![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
54 Cards in this Set
- Front
- Back
|
What are some important considerations in choosing a cloning vector? |
Amount of product desired? Nature of the protein to be expressed? getting the protein out of the vector? |
|
|
What is conjugation in cloning? |
using a plasmid to insert new genes into a bacteria |
|
|
Transduction in cloning |
A phage injects DNA into a cell |
|
|
How is mRNA cloned? |
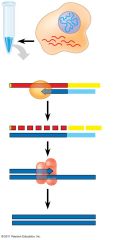
turn mRNA into cDNA which is then used for cloning. Isn't ideal to clone mRNA directly. Can be very useful for putting mammalian genes into prokaryotes, but they must be free of introns |
|
|
what is an oligonucleotide? |
Short sequences of nucleotides that can be used for genetic testing - designed to hybridize to specific sequences |
|
|
How could you encode an oligonucleotide using reverse translation? Would this yield one or multiple options? |
You could run different nucleotides over the protein sequence for the mRNA possibilities, then reverse transcribe possible probes. Would end up with different ones because different organisms would have different codon preferences |
|
|
what is a bacteriophage? |
a virus that infects bacteria. |
|
|
What are restriction enzymes and where do they originate in bacteria? |
Restriction enzymes target specific DNA sequences and cleave them. In bacteria - they target phage specific sequences and allow them to break them down so the phage doesn't kill the bacteria |
|
|
How do bacteria not cleave their own DNA with their restriction enzymes? |
Their DNA is supercoiled and harder for the enzymes to access. Phage DNA is relatively naked so it can be easily transcribed. |
|
|
What are restriction endonucleases sites? |
the areas targeted by restriction enzymes. Cleave DNA into sticky or blunt ends |
|
|
How can you use restriction enzymes to make recombinant DNA? |
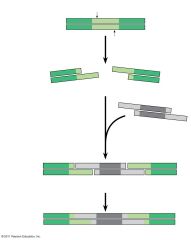
use the enzyme to cut sticky ends then add in DNA fragment (cut with same enzyme) with beginning and end parts that match the sticky ends |
|
|
what is a DNA probe? |
a complementary sequence that seeks out it's DNA sequence and hybridizes to it |
|
|
what is agarose gel electrophoresis |
takes advantage of the negatively charged phosphate backbone of DNA to provide directional pull on the fragment towards positively charged anode with larger fragments moving slower |
|
|
PCR (polymerase chain reaction) |
DNA strands, specific primers for start of gene, and nucleotides are heat cycled to cause denaturing, annealing, and replication of gene strands. Yields whole pure gene based on primer. |
|
|
Starting information or resources for cloning? |
protein sequence mRNA species/sequence cDNA libraries DNA sequences known/unknown genomic dna libraries PCR product |
|
|
what is CRISPR (clustered regularly interspaced short palindromic repeats) and what is RNA guided endonucleases (Cas9) |
CRISPR refers to a part of the bacterial genome that has short palindromic repeats then spacer DNA (non-coding) from the virus. During the first phage attack, part of the phage DNA is incorporated here as a spacer. During second attack, Cas9 endonuclease combines with viral RNA, incorporates into triple helix in viral DNA upon seeing PAM sequence and cleaves it at a particular locus. Before, it was very difficult to make double stranded breaks at the desired location! |
|
|
Why is CRISPR better than zinc finger editing? |
Doesn't require specific protein to be made for single site. Instead you just change the mRNA sequence a bit. Much easier |
|
|
What are plasmids? What is a limitation to using them to clone? |
small circular or linear DNA segments that carry a variety of genes - transferred between bacteria. if the DNA contained is closely related, it won't take. |
|
|
How would you clone into a plasmid? (use it as vector) |
cut using restriction enzyme, insert DNA segment cut with same enzyme, add DNA ligase to combine segments. small, easy to manipulate, can handle multiple copies of the gene. Can insert nonsense DNA to break up a gene or addition of new function |
|
|
What features are important to DNA cloning Vectors? |
Size of the vector vs. size of insert Make sure it has an origin of replication so it gets replicated Needs to be unique in endonuclease target site Marker gene ideal RNA polymerase promoter sequences for transcription initiation minimum amount of non-essential DNA to optimize cloning |
|
|
What is blue-white selection? |
disruption of a blue phenotype gene by insertion of another section of DNA to prevent it from expressing. then grow up the vector in culture and select visually for the mutant |
|
|
What is an expression vector? |
usually a plasmid or virus DNA that cone desired gene along with regulatory sequences to promote it being expressed in the host cell |
|
|
What is a shuttle vector? |
A cloning vector that can replicate in two different organisms (ex. E. coli and yeast) |
|
|
Are plasmids shuttle vectors, expression vectors or both? |
Both! They contain expression guiding constructs. Plus some have been developed that can go between two organisms. |
|
|
what is site-directed mutagenesis? |
Making pointed changes to a DNA sequence then cloning it into a vector or plasmid. Can be done synthetically |
|
|
What is gene disruption? |
Entering a "cassette" (DNA fragments) into a gene to disrupt it's function |
|
|
How could you study the function of a specific gene? |
disrupt it or clone it into another organism |
|
|
What is a fusion protein? Why would you make them? |
a protein created by the fusion of two different genes (usually needs one similar construct at least). Retains properties of each gene. Could help stabilize the protein or change solubility, could remove immune stimulating portions (organism specific) and replace with host proteins so the medicine isn't perceived as foreign |
|
|
What is an oncogene? |
A gene that can potentially cause cancer |
|
|
What is a Bacterial Artificial Chromosome? |
A large plasmid that can carry a large DNA insert (a few 100kb), also used for DNA library construction. Can insert large segment or individual segments (large plasmid) |
|
|
If you have low expression levels in a bacterial system, what could you do? |
change promoter, change plasmid, change cell type, add far tRNAs for rare codons on second plasmid |
|
|
If you have severe protein degradation in a bacterial system, what could you do? |
Use proteasome inhibitors and other protease inhibitors Try induction at a lower temperature |
|
|
Misfolded protein (inclusion bodies), what could you do? |
co-express with a chaperone protein or try refolding buffers |
|
|
What are baculoviruses? |
viruses that target insect cells. Shown to have very high yields! |
|
|
Can viruses hold as much DNA as bacterial artificial chromosomes? |
No! only about 20 kb of DNA |
|
|
Yeast artificial chromosomes - what are they? and how much data can they carry? |
linear plasmids essentially - linear DNA vectors that resemble normal yeast chromosomes. Contain telomeres that stabilize chromosome ends. They're shuttle vectors with a bacterial origin of replication. Can do hundreds of thousands of base pairs |
|
|
Retroviral vectors - what are they and how are they used? |
they're mRNA viruses that convert into cDNA by viral reverse transcriptase and is integrated into the host genome. Can reside there practically forever. Retroviral vectors are used for studying oncogenes and other human genes. Include lentiviruses |
|
|
Inducible expression (how does Tet on/ Tet off work) |
promoter is triggered by presence of a chemical - in this case, tetracycline. When Tet-on promoter used, presence of Tet turns it on, absence turns it off. In Tet-off, the reverse is true. Allows for specific control |
|
|
particle gun introduction of DNA to a cell |
Shooting DNA into the cell |
|
|
Transduction method of integrating DNA into vector cell. Generalized and efficient |
Phage transfers a portion of the host DNA to a recipient host cell. Generalized the virus incorporates fragments of the host at low efficiency, specialized it takes the adjacent genes so it can transduce more efficiently in the future |
|
|
Transformation method of integrating DNA in cell |
DNA is put around the cell and the cell uptakes it and transforms it's own DNA to incorporate it (similar to what plasmids can do). Could give the cell features of a dead cell |
|
|
Conjugation method of inserting DNA |
Bacterial conjugation - when two bacteria form a bridge and share DNA |
|
|
Microinjection method of introducing DNA or protein? |
use tiny needle to inject something into the soma of the cell |
|
|
Transduction in prokaryotes vs eukaryotes? |
prokaryotes - can transfer bacterial genes that provide antibiotic resistance, change nutritional requirements, etc (change niche in which bacteria can survive) eukaryotes - can transmit oncogenes (i.e. HBV and human and animal retroviruses |
|
|
Why is transduction important to phages in genetic engineering? |
It allows them to grab DNA from the host and take it to other cells after rapid replication |
|
|
Electroporation method of getting DNA into cells |
put plasmid DNA and bacteria in the same solution and supply brie electrical shock to create transient holes in the membrane. Allows plasmids to enter, then culture and select. |
|
|
What are DNA libraries? (what are the two types) |
they're collections of cloned DNA fragments from a particular organism contained in a bacteria or virus host.
Screened to pick different genes of interest Can use libraries to determine entire sequence of a gene when you only know the partial
Genomic DNA libraries and cDNA libraries |
|
|
Genomic libraries - what do they do/how are they made |
Take DNA fragments representing the entire genome of an organism, cut it with restriction enzyme and inserted in DNA cut with the same enzyme (ligase added), then recombinant vector is used to transform bacteria. Disadvantage: non-protein coding introns are clone as well, so will be mostly non-coding. Most animals have a large genome so searching for a single one is exhaustive |
|
|
cDNA libraries - how made? Advantages/disadvantages? |
mRNA reverse transcribed for cDNA, mRNA degraded, DNA polymerase added to make second strand of DNA, then cut with restriction enzyme, put in plasmid vector, and used to transform bacteria. Advantages: helps narrow down what you're looking for - know where the mRNA shows up, doesn't include introns. Disadvantages - can be difficult to make if you can't find mRNA in significant amount |
|
|
How could you test for gene expression in colonies? (hint - involves replica plating) |
Replica plate the cultured colonies. On one plate, lyse bacteria, denature DNA, add RNA or DNA probe washout unbound radioactivity. Read on x-Ray film to determine active colonies. On plate two, partially lyse cells and add antibody to bind to desired protein in radio labeled form. |
|
|
Advantages/disadvantages to E. coli and Bacillus subtilis as cloning hosts? |
E.coli - well developed genetics, many strains, best known prokaryote, but: potentially pathogenic and periplasm could trap proteins B. subtilis - easily transformed, nonpathogenic, naturally secretes proteins, forms endospores (easy culturing), BUT: genetically unstable and less developed than E. coli. |
|
|
Advantages/disadvantages to yeast? Saccharomyces cerevisiae |
Well developed genetics, can process mRNA and proteins (for higher organism gene products), easy to grow, BUT plasmids are unstable and will not replicate most prokaryotic plasmids |
|
|
Mammalian cells for genes? how could you use baculoviruses to overcome certain downfalls? |
Great for studying protein structure and function in genetics, cancer, diseases, etc. but expensive to maintain and levels of recombinant gene expression is often low. Can increase expression with baculoviruses in insect cell lines. |
|
|
Reporter gene - what is? |
It's a gene embedded in the DNA of a vector that encodes a protein that expresses some sort of signal. It's besides the promoter and measures the existence of the protein (potentially mRNA?) instead of the location of the gene. Could be on a rate limiting molecule to determine protein production |

