potential |
A stronger stimulus maintains strength above threshold as it moves down the cell body and once it reaches the trigger zone is able to trigger an action potential. |
|
|
Trigger zone |
Axon hillock. Integrating center of the neuron. Has a high concentration of sodium ion gated channels. |
|
|
Depolarizing action potentials |
Excitory |
|
|
Hyperpolarising action potentials |
Inhibitory |
|
|
Events of an action potential |
1. Depolarising stimulus reaches trigger zone. 2. Voltage gated sodium ion channels open causing sodium to rush in to the cell, causing more sodium ion gated channels to open etc (positive feedback loop). 3. Membrane depolarises - rising phase. 4. At peak voltage gated sodium ion channels begin to inactivate and the slower opening potassium ion channels begin to open. This stimulus terminates the positive feedback loop. 5. Opening of potassium ion channels allows potassium ions to leave the cell causing the cell to begin repolarising. 6. Potassium ion channels keep opening which creates an overshot and then hyperpolarisation. 7. When the potassium channels close the cell returns to its resting membrane potential. |
|
|
Depolarising at peak |
+30mV |
|
|
Hyperpolarising at peak |
-80mV |
|
|
Action potential summary |
The action potential is a change in membrane potential. It occurs when voltage gated ion channels in the membrane open, increasing the cells permeability, first to sodium ions (in) and then potassium ions (out). The influx of sodium ions depolarises the cell. This is followed by an efflux of potassium ions which first hyperpolarise the cell and then repolarises the cell to its resting membrane potential. |
|
|
Absolute refectory period |
The time when an action potential will NOT fire regardless of stimulus strength. Action potentials will not travel backwards and can not overlap. Voltage gated sodium ion channels take time to become active again after inactivation. During this time the membrane is hyperolarised and potassium is still leaving the cell. No action potential can occur. |
|
|
Relative refractory period |
Occurs after absolute refractory period. Some sodium ion gated channels are active again shower the cell is still hyperpolarised. As a result an extra strong stimulus (above threshold) is required to generate an action potential. |
|
|
Larger neurons |
Conduct faster action potentials. The larger diameter of the axon the more leak resistant the membrane is. E.g. the larger and wider the pipe the faster the water can flow. |
|
|
Myelination |
Increases conduction rate. Myelinated axons limit the amount of axon in contact with the ECF which reduces leak. Action potentials jump between nodes of ranvier = saltatory conduction |
|
|
Chemical synapse |
The electrical signal of a presynaptic cell is converted in to a neurocrine signal that crosses the synaptic cleft and binds to receptors on the target cell (post-synaptic cell). |
|
|
Synapse |
Each synapse has 2 parts. 1. The axon terminal of the pre-synaptic cell. 2. The membrane of the post synaptic cell. Most synapses are CHEMICAL. |
|
|
Electrical synapses |
Pass an electrical signal directly from the cytoplasm of one cell to another via gap junctions. Information flows in both directions. |
|
|
Neurotransmitters |
Create rapid responses. Released in vesicles (synaptic vesicles). Act as paracrine signals - target cells are close to the neuron secreting them. |
|
|
Neurohormone |
Neuron releases neurotransmitter into the bloodstream, acting as a hormone. |
|
|
Axon terminal membrane |
Calcium ion gated channels |
|
|
Neurotransmitter release |
Neurotransmitters in the axon terminal are stored in vesicles. They are released in to the synaptic cleft by exocytosis. This process occurs faster in neurons. |
|
|
Events at chemical synapse |
1. Action potential arrives at the axon terminal resulting in depolarisation of the membrane. 2. Depolarisation triggers calcium ion gated channels to open, allowing calcium ions to enter the cell. 3. Calcium ions entering the cell triggers the release of neurotransmitter vesicles. Vesicles move to the cell membrane where they exocytose in to the synaptic cleft. 4. Neurotransmitters diffuse across the synaptic cleft and bind with receptors on the post-synaptic cell. 5. Neurotransmitter binding results in a response in the post-synaptic cell. |
|
|
Neurotransmitter termination |
Neural signalling is a short process. Neurotransmitter action terminates when the chemicals are: 1. Broken down 2. Taken in to the post-synaptic cell or back in to the pre-synaptic cell. 3. Diffuse away in the ECF. If unbound transmitter is removed the synapse receptors release bound neurotransmitter which maintains equilibrium and terminates activity. |
|
|
Fast synaptic responses |
Fast synaptic responses are associated with the opening of ion gated channels. Ions move between the post-synaptic cell and ECF resulting in a change of membrane potential. Fast synaptic responses begin rapidly & last milliseconds. If the synaptic potential is depolarising it is EXCITORY. If the synaptic potential is hyperpolarised it is INHIBITORY. |
|
|
Excitory post-synaptic potential |
An excitory post-synaptic potential is more likely to result in an action potential in the post-synaptic cell. |
|
|
Inhibitory synaptic potential |
As a result of a hyperpolarisation potential of the post-synaptic cell. Considering INHIBITORY as it moves the membrane away from potential. |
|
|
Neurotransmitter receptors |
Ligand-gated ion channels = IONOTROPIC. G protein-coupled receptor = METABOTROPIC |
|
|
Ionotropic neurotransmitter receptors |
Ligand-gated ion channels. Rapid response. Ions move in/out of the cell creating graded potentials. Potentials can be excitory (depolarising) or inhibitory (hyperpolarising). |
|
|
Metabotropic neurotransmitter receptors |
G protein-coupled receptors. Slower responses. Use second messenger pathways to create response in the post-synaptic cell. Some receptors will result in the opening and closing of ion channels. Associated with metabolic processes. |
|
|
Divergence |
In a divergent pathway ONE pre-synaptic neuron branches to affect a large number of post-synaptic neurons. This pattern is similar to second messenger pathways. |
|
|
Convergence |
In a convergent pathway many pre-synaptic neurons provide input to influence a small number of post-synaptic neurons. |
|
|
Neurocrines |
Neurocrines function as neurotransmitters, neurohormones & neuromodulators. 7 classes. CNS releases MANY. PNS only releases 3: 1. Acetylcholine 2. Norepinephrine 3. Epinephrine. |
|
|
Acetylcholine (ACh) |
Neurons that secrete or bind to ACh are Cholinergic receptors. ACh is synthesised from choline and acetyl CoA at the synaptic terminal. Cholinergic receptors are either: Nicotinic - skeletal muscle, Autonomic division PNS and CNS. Opens ion channels>cell depolarises. Muscarinic - CNS & Parasympathetic PNS. 5 subtypes with various second messenger pathways. Metabotropic. . |
|
|
Epinephrine & norepinephrine |
Neurons that secrete or have receptors for epinephrine & norepinephrine are Adrenergic receptors. 2 classes alpha and beta with multiple sub-types of each. G protein-coupled receptors working with different second messenger pathways. Some result in excitory responses and others inhibitory responses. Derived from amino acid tyrosine.
Norepinephrine acts as a NEUROTRANSMITTER Epinephrine acts as a NEUROHORMONE. |
|
|
Temporal summation |

Occurs when 2 graded potentials from 1 pre-synaptic neuro noccur close together in time. If 2 sub-threshold graded potentials are too far apart in time they will not generate an action potential. If 2 sub-threshold graded potentials arrive at the trigger zone within a short period of time they may initiate an action potential. |
|
|
Spatial summation |

Spatial summation occurs when currents from nearly simultaneous graded potentials from a pre-synaptic neuron combine on a post-synaptic neuron. The summation of several sub-threshold graded potentials simultaneously results in an action potential. |
|
|
Pre-Synaptic Inhibition |

One inhibitory post-synaptic potential sums with two excitory post-synaptic potentials to prevent an action potential from occuring in the post synaptic cell. |
|
|
Global pre-synaptic inhibition |

Excitory and inhibitory pre-synaptic neurons fire but due to the inhibitory neuron, summed signal is below threshold and no action potential is generated. All targets are inhibited. |
|
|
Selective pre-synaptic inhibition |

An excitory neuron fires generating an action potential. An inhibitory neuron fires at the pre-synaptic axon terminal of one collateral which blocks the release of neurotransmitter release at one synapse. |
|
|
Pre-synaptic modulation |
A pre-synaptic neuron synapses on to an axon terminal of a neuron and alters the action potential teaching the pre-synaptic terminal. This alters the release of neurotransmitters from the axon terminal. |
|
|
Presynaptic inhibition |
Decreases the release of neurotransmitter at the synapse |
|
|
Pre-synaptic facilitation |
Increases neurotransmitter release from synapse. |
|
|
Neuromodulator |
Alters the responsiveness of a target cell to neurotransmitters. Neuromodulators can change the type & number of receptors as well as affinity of receptors for a neurotransmitter. |
|
|
Isoform |
Closely related proteins which function in a similar manner as ligands, however affinity differs. E.g. hemoglobin has multiple isoforms (adult & foetal) |
|
|
Protein activation |
Some proteins are inactive when they are synthesised in the cell, in order for the protein to be activated enzymes must remove a part of the molecule. Protein hormones & enzymes commonly undergo activation. |
|
|
Proteolytic activation |
A protein is inactive and requires an enzyme to activate it |
|
|
Co-factor activation |
When an ion or small functional group must bind to an active site before the ligand will bind and become active. Many enzymes do not function without co-factors. |
|
|
Protein modulation |
The ability of a protein to bind to a ligand & initiate a response can be altered by various factors, including: temp, pH and other molecular interactions. |
|
|
Protein modulator |
Can either change a proteins ability to bind to a ligand or changes the proteins ability to create a response. |
|
|
Chemical modulators |
Molecules that bind covalently or non-covalently to proteins and as a result alters their binding ability or activity. Chemical modulators can activate or enhance ligand binding, decrease binding or completely inactivate a protein. |
|
|
Protein inactivation |
Can be irreversible or reversible |
|
|
Antagonists |
Inhibitors. Chemical modulators that decrease activity. Often molecules that bind and block active sites. |
|
|
Competitive ligand inhibitors |
Reversible antagonists that compete with ligands for binding sites |
|
|
Irreversible ligand inhibitors |
Bind tightly to the active site and can not be displaced. |
|
|
Allosteric modulators |

Bind reversibly to a protein at a regulatory site which alters the protein binding site. Allosteric modulators can inhibit or activate. |
|
|
Covalent modulators |
Are atoms or functional groups that bind covalently to proteins = altered protein properties resulting in increased or decreased activity and binding ability. E.g. phosphorylation is the covalent bonding of a protein and a phosphate group. |
|
|
Physical factors & effect on ligand binding |
Temp & pH can have dramatic effects on protein structure and function. When these variables exceed a critical level they disrupt the bonds holding the protein in its tertiary structure. When this occurs the proteins shape is altered, altering the binding site which ultimately results in a loss of function |
|
|
Up-regulation |
The programmed production of new protein receptors, transporters and enzymes is called up-regulation. |
|
|
Down-regulation |
The programmed removal of proteins from the cell membrane is called down-regulation. Membrane receptors, transporters and enzymes are exocytosed. |
|
|
Conjugated proteins |
Protein molecules combined with another biomolecule |
|
|
Lipoprotein |
Lipid + protein |
|
|
Glycosated molecules |
Molecules where a carbohydrate has been attached |
|
|
Glycoprotein |
Carbohydrate + protein |
|
|
Glycolipid |
Carbohydrate + lipid |
|
|
Solution |
A homogenous mixture of 2 or more substances. Substances must be electrically attracted to water (polar) to dissolve. Like disolves like. |
|
|
pH of ECF |
Homeostatic range of 7.35-7.45 |
|
|
pH of ICF |
Homeostatic range of 7.0-7.2 |
|
|
Fluid mosaic model of biological membranes |
Describes the plasma membrane as a fluid combination of phospholipids, cholesterol & proteins. |
|
|
Cytoplasm |
Includes all material inside the cell membrane EXCEPT the nucleus. |
|
|
Cytosol |
Intracellular fluid. Semi gelatinous. Contains dissolved nutrients, proteins, ions and waste products. |
|
|
Cytoskeleton |
The internal support system of a cell. Consists of: Microvilli - increase S.A Microfillaments - form a network inside the cell membrane to support and maintain shape. Microtubules - largest cytoskeleton fiber. Hold organelles in place and used in mitosis. Intermediate filaments: myosin & keratin fibers. |
|
|
Mitochondria |
Double membrane - cristae Inner region - mitochondrial matrix matrix intermembrane space site of ATP production in the ETS. |
|
|
Golgi apparatus |
Series of hollow curved sacs (cisternae) stacked on top of each other. Surrounded by vesicles. Receives proteins from E.R on the cis face. Participates in post-translational modification of proteins. Packages modified proteins and transports out in vesicles via the trans face. |
|
|
Endoplasmic reticulum |
Rough - studded with ribosomes. Site of protein synthesis. Smooth - site of lipid synthesis and calcium stores. |
|
|
Nucleus |
Double membrane called the nuclear envelope. Pores within the membrane allow for communication with the cytoplasm. Outer membrane is continuous with the E.R. Nucleolus region contains the genes and proteins for synthesis of RNA & ribosomes. Nucleus houses chromatin (DNA & associated proteins). |
|
|
Lysosomes |
Digestive enzymes. Membrane bound vesicles. Digest bacteria, cellular waste and old organelles. |
|
|
Peroxisomes |
Smaller than lysosomes. Membrane bound vesicles. Generate hydrogen peroxide. Degrade fatty acids & toxins. Reactions inside the peroxisome convert hydrogen peroxide in to oxygen and water. |
|
|
Ribosomes |
Proteins. RNA structures. Float freely in cytosol or embedded in the rough E.R. Not membrane bound. Used in protein synthesis > translation |
|
|
Protein synthesis |
Conversion of DNA in to functional proteins. |
|
|
Codon |
Triplet of bases that code for a specific amino acid |
|
|
Gene |
Specific region of DNA that contains the information needed to make a functional piece of RNA. |
|
|
Protein synthesis: Gene activation |
Occurring in the nucleus of the cell. A regulatory protein known as a 'transcription factor' binds to the DNA and activates a promoter. The promoter then attracts the transcription enzyme RNA POLYMERASE where to bind to the DNA. The RNA polymerase moves along the DNA unwinding the double helix by breaking the hydrogen bonds. |
|
|
Protein synthesis: Transcription |
Occurs in the nucleus of the cell. One strand of unzipped DNA acts as a template strand. The promoter region is not transcribed. Each nitrogenous base in the DNA template is paired with its complimentary RNA base pair (G + C, T + A, A + U). This process is similar to the formation of the DNA double helix. As RNA bases bind to the DNA template they form a functional piece of mRNA. The RNA polymerase reaches a stop codon (UAG, UAA or UGA). No more bases are added and the mRNA strand is released. mRNA consists of nitrogenous bases, a sugar and a phosphate backbone.
Processing is the following step. |
|
|
Protein synthesis: Processing |
Occurs in the nuclus of the cell. mRNA is now inactivated/destroyed or it is processed by alternative splicing. Alternative splicing is the clipping of mRNA strands by enzymes. The mRNA is clipped into segments: Introns: non functional units which are discarded, Exons: functional segments which are put back together to form a shorter functional piece of mRNA. The mRNA can now leave the nucleus and head to the ribosomes for translation. |
|
|
Basic signal transduction |

|
|
|
Second messenger pathways |

|
|
|
Protein synthesis: Translation |
mRNA binds to ribosomes in the cytosol. Each ribosome has 2 sub-units. The small ribosomal sub-unit binds to the mRNA, the larger sub-unit then binds on top. The mRNA is sandwiched in the middle. This mRNA and ribosome complex is now ready for translation. During translation the mRNA codons are matched with an amino acid. This process occurs with the assistance of tRNA. tRNA contains anticodons which complement the codons on the mRNA. The mRNA codons attach to their complimentary tRNA anticodons, the correct pairing mRNA and tRNA codons puts amino acids in to their correct orientation which results in the formation of a peptide chain. Once the last amino acid has been added to the chain, it is released. The now empty tRNA is released where it can now attach to another amino acid molecule. The ribosomal sub-units separate ready for a new round of protein synthesis. The mRNA is broken down by enzymes. The peptide chain of amino acids are now sorted. The protein may have a sorting signal which means it will automatically move where it is needed> organelles or to the membrane to be exocytosed. Proteins without a signal remain in the cytosol. |
|
|
Protein synthesis: post translational modification |
The amino acid sequence that is produced during translation is the primary structure of a protein. The protein can now form different types of bonds - fold, spilt, bind to functional groups, form globular proteins. This process mainly occurs in the golgi. |
|
|
Respiration |
The process which animals extract energy from biomolecules. Animals consume oxygen and produce carbon dioxide and water as a bi-product. When more energy is ingested than required it is stored in chemical bonds. |
|
|
Main energy source for humans |
Glucose and lipids Triglycerides |
|
|
Energy |
The capacity to do work. Chemical Transport Mechanical |
|
|
Mechanical work |
Movement at a cellular level - Cells changing shape. Movement at macroscopic level - muscle contraction |
|
|
Chemical work |
Making and breaking of chemical bonds |
|
|
Transport work |
Movement of ions, molecules and large particles through the cell membrane and through the membranes of organelles. |
|
|
Kinetic energy |
The energy of motion. Molecules moving down a concentration gradient produce kinetic energy. |
|
|
Potential energy |
Stored energy. Potential energy is stored chemical bonds - high energy electrons. |
|
|
Activation energy |
The energy required to initiate a chemical reaction |
|
|
Enzymes |
Biological catalysts Lower activation energy Enzymes are NOT used in the chemical reaction. Most enzymes are proteins. |
|
|
exergonic reactions |
ENERGY IS RELEASED. The products of the reaction have LESS stored energy than the reactants. Give off heat |
|
|
Endergonic reactions |
Energy is stored. Products have MORE energy than the reactants. Synthesis reactions - complex molecules made from smaller molecules. |
|
|
Coupling endergonic and exergonic reactions |
Energy released exergonically can be trapped and stored endergonically. |
|
|
ATP |
Adenosine triphosphate Energy transfer molecule. Energy trapped in phosphate groups. When the bond is broken, energy is released and ATP forms ADP & phosphate. ATP is composed of Nucleotide AMP and 3 phosphate groups. |
|
|
FAD & NADH |
Molecules that trap high energy electrons |
|
|
Metabolism |
The sum of all chemical and physical processes that take place in the body |
|
|
Catabolism and anabolism |
Catabolism - reactions release energy via the breakdown of LARGE BIOMOLECULES > energy OUT Anabolism - reactions build large molecules using energy (ATP) > energy IN. Catabolic reactions drive anabolic reactions. |
|
|
Glycolosis |
Aerobic pathway to ATP production. Occurs in the cytosol. 1 x 6 carbon glucose is phosphorylated to form glucose-6-phosphate. A series of enzymatic reactions occur Resulting in = 2 x 3 carbon pyruvate molecules and 2 x ATP. Glycolsis does NOT require oxygen. |
|
|
Citric acid cycle. |
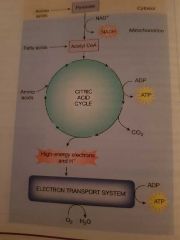
Step 1. If a cell has adequate oxygen the 2 x 3 carbon pyruvate molecules formed during glycolosis move in to the mitochondrial matrix where they react with coenzyme A (CoA) to form ACETYL CoA, 1 x NADH and 1 carbon dioxide.
Step 2. The 2 carbon acyl unit now enters the citric acid cycle. It combines with a 4 carbon oxaloacerate, this process results in a series of enzymatic reactions forming a continuous cycle. With each turn of the cycle produces 2 ATP, 3 x NADH, 1 x FADH and carbon dioxide.
These steps occur twice in every cycle producing a net yield of 8 x NADH, 2 x FADH, 2 ATP and 6 carbon dioxide |
|
|
Electron transport system |
Final step of aerobic ATP production. The energy transfer from high energy electrons to ATP. Requires the electron transport system which is located in the space between the 2 mitochondrial membranes. The process of ATP synthesis is called oxiative phosphorylation. NADH & FADH release their high energy electrons and hydrogen ions to the electron transport system. The ETS is a series of proteins which span the inner mitochondrial membrane. The high energy electrons move along the proteins and their energy is released kinetically. This energy is used to move hydrogen ions against their concentration gradient OUT of the mitochondrial matrix into the intermembrane space. The hydrogen ion concentration gradient is a source of potential energy. Each pair of electrons released by the ETS binds to 2 hydrogen ions and an oxygen to form water. The potential energy formed in the hydrogen ion concentration gradient is released when the hydrogen ions flow back down their gradient back in to the mitochondrial matrix through a protein channel called ATP SYNTHASE. As the hydrogen move in ATP SYNTHASE harvests the released kinetic energy and synthesises the ATP molecule. Energy is transfered to the phosphate bond. A small portion of this energy is lost as heat. For every 3 hydrogen ions that pass through ATP SYNTHASE one ATP is created. Net yield = 30-32 ATP 6 water molecules 6 carbon dioxide. |
|
|
Aerobic metabolism net equation |

|
|
|
Beta oxidation |
The process of fatty acids being catabolised in the mitochondria to be converted to acetyl CoA which can then enter the citric acid cycle. |
|
|
Fatty acids can only be used for energy aerobically |
When energy is required large molecules such as glycogen can be broken down in to components for glycolosis etc.. Proteins can be broken down in to amino acids Lipids can be broken down into glycerol and fatty acids. Glycerol feeds into glycolosis and fatty acids are used by acetyl CoA in the citric acid cycle. MORE ENERGY IS STORED IN FATS THAN CARBOHYDRATES AND PROTEINS. |
|
|
Membrane dynamics |
Water is the only molecule that can move freely between cells and the ECF. Potassium ions can LEAK out and Sodium ions can LEAK in but in order for them to return to their original compartments energy is required in the form of ATP. Homeostatic mechanisms maintain a relatively stable condition in the ECF - continual input of energy is required to maintain solute concentrations. |
|
|
Osmotic equilibrim |
Free movement of water between ICF and ECF compartments. |
|
|
Intracellular fluid |
2/3 of the bodies water volume. Slight negative charge due to large negatively charged anions. |
|
|
Extracellular fluid |
1/3 of bodies water volume
Slight positive charge. |
|
|
Osmosis |
The movement of water across a semi permeable membrane in response to solute concentrations. In osmosis water moves from it's area of high concentration to an area of low concentration. This relates to solute concentrations as an area with HIGH amounts of non-penetrating particles has a low free water concentration. An area with LOW amounts of non-penetrating particles has a high concentration of free water. When these solutions are separated by a semi permeable membrane the free water in the solution with low solute concentration will pass across the membrane DOWN it's concentration gradient in to the solution with high non-penetrating particles (low free water). The water will dilute the solution until osmotic equilibrium has been reached. been reached. |
|
|
Diffusion |
Net movement of molecules from an area of high concentration to an area of low concentration. |
|
|
Simple diffusion |
Passive diffusion. Molecules diffuse across the cell membrane directly. In order to do this they must be SMALL, NON-POLAR AND LIPID SOLUBLE. |
|
|
Facilitated diffusion |
Molecules diffuse across the cell membrane via transporters and channels. |
|
|
Active transport |
Requires energy input. It is the movement of substances AGAINST their concentration gradients. |
|
|
Primary active transport |
Uses ATP. e.g. sodium potassium pump. |
|
|
Secondary active transport |
Uses the kinetic energy of one molecule moving down its concentration gradient to transport another molecule against its concentration gradient. |
|
|
Omolarity |
The number of particles in a solution. |
|
|
Osmotic pressure |
The force required to prevent movement of water across a membrane. The higher the osmotic pressure the more forcefully the water is pulled across. Larger concentration gradients produce higher osmotic pressure. |
|
|
Tonicity |
Solute concentration and it's effect on water movement in and out of the cell. |
|
|
Isotonic |
Solution has the same amount of non-penetrating particles in and out of the cell. Water moves freely back and forth with no net gain or loss |
|
|
Hypotonic |
Solution has a lower concentration of particles OUTSIDE Which results in a high free water concentration outside of the cell. Water rushes down its concentration gradient IN to the cell where the free water is low. This causes the cell to swell. |
|
|
Hypertonic |
The solution has a high concentration of non-penetrating particles outside of the cell and low free water. The water rushes down its concentration gradient out of the cell causing the cell to shrink. |
|
|
Channel proteins |
Create water filled passageways that link the ICF and ECF compartments. Movement through these channels is restricted to water and ions. They are named according to the substance that can pass through. E.g. sodium ion channels. Channels are selective. This is determined by its size and electrical charge of the ions. Channels can be: Open - leak channels Gated - open in response to signals. Chemical - ligand binding Voltage - change in membrane potential Mechanical - physical forces |
|
|
Carrier proteins TRANSPORTERS |
Transporters bind with particular substrates and carry them across the membrane by changing formation. Carries small organic molecules such as glucose and amino acids.
Uniport carriers - 1 type of molecule
Symport carriers - 2 molecules in the same direction
Antiport carriers - 2 molecules in opposite directions. Carrier proteins exhibit: Specificity - bind to specific molecules Competition - substrates compete for binding sites. Competitive inhibitors can slow transport. Saturation - when all active sites are full and working at capacity. Cell may UP-REGULATE. |
|
|
Vesicular transport |
Transport molecules that are too large for channels and carriers. Use ATP. Phagocytosis - membrane bound vesicle engulfs bacteria and other particles. The vesicle fuses with lysosomes where digestive enzymes destroy the contents. Pinocytosis - membrane bound vesicle full of fluid. Exocytosis - movement of ICF and substances out of the cell in vesicles. |
|
|
Epithelial transport |
Transport of substances entering or leaving the body and between compartments must cross a layer of epithelial cells. Cells are joined by tight junctions. The surface of the cell facing the lumen = apical surface. Surface facing ECF is the basolateral membrane. Transport from the lumen of an organ to the ECF is ABSORBTION. Transport of material from the ECF in to the lumen (or release of substances from a cell) SECRETION. Paracellular transport - transport between adjacent epithelial cells through leaky tight junctions. Transcellular transport - substances cross both apical and basolateral membranes using a combination of active and passive transport. |

