![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
237 Cards in this Set
- Front
- Back
|
What is the major site for ATP production in the eukaryotic cell?
|
mitochondria
|
|
|
Where does transcription occur in a eukaryotic cell?
|
nucleus
|
|
|
Which features of eukaryotic cells are believed to have originated from the engulfing of a prokaryotic cell?
|
mitochondria and chloroplasts
|
|
|
Which of the following IS NOT found in the nucleus?
A) DNA B) RNA C) Proteins D) Nucleotides E) All are found in the nucleus. |
All are found in the nucleus
|
|
|
Which is NOT a function of a eukaryotic cell membrane (the membrane which surrounds the cell?
A) Transport of wastes and nutrients B) Maintain gradients C) Cell Motility D) Cell-Cell Communication E) Production of ATP energy |
Production of ATP energy
|
|
|
Which of the following is NOT true about prokaryotes?
A) One type is bacteria B) One type is archaea C) Some can do photosynthesis D) One type is yeast E) Some can use oxygen to oxidize food molecules |
One type is yeast
|
|
|
The Golgi _____.
A) Is involved in sorting proteins to various membrane-bound organelles B) Receives proteins from the rough ER C) Packages proteins that are to be secreted from the cell D) Is a stack of membrane-bound flattened sacs E) All of the above |
All of the above
|
|
|
The major function of the rough endoplasmic reticulum is _____.
A) Generation of energy B) Synthesis of cytosolic proteins C) Cell motility D) Generation of phospholipids E) Synthesis of secreted and membrane-associated proteins |
Synthesis of secreted and membrane-associated proteins
|
|
|
Which of the following does NOT pertain to the cytoskeleton?
A) Contains actin filaments, microtubules and intermediate filaments B) Provides structure to the cytoplasm of the cell C) It gives shape and strength to the cell D) It is involved in cell movement. E) It helps generate ATP in the cell |
It helps generate ATP in the cell
|
|
|
The term "compartmentalization" refers to:
A) The presence of membrane-bound organelles in eukaryotic cells B) The ability of eukaryotic cells to secrete proteins C) The ability of eukaryotic cells to make contact with each other D) The fact that eukaryotic cells produce energy E) The ability of a eukaryotic cell to contain a prokaryotic cell |
The presence of membrane-bound organelles in eukaryotic cells
|
|
|
The four groups that amino acids are classified into are _____.
|
basic
acidic uncharged polar nonpolar |
|
|
The THREE amino acids that carry positive charges at neutral pH are _____.
|
lysine
arginine histidine |
|
|
Which of the following short proteins chains (or peptides) would be considered the most HYDROPHOBIC?
a) lysine-arginine-histidine-glycine-lysine b) glutamic acid-glycine-aspartic acid-leucine-glutamic acid-aspartic acid c) proline-leucine-valine-proline-alanine-methionine d) cysteine-methionine-cysteine-valine-glutamic acid e) None of the above. |
proline-leucine-valine-proline-alanine-methionine
|
|
|
Which of the following amino acids would likely be found on the SURFACE of a folded protein?
a) Glutamic Acid b) Proline c) Glycine d) Leucine e) Alanine |
Glutamic acid
|
|
|
An acidic, basic, or uncharged polar amino acid would likely be found here in a folded protein.
|
On the surface
|
|
|
The nonpolar amino acids are would be found here in the folded protein.
|
In the inside of the folded protein, away from the aqueous cytoplasm.
|
|
|
The nonpolar amino acids are found in the inside of the folded protein, because they are:
|
hydrophobic and are kept away from the aqueous cytoplasm
|
|
|
Which of the following amino acids contain a sulfur atom?
a) Tryptophan b) Cysteine c) Methionine d) Arginine e) more than one of the above |
more than one of the above
|
|
|
What would be the NET charge of the following peptide in a neutral buffer (pH = 7.4)? ARG-LYS-ARG-GLU-LYS-LYS-GLY-VAL.
a) +4 b) −4 c) +2 d) −2 e) 0 |
+4
|
|
|
If a protein contained the sequence: glutamic acid-glycine-aspartic acid–leucine-glutamic acid-aspartic acid, which of the amino acid chains below would it most likely form ionic bonds with?
a) lysine-arginine-histidine-glycine-lysine b) glutamic acid-glycine-aspartic acid-leucine-glutamic acid-aspartic acid c) proline-leucine-valine-proline-alanine-methionine d) cysteine-methionine-cysteine-valine-glutamic acid e) None of the above |
lysine-arginine-histidine-glycine-lysine
|
|
|
According to lecture, the macromolecules found in a cell are all _____.
|
polymers
|
|
|
Phospholipid molecules in water might be expected to stick to one another by virtue of _____ interactions occurring between their tails.
|
hydrophobic
|
|
|
Remember that hydrogen atoms forming nonpolar covalent bonds (for example, with C) are NOT ideal for:
|
hydrogen bonding
|
|
|
Remember that hydrogen atoms forming _______ (for example, with C) are NOT ideal for hydrogen bonding.
|
nonpolar covalent bonds
|
|
|
Which of the following can be used by cells to produce ATP energy?
a) sugars b) fatty acids c) amino acids d) sugars and fatty acids e) sugars, fatty acids, and amino acids |
All of them
|
|
|
How many elements are found in nature?
|
92
|
|
|
These elements make up 98% of all elements in a cell
|
C, H, O
|
|
|
What is the most abundant substance in cells?
|
Water, makes up 70% of the cell
|
|
|
Hydrophilic molecules have a _____ charge.
|
partial or full plus or minus charge. they are polar
|
|
|
Hydrophilic molecules are polar or non-polar?
|
polar
|
|
|
Hydrogen bonding is formed by hydrophilic or hydrophobic molecules?
|
hydrophilic
|
|
|
Hydrophobic molecules have a _____ charge.
|
neutral. they are non-polar
|
|
|
Hydrophilic molecules form hydrogen bonds: T/F
|
T
|
|
|
Hydrophobic molecules form hydrogen bonds: T/F
|
F
|
|
|
These increase the level of H+
|
acids
|
|
|
These decrease the level of H+
|
bases
|
|
|
Acids increase the level of ___
|
H+
|
|
|
Bases decrease the level of ___
|
H+
|
|
|
Bases decrease the level of H+ by:
|
direct: acquriing H+
indirect: release of OH- |
|
|
This is an important cellular base which forms NH3+ and OH-
|
NH2
|
|
|
Cells contain these 4 major building blocks of small organic molecules which makeup these larger units:
|
Monosaccharides --> Polysaccharides
Fatty Acids --> Fats, Lipids, Membranes Amino Acids --> Proteins Nucleotides --> Nucleic Acids |
|
|
The four building blocks of the cell have different fates inside the cells; the overall distribution is toward formation of:
|
large macromolecules
|
|
|
The four building blocks of the cell have different fates inside the cells; the overall distribution is toward formation of large macromolecules. This is known as:
|
polymer chemistry
|
|
|
The formula for monosaccharides is:
|
(CH2O)n
n= 3-7 |
|
|
Monosaccharides form polysaccharides by covalent linkages through:
|
-OH
|
|
|
Monosaccharides undergo this type of reaction to form polysaccharides:
|
condensation (dehydration)
(water expelled) |
|
|
Polysaccharides undergo this type of reaction to form monosaccharides:
|
hydrolysis
(water consumed) |
|
|
These are the functions of sugars:
|
-production of storage and energy
-cell walls (cellulose) -extracellular matrix -DNA/RNA -linked to protiens |
|
|
Fatty acids have two chemically distinct regions:
|
long hydrocarbon chain - hydrophobic
carboxyl head group - hydrophilic |
|
|
Three fatty acid chains linked to glycerol is known as:
|
Triacylglycerols
|
|
|
Two fatty acid chains linked to glycerol/PO4/head group is known as:
|
Phospholipids
|
|
|
Triacylglycerols are made up of:
|
3 fatty acid chains linked to glycerol
|
|
|
Phospholipids are made up of:
|
2 fatty acid chains linked to glycerol/PO4/head group
|
|
|
Fatty acids are stored as ________ through an ester linkage to glycerol to form triacylglycerols.
|
an energy reserve
|
|
|
Phospholipids are the major constituents of:
|
cell membranes
|
|
|
These are the functions of fats:
|
-production and storage of energy
-cell membranes -cell signaling (steroids, signal transduction) |
|
|
Fats provide _______ than sugars by weight.
|
2x more energy
|
|
|
Monosaccharides make covalent linkages through -OH groups to form:
|
polysaccharides
|
|
|
Amino acids are made up of:
|
-Carboxylic acid group
-Amino group -Side chain (R) -Alpha Carbon |
|
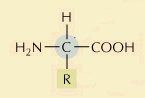
What is this molecule?
|
Amino Acid
|
|
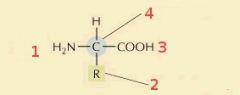
Name these parts of the amino acid:
|
1) Amino group
2) Side-chain group 3) Carboxyl group 4) Alpha Carbon |
|
|
Both the amino and carboxyl groups are ionized in an amino acid at this pH:
|
7
|
|
|
At pH 7 both the amino and carboxyl groups of an amino acid are:
|
ionized
|
|
|
At pH 7, an amino acid will ionize leaving the molecule charged in this way:
|
amino group becomes NH3+
carboxyl group becomes COO- |
|
|
How does an amino acid look at pH 7?
|
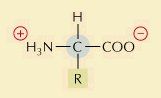
|
|
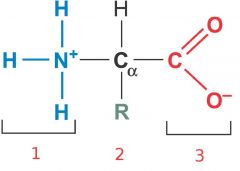
Name the parts of the amino acid.
|
1) Amino group
2) Side chain 3) Carboxyl group |
|
|
There are ____ distinct amino acids
|
20
|
|
|
This part of the amino acid confers its functionality (nonpolar, uncharged-polar, acidic, basic)
|
Side chain
|
|
|
Only this form of amino acid is used:
|
L form
|
|
|
Amino acids are linked via:
|
peptide bonds
|
|
|
These are the functions of proteins:
|
Generation of proteins
Production of energy |
|
|
Nucleotides are made up of:
|
-Nitrogen rings (pyrimidines, purines)
-Sugar (ribose or deoxyribose) -Phosphate group (1, 2 or 3) |
|
|
These are the types of Nitrogen Rings:
|
Purines (A, G)
Pyrimidines (T, U, C) |
|
|
These are the functions of nucleotides:
|
DNA (A, G, C, T)
RNA (A, G, C, U) Carriers of chemical energy (mainly ATP) Signal molecules Coenzymes |
|
|
All macromolecules are made with the same basic chemical reaction:
|
dehydration (condensation)
|
|
|
For a protein of 200 amino acids, there is this many possible sequences:
|
20^200
|
|
|
This specifies the shape of a macromolecule:
|
noncovalent bonds
|
|
|
These types of bonding specify the shape of a macromolecule:
|
Van der Waals
Hydrogen Hydrophobic Ionic |
|
|
At very short distances, any two atoms show a weak interaction due to fluctuations in charges. This is known as:
|
Van der Waals Forces
|
|
|
Describe Van der Waals forces:
|
At very short distances, any two atoms show a weak interaction due to fluctuations in charges
|
|
|
A bond where a hydrogen is sandwiched between two electron-attracting ions. This is known as:
|
Hydrogen bond
|
|
|
Describe hydrogen bonding:
|
A bond where a hydrogen is sandwiched between two electron-attracting ions
|
|
|
Hydrogen bonding is ____ the strength of a covalent bond.
|
1/20th
|
|
|
Hydrophobic groups are forced together to minimize their disruption of hydrogen bonds in water. This is known as:
|
Hydrophobic interactions
|
|
|
Describe Hydrophobic interaction:
|
Hydrophobic groups are forced together to minimize their disruption of hydrogen bonds in water
|
|
|
The transfer of electrons to create a bond occurring between fully charged or partially charged groups.
|
Ionic bonding
|
|
|
When considered individually, _____ are very weak, but multiple bonds result in a very strong interaction, between different parts of the molecule.
|
non-covalent bonds
|
|
|
_____ bonds allow intermolecular interactions.
|
Non-covalent
|
|
|
_____ bonds allow the assembly of multiple macromolecules into a single structure.
|
Non-covalent
|
|
|
The assembly of multiple macromolecules into a single structure through non-covalent bonds is called:
|
macromolecular assembly
|
|
|
Macromolecules are made by these kinds of bonds:
|
covalent
|
|
|
3-D structure of macromolecules is formed through:
|
non-covalent bonds
|
|
|
The pH of a cell is:
|
7.4
|
|
|
Hydrogen bonds are the strongest when:
|
the three atoms involved are in a straight line
|
|
|
The second law of thermodynamics states that:
|
In any system the degree of disorder (entropy) will increase over time.
|
|
|
How do cells generate order?
|
-There is a constant input of energy into the system.
-Cells release heat to their surroundings, increasing entropy in the environment. |
|
|
Cells release ____ to their surroundings, _____ entropy in the environment and _____ entropy in the cell.
|
heat
increasing decreasing |
|
|
The integrated network of chemical reactions that regulate life:
|
metabolism
|
|
|
Metabolism is:
|
the integrated network of chemical reactions that regulate life
|
|
|
This part of metabolism involves the breakdown of molecules and typically releases energy
|
catabolism
|
|
|
This part of metabolism involves the building or biosynthesis of molecules and typically requires energy
|
anabolism
|
|
|
An initial compound in a reaction which binds to an enzyme is known as:
|
substrate
|
|
|
A compound generated from the substrate which will likely become substrate itself for a new reaction.
|
product
|
|
|
Intrinsic energy within a compound, expressed in terms of calories (kcal)
|
Free Energy (G)
|
|
|
Free Energy is defined as:
|
intrinsic energy within a compound, expressed in terms of calories (kcal)
|
|
|
If ∆G is negative, the reaction will:
|
proceed spontaneously
A --> B |
|
|
If ∆G is positive, the reaction will:
|
not be spontaneous
A <-- B |
|
|
If ∆G is zero, the reaction will:
|
be at equilibrium
A <==> B |
|
|
The amount of energy generated by
A <==> B is equal to: |
∆G = G_B - G_A
|
|
|
This means that the number of A molecules being converted to B is the same as B to A
|
equilibrium
|
|
|
Equilibrium means that there is the same number of A molecules and B molecules. T/F
|
F
|
|
|
Equilibrium is defined as:
|
A reaction that the number of A molecules being converted to B is the same as B to A
|
|
|
This is independent of the pathway of the reaction, and tells us nothing about the rate.
|
∆G
|
|
|
∆G is independent of the pathway of the reaction and tells us nothing about the rate. T/F
|
T
|
|
|
This can be used to predict the direction a reaction goes to
|
∆G
|
|
|
This can be used to find how much energy will be liberated or used in a reaction:
|
∆G
|
|
|
∆G can be used to predict:
|
The direction a reaction goes (spontaneous or not)
How much energy will be liberated or used. |
|
|
∆G° + 0.616 kcal/mol • ln [B]/[A] = ?
|
∆G
|
|
|
∆G° represents:
∆G represents: |
standard free energy change (in test tube)
change in free energy (in cell) |
|
|
We can relate ∆G of biological systems to the relative SUBSTRATE and PRODUCT concentrations by doing this:
|
Mix equal amounts of A and B in a test tube under STANDARD STATE CONDITIONS, and let it go to equilibrium.
Then, evaluate the concentrations of A and B. |
|
|
K_eq is defined as:
|
the equilibrium constant, the value of [B]/[A] at equilbrium
[B] = concentration of product [A] = concentration of substrate |
|
|
Enzymes provide:
|
catalytic power
|
|
|
Enzymes accelerate reactions by ______ times or more
|
1,000,000
|
|
|
Enzymes decrease the ______ required for reactions to occur.
|
activation energy
|
|
|
Enzymes decrease the activation energy by:
|
bringing the substrate into a favorable conformation
|
|
|
Enzymes do not affect the equilibrium, they accelerate the attainment of equilibrium. T/F
|
T
|
|
|
∆Go is known as:
|
the free energy under standard state in the test tube
|
|
|
Define the purpose, approach, equation, and variables with regards to free energy in the test tube.
|
Purpose: to solve for G°
Approach: reaction GOES TO equilibrium Equation: ∆G° = −0.616 kcal/mol (ln Keq) Variables: ∆G° and Keq |
|
|
Define the purpose, approach, equation, and variables with regards to free energy in the cell.
|
Purpose: to solve for G
Approach: reaction goes TOWARD equilibrium Equation: ∆G = ∆G° + 0.616 kcal/mol • ln [B]/[A] Variables: ∆G, ∆G°, and [B]/[A] (not Keq) |
|
|
In biological systems a chemical reaction that is not at equilibrium is capable of providing energy as it attains equilibrium state. This is known as:
|
coupling
|
|
|
Coupling is defined as:
|
In biological systems a chemical reaction that is not at equilibrium is capable of providing energy as it attains equilibrium state.
|
|
|
We collect free energy (∆G) in two basic forms:
|
electrons (e−)
High energy phosphate (–PO4 2−) |
|
|
The overall free energy change for chemically coupled reactions (serial or independent rxns) is equal to:
|
the SUM of the free energy changes of the individual steps.
∆Gs are ADDITIVE! |
|
|
How do reactions proceed in a timeframe that allows organisms to thrive?
|
enzymes
|
|
|
How do we collect the free energy from chemical reactions?
|
oxidation
|
|
|
This is an example of what kind of reaction?
C6H12O6 + O2 → H2O + CO2 |
oxidation
|
|
|
The oxidation of these fuel molecules generates the free energy to drive reactions:
|
glucose, fatty acids, amino acids
|
|
|
The oxidation of glucose, fatty acids, and amino acids generates:
|
the free energy to drive reactions
|
|
|
Glucose, fatty acids, and amino acids undergo this type of reaction to generate the free energy to drive reactions:
|
oxidation
|
|
|
Oxidation is the elimination of:
|
H- (electrons)
H+ |
|
|
The elimination of electrons is known as:
|
oxidation
|
|
|
Free energy is represented by:
|
chemical bonds (electron pairs)
|
|
|
These are the 'currency' of oxidation:
|
electrons
|
|
|
These serve as electron carriers:
|
NAD+
FAD NADP+ |
|
|
These transfer electrons to the oxidative phosphorylation enzymes in the mitochondria:
|
NAD+
FAD |
|
|
NAD+ and FAD transfer electrons to the:
|
oxidative phosphorylation enzymes in the mitochondria
|
|
|
The energy in the electrons is used to produce a H+ gradient, so that:
|
ATP can be synthesized
|
|
|
The energy in the electrons is used to produce a ______, so that ATP can be synthesized
|
H+ gradient
|
|
|
These amino acids have basic side chains:
|
lysine
arginine histidine |
|
|
These amino acids have acidic side chains:
|
aspartic acid
glutamic acid |
|
|
Amino acids are commonly joined together by an amide linkage, called a:
|
peptide bond
|
|
|
These amino acids have uncharged polar side chains:
|
asparagine
glutamine serine threonine tyrosine |
|
|
These amino acids have nonpolar side chains:
|
alanine
valine leucine isoleucine proline phenylalanine methionine tryptophan glycine cysteine |
|
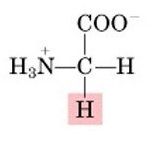
Name this amino acid.
|
glycine
|
|
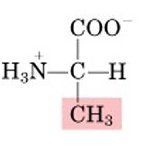
Name this amino acid.
|
alanine
|
|
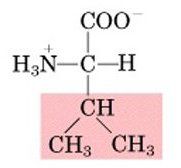
Name this amino acid.
|
valine
|
|
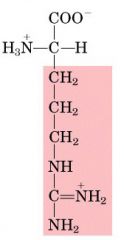
Name this amino acid.
|
arginine
|
|
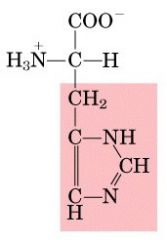
Name this amino acid.
|
histidine
|
|
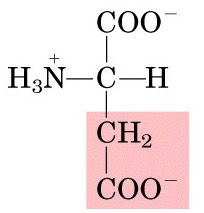
Name this amino acid.
|
aspartate
|
|
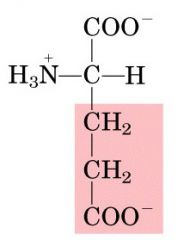
Name this amino acid.
|
glutamate
|
|
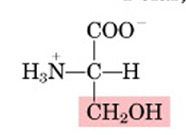
Name this amino acid.
|
serine
|
|
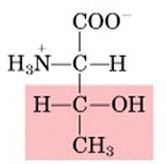
Name this amino acid.
|
threonine
|
|
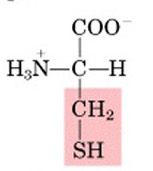
Name this amino acid.
|
cysteine
|
|
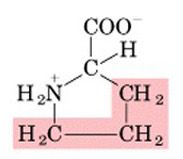
Name this amino acid.
|
proline
|
|
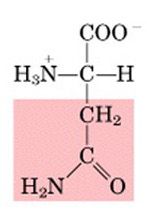
Name this amino acid.
|
asparagine
|
|
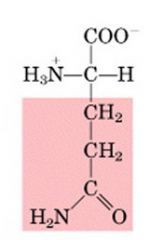
Name this amino acid.
|
glutamine
|
|
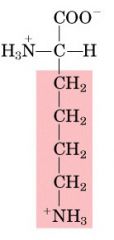
Name this amino acid.
|
lysine
|
|
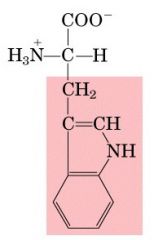
Name this amino acid.
|
tryptophan
|
|
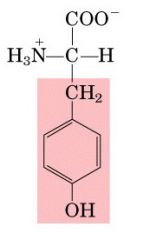
Name this amino acid.
|
tyrosine
|
|
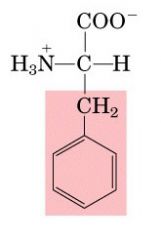
Name this amino acid.
|
phenylalanine
|
|
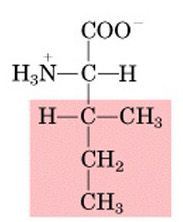
Name this amino acid.
|
isoleucine
|
|
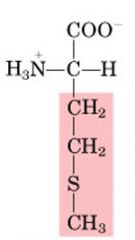
Name this amino acid.
|
methionine
|
|
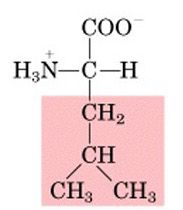
Name this amino acid.
|
leucine
|
|
|
Only this form of amino acid is found in proteins:
|
L-form
|
|
|
Only the D form of amino acids are found in proteins: T/F
|
F
|
|
|
Only the L form of amino acids are found in proteins: T/F
|
T
|
|
|
This is the major electron donor in reductive biosynthesis and photosynthesis:
|
NADPH
|
|
|
The free energy of electrons is used to generate:
|
ATP
|
|
|
The universal currency of free energy in biological systems:
|
ATP
|
|
|
What is the basis of ATP's free energy?
|
Electrostatic repulsion between the negative charges of −PO4 2−
Resonance stabilization |
|
|
The hydrolysis of ATP changes the ____ by a very large number, allowing thermodynamically unfavorable reactions to occur.
|
Keq
|
|
|
The hydrolysis of _____ changes the Keq by a very large number, allowing thermodynamically unfavorable reactions to occur.
|
ATP
|
|
|
The hydrolysis of ATP changes the Keq by a very large number, allowing:
|
thermodynamically unfavorable reactions to occur.
|
|
|
List some important compounds with high phosphoryl transfer potential:
|
Phosphenolpyruvate (PEP) ∆Go −14.3
Creatine Phosphate ∆Go −10.3 ATP ∆Go −7.3 Glucose-1-phosphate ∆Go −5.3 Glucose-6-phosphate ∆Go −3.3 |
|
|
Small molecules which bring unique chemical functionality to certain enzyme reactions.
|
coenzyme
|
|
|
This is the universal carrier of acyl groups:
|
Coenzyme A (CoA)
|
|
|
This is the reactive site of Acetyl Coenzyme A:
|
terminal sulfhydral group
|
|
|
The reduced form of NADP+ is:
|
NADPH
|
|
|
The oxidized form of NADPH is:
|
NADP+
|
|
|
The primary structure of a protein consists of:
|
the amino acid sequence
|
|
|
Once this is established, the protein will FOLD into a single conformation with the lowest energy state.
|
primary structure
|
|
|
Once a primary structure is established, the protein will FOLD into a single conformation with the lowest energy state. This is mediated by:
|
non-covalent bonds
|
|
|
Once a primary structure is established, the protein will FOLD into a single conformation with:
|
the lowest energy state.
|
|
|
The secondary structure of a protein consists of:
|
STRETCHES of amino acids that fold into functional structures, based on the sequence of amino acids (primary structure)
|
|
|
The secondary structure of a protein consists of stretches of amino acids that fold into functional structures, based on:
|
the sequence of amino acids (primary structure)
|
|
|
A protein's secondary structure can have these formations:
|
Alpha Helix
SINGLE REGION of amino acids Turns every ~ 4 amino acids Beta Sheet MULTIPLE REGIONS Antiparallel OR Parallel |
|
|
The non-covalent bonds involved in the secondary structure of a protein are:
|
hydrogen bonds
|
|
|
In the secondary structure of a protein, alpha helices and beta sheets arise through:
|
H-bonds between N-H and C=O groups in peptide backbone.
|
|
|
Side chains of amino acids are involved in the hydrogen bonds which form alpha helices and beta sheets in the secondary structure of proteins. T/F
|
F
|
|
|
Protein secondary structure helices are important because they:
|
Form membrane-spanning regions
Coiled-coils Provide functional domains to many proteins |
|
|
Any part of a polypeptide chain that can fold independently into a compact, stable structure:
|
functional domain
|
|
|
These are examples of protein functionality:
|
Protein-Protein Interaction
Protein-DNA Interaction Substrate-Binding Site |
|
|
The tertiary structure of a protein consists of:
|
The 3-D structure of the functional protein molecule (Conformation)
Involves all non-covalent bonding (ionic, hydrogen, van der Walls, hydrophobic) |
|
|
Each protein normally folds into a single conformation with the lowest energy state but changes in shape can be caused by:
|
binding of additional molecules
environmental changes phosphorylation other modifications |
|
|
Define Allosteric Protein
|
a protein that has multiple conformations
|
|
|
A protein that has multiple conformations. Generally, one form is active and the other form is inactive
|
allosteric protein
|
|
|
Many functional proteins often have more than one polypeptide chain. Each of these are called:
|
subunit
|
|
|
The quaternary structure of a protein consists of:
|
A combination of MORE THAN ONE
independent polypeptide chains Involves all non-covalent bonding detailed previously. (ionic, hydrogen, van der Walls, hydrophobic). |
|
|
Individual protein subunits can make these formations:
|
filaments
sheets spheres |
|
|
The majority of bonds that influence protein folding and structure are:
|
non-covalent
|
|
|
Disulfide bonds can influence protein structure. T/F
|
T
|
|
|
Non-covalent bonds are the only that influence protein folding and structure: T/F
|
F
|
|
|
Protein folding and structure is influenced by:
|
non-covalent bonds
disulfide bonds |
|
|
List the structures of protein and the bonds that influence them.
|
Primary
covalent Secondary Non-covalent Tertiary Non covalent and SH (covalent) Quaternary Non covalent and SH (covalent) |
|
|
Ligand binding occurs through this kind of bond:
|
non-covalent
|
|
|
An enzyme can increase the rate of a reaction in these ways:
|
1) Brings TWO substrates together in a precise orientation.
2) Realignment of charges within substrate. 3) Physical stress of substrate to generate more favorable state. |
|
|
If KM is SMALL Number:
|
Then it takes low level of [S] to fill the active site, and the ES complex is STRONG.
|
|
|
If KM is LARGE Number:
|
Then it takes high level of [S] to fill active sites, and the ES complex is WEAK.
|
|
|
K_M is analogous to:
|
Affinity
|
|
|
Allows comparison of enzymes and
prediction of enzyme kinetics, based on cellular concentrations of substrates |
K_M
|
|
|
When a compound binds to the active site and displaces or outcompetes the ligand for binding:
|
COMPETITIVE enzyme inhibition
|
|
|
When the enzyme is modified at a site different from the active site, to reduce the function of the enzyme.
|
NON-COMPETITIVE enzyme inhibition
|
|
|
This inhibition cannot be overcome by the addition of more ligand:
|
NON-COMPETITIVE enzyme inhibition
|
|
|
Non-competitive inhibition can be overcome by the addition of more ligand. T/F
|
F
|
|
|
Competitive inhibition causes this in Vmax and K_M:
|
Vmax same
K_M changes |
|
|
Non-competitive inhibition causes this in Vmax and K_M:
|
Vmax changes
K_M same |
|
|
The most common method of enzyme regulation involves feedback: T/F
|
T
|
|
|
Most common method of enzyme regulation. Usually involves control by amount of product formed. Reduces activity of enzymes in the pathway.
|
feedback inhibition
|
|
|
In a positive feedback loop this occurs:
|
Usually, product feeds back to increase its own production
|
|
|
In positive regulation this occurs:
|
Increased activity through binding by ligand.
|
|
|
How is enzyme activity changed?
|
allosteric changes
change in conformation due to covalent modification or binding of ligand |
|
|
The addition or removal of phosphoryl groups (PO4) from a protein is the most common modification used to affect protein function: T/F
|
T
|
|
|
The addition or removal of _______ from a protein is the most common modification used to affect protein function
|
phosphoryl groups (PO4)
|
|
|
Catalyzes addition of PO4 to serine/threonine/tyrosine
|
kinase
|
|
|
Removes PO4 moiety
|
phosphatase
|
|
|
This is the most important chemical modification in cells.
|
Phosphorylation / dephosphorylation
|
|
|
What is the difference between fibrous and globular proteins?
|
These are the quaternary shapes formed by proteins. Globular are spherical and soluble. Fibrous long filaments and insoluble.
|

