![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
86 Cards in this Set
- Front
- Back
- 3rd side (hint)
|
Domain Bacteria |
-Prokaryotic -Habitats: terrestrial and acquatic -Reproduction: asexual Size range: 0.3 --> 100 micrometers |
|
|
|
Bacteria types of shapes and patterns (vocabulary) |
Coccus = sphere diplococcus = pairs streptococcus = chains staphylococcus = grape-like clusters tetrads = 4 cocci in a square |
_______ = sphere diplococcus= pairs |
|
|
Shapes and arrangements (vocab) |
Bacillus = rod vibrio = curved rod spirillum = rigid helix spirochete = flexible helix pleomorphic = variable hyphae = long filaments mycelium= network of hyphae |
|
|
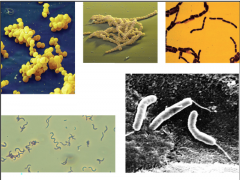
identify type (lecture two) |
1) staphylococcus 2) streptococcus 3) bacillus 4) spirillum 5) vibrio |
|
|
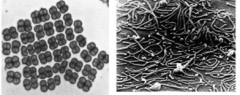
identify type (lecture two) |
1) tetrads (genus deinococus) 2) pleomorphic (genus mycoplasma) |
|
|
|
Advantages of being small (figure 3.5 in textbook) |
- smaller cells, higher surface to volume ratio - efficient nutrient uptake and diffusion in cell -rapid growth rate |
|
|
|
Functions of the cell membrane |
- separates cell from environment -main site energy generation -selectively permeable barrier -transport systems (contains the transport proteins) |
|
|
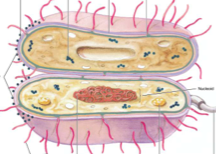
learn cell envelope and plasma membrane |
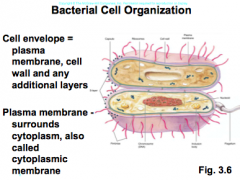
|
|
|
|
plasma membrane (makeup and function) |
- lipid and protein -- lipids form bilayer -- with embeded protiens - organized, asymmetric, flexible, dynamic |
|
|
|
Lipid interaction picture |
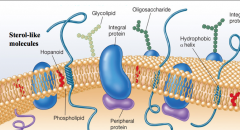
|
hopanoids are sterol-like molecules |
|
|
Bacterial cell wall (definition and functions) |
Definition: rigid structure, found outside the membrane functions: - determines the shape of cell - protection from toxic substances (osmotic lysis) - osmosis: water moving across a membrane - makes sure that cell wont burst due to excess water. |
relevant (hypo vs hypertonic solutions) |
|
|
Bacteria are most often in hypo/vs/hyper tonic solutions? |
hypotonic [solute] outside < [solute] inside
|
|
|
|
Cell walls of bacteria (definition) |
- bacteria are divided into two major groups based on the response to gram-stain procedure (gram + = stain purple) (gram - = stain pink) |
|
|
|
gram positive vs gram negative cell walls (picture in ans slide) |
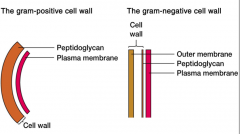
|
gram positive stains purple due to crystal violet gram negative stains pink due to saffron |
|
|
Peptidoglycan structure |
-important component of cell wall in gram + and gram - bacteria -polysaccharide formed from subunits -two alternating sugars (N-acetylglucosamine[NAG} and N-acetyimurami acid [NAM]) -sugar backbones cross-linked by peptides |
|
|
|
Peptidoglycan key features |
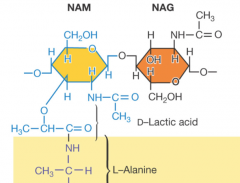
|
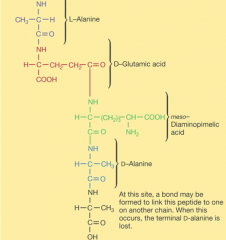
|
|
|
peptidoglycan picture in sequence |
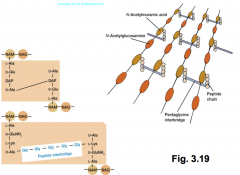
|
-transpeptidation: process of connecting one chain of amino acids to another chain of amino acids |
|
|
Gram positive cell walls |
-primarily peptidoglycan -also teichoic acids (polymers of glycerol or ribitol)
|
|
|
|
Gram negative Cell walls |
- thin layer peptidoglycan surrounded by outer membrane of lipids (lipopolysaccharide(LPS)) -no teichoic acids |
|
|
|
Importance of Lipopolysaccharides (picture in hint icon) |
-protection from host defenses -- O antigens vary (example, E.coli O157 H7) -attachment -stability lipid A portion of LPS can act as a toxin (called endotoxin) causes fever, septic shock |
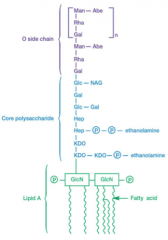
three parts: lipid A core polysaccharide o side chain (o antigen) |
|
|
Gram+ vs Gram- (hpic for Gram-) Teichoic Acid Periplasmic Space Plasma Membrane Outer Membrane Lipopolysaccharide Peptidoglycan Capsule |
Gram+ Teichoic Acid: yes Periplasmic Space: Plasma Membrane: Outer Membrane: Lipopolysaccharide: Peptidoglycan: Capsule: yes |
Gram- Teichoic Acid: no Periplasmic Space: Plasma Membrane: Outer Membrane: Lipopolysaccharide: Peptidoglycan: Capsule: yes |
|
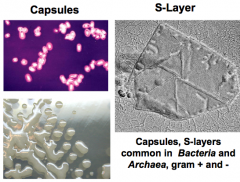
Layers outside the cell wall (hpic has functions) |
Capsules: polysaccharides, organized, not easily removed. Slime layers: polysaccharides, diffuse, unorganized, easily removed. S-Layers: proteins, organized
|
Functions: attachment, protection, resistance to chemicals and host defense. dessication |
|
|
Cytoplasm of bacteria and archaea (hpic) |
substance in which includes chromosomes and ribosomes. mostly water |
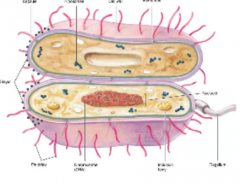
|
|
|
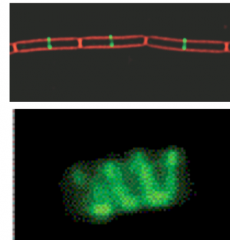
Cytoskeleton (highly organized) hpic |
-FtsZ (tubulin-like protein, forms contractile ring, septum formation and cell division) -MreB and Mbl (actin-like proteins, may form coils in rod-shaped cells, like a spring) |
|
|
|
Cytoplasm (inside one) |
inclusions: granules of organic or inorganic material stockpiled by cell for future use. can be membrane-enclosed. |
|
|
|
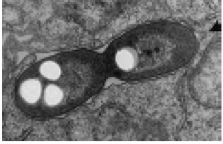
Storage inclusions hpic |
Storage inclusions: stores energy and carbon, like glycogen, poly-B-hydroxybutyrate. also stores phosphate and sulfur. Carbon and nitrogen, in cyanophycin granules.
|
|
|
|
Microcompartments hpic |
-Carboxysomes: polyhedral shape, protein coat. enzymes inside carbonic anydrase takes carbonic acid and makes into CO2 rubisCO fies CO2 into sugar uses energy from photosynthesis
|

|
|
|
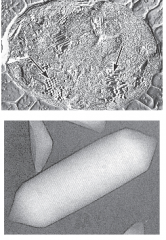
Inclusions (gas vacuoles) hpic |
found in acquatic photosynthetic bacteria and archaea |
|
|
|
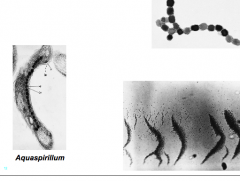
Magnetosomes (hpic) |
type of inclusion, iron in the form of magnetite (Fe3O4) |
|
|
|
Translation and transcription in Cytoplasm |
RNA polymerase transcribes DNA to mRNA Ribosomes translate mRNA to protein in bacteria and archaea processes occur simultaeously |
|
|
|
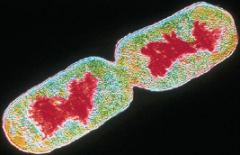
Things found in cytoplasm hpic |
Nucleoid: region containing chromosomes (closed, circular doublestranded DNA) not membrane enclosed. Plasmids: small, closed circular DNA, they exist and replicate independently from chromosomes, may carry genes that confer advantages (Conjugataive plasmid, R-plasmid) R=resistance |
|
|
|
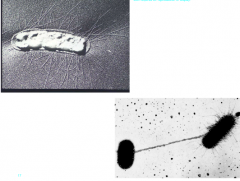
External structures of the Cell wall (hpic) |
functions: attachment, horizontal genetransfer, and movement. Pilli (thin protein appendages) -attachment protein (shown in hpic)
|
|
|
|
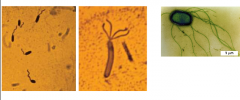
Bacterial Flagella |
motility organelles found in all organic life. (from left to right) Polar,Lophotrichous,Peritrichous (monotrichous = one flagella lophotrichous,= cluster on one end, peritrichous, all over) |
|
|
|
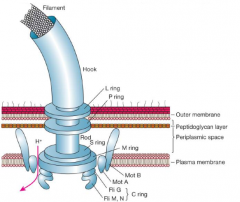
Bacterial flagella, structural components. |
|
|
|
|
Chemotaxis(hpicnotes) |
Chemotaxis: movement toward or away from a chemical attractants cause C-clockwise rotation, flagella bundle Repellents = clockwise, flagella fly apart and cause a "tumble" that changes direction |
chemoreceptors detect attractant CCW rotations are favored biased random walk net movement is towards the attractant |
|
|
Carboxysomes |
found in photosynthetic cyanobacteria, but also in other non-photosynthetic bacteria that fix CO2 |
|
|
|
Chemotaxis motility |
CCW rotation of flagellum propels forward CW rotation propels backwards Bacteria with single flagellum dont tumble |
|
|
|
Archaea structure and function
(hpic notes) |
-prokaryotic
-asexual -terrestrial and aquatic (found in both land and sea) -have circular doublestranded- DNA chromosomes and plasmids -has histone proteins, similar to ours |
-some are thermophiles (can grow/survive at high temperatures)
-some are methanogens (generate CH4 methane) -have gram +/- diversity -have plasma membranes, some have cell walls -NO PEPTIDOGLYCAN! -some have no cell wall, S-layer instead |
|
|
Archaeal lipids and membranes (hpic notes) |
Ether linkage in membrane is more stable but less flexible than ester linkages found in bacterial membranes. |
comes in both bilayers and monolayers with tetraethers. |
|
|
Archaeal membranes |
membranes provide enhanced stability for survival and growth at high temperatures. |
|
|
|
Viruses(hpic notes) |
-not cells -not living -infects living cells to replicate -bacteriophages type a virus to infect bacterial cells -depends on host metabolism |
-protein and nucleic acid -simple mechanism -small: range of nm
|
|
|
General properties of viruses -structure (hpic) |
Virion: complete virus particle Capsid: protein coat around the genome Nucleocapsid: Nucleic Acid covered by the capsid Protomer: protein subunit of capsid |
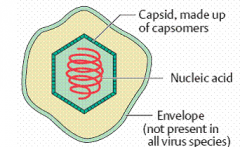
|
|
|
Morphological types (shapes of viruses) hpic |
polyomavius: icosahedral tubulovirus: helical herpesvirus: enveloped T-even coliphage: binal |
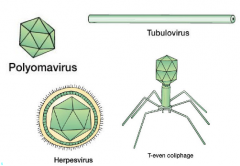
|
|
|
Helical Capsids hpic |
- hollow tubes with protein walls (picture) has RNA rahter than DNA, helical pattern -Influenza virus also has helical capsid structure |
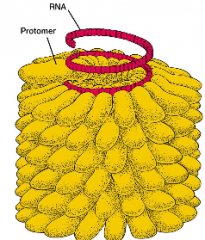
|
|
|
icosahedreal capsids hpic |
-efficeint way to enclose a space -20 triangular faces -ring-shaped units - capsomers -capsomers = 5-6 protomers. |
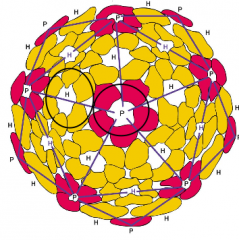
|
|
|
Binal capsids (T0even coliphage) hpic |
tail pins and tail fibers cling to cell, and chromosomes for replication travel from head, down core/tube and enter the cell |
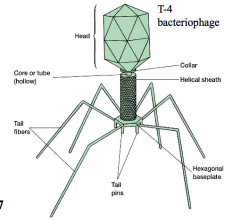
|
|
|
viral envelope and spikes hpic |
- the capsid of some viruses is enclosed in an envelope from host cell membrane -envelopes has protein spikes, encoded by virus (these spikes are to help attach to cells) -between envelope and capsid, tegument proteins sometimes found |
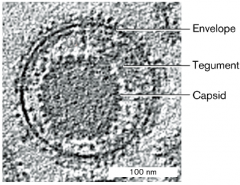
|
|
|
structure of HIV hpic |
-gp 120 very important to how HIV attacks humans - genome is RNA (2 copies) -some viruses also carry enzymes with them |
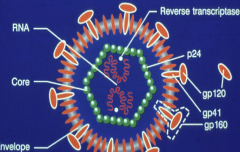
|
|
|
Influenza Virus hpic |
- envelope has spike proteins -Neuraminidase spike: has an enzymatic function of spreading the infection notice the RNA replicase and segemented genome -Hemagglutinin spike, initiates initial interaction to host cell |
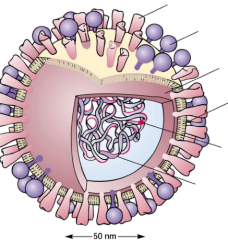
-Hemagglutini spike -neuraminidase spike -matrix protein -envilope -RNA rep -Seg RNA genome |
|
|
Viral genomes hpicnotes |
-Either DNA or RNA contained inside -Single (ss) or double stranded (ds) -linear or circular (shapes and morphologies) -Encode viral proteins (ex gp120)
|
EX) retrovirus genome: 3 major parts of genome capped by LTR on each side gag: capside protein genes pol: polymerase protein genes (reverse transcriptase encoded here, also integrase & protease) env: envelope protein genes found here.
|
|
|
Viral multiplication/infections cycle |
1) virus attaches to host cell 2) entry and uncoating (reveals genetic material still within virus) 3)synthesis of viral proteins and nucleic acid with host cell 4) assembly of capsids to cover genomes 5) release of virions within host cell |
|
|
|
Animal Virus Attachment (hpicnotes) |
- viral surface proteins mediate attachment to host receptors such as carbs, proteins, and lipids. |
ex) hemaglutinin binds host sialic acid HIV gp120 binds to CD4 receptor and CCR5 co-recepter in human T-cells Tropism: targeting of a particular virus to host cell or tissue |
|
|
Animal virus Entry (hpic) |
1) fusion with host membrane 2) endocytosis (hijacking the way our own cells uses endocytosis to gain entry) pinched off inside of a vacuole |
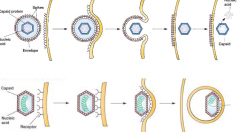
|
|
|
Basics of Viral replication (hpic) |
DNA replicates transcription RNA is made Translation proteins are made |
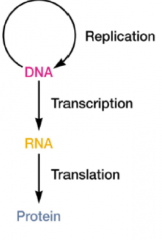
|
|
|
Life cycle facts of viral genome (hpicnotes) |
DNA viruses -typically replicate in the nucleus -use host DNA polymerase -exception: Herpes uses their own DNA polymerase rather than ours |
RNA viruses -typically replicate in cytoplasm -use RNA replicases |
|
|
Animal virus synthesis and assembly |
-Synthesis -All viruses make proteins using host ibosomes -Translation in cytoplasm (requires that nucleus of virus must enter host cell) -Assembly -Capsid + genome (use the host cell as a scaffold to incorporate their own genetic material) takes place usually in cytoplasm, but also in nucleus -spike proteins insert into membrane |
|
|
|
Animal virus release mechanisms (hpic) |
-Lysis (the breaking of membrane to release into host) -Budding: membrane lipids surround capsid to form envelope |
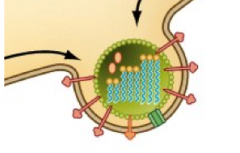
|
|

Even smaller than a virus |
-Viroids: RNA only (not viruses) - Do not encode proteins - RNA may pair with plant RNA, this will create a Doublestranded RNA. (RNA silencing) -Prions: infections proteins (not viruses, b/c no RNA, just proteins) (associated with neurodegenerative diseases, ex mad cow) - Abnormal protein form -plaque formation, that causes cell death |
|
|
|
Microbial nutrition |
-To obtain energy and construct new cellular components, microbes must have a supply of raw materials and nutrients
-nutrients - substances used in biosynthesis and energy release - required for growth |
|
|
|
Major elements of microbial cell (makes up 95% of its weight) |
-Macronutrients (needed in large amounts) (C, O, H, N, S, P)
-Micronutrients: trace elements (very little required) (cobalt, copper, zinc, manganese) |
|
|
|
Growth Factors (that some microbes require to synthesize certain compounds) |
-Amino acids -Purines and pyrimidines -Vitamins ex) microbes in intestinal tract |
|
|
|
Sources of Nitrogen to microbes |
-Ammonia(NH4): can diffuse into cells, incorporated into cell material -Nitrate (NO3): first reduced to ammonia by assimilatory nitrate reduction - then incorporated - Nitrogen Gas (N2): 79% of the atmosphere, requires nitrogen fixation for microbes to utilize atmospheric nitrogen (N2 reduced to ammonia) |
|
|
|
Nitrogen Fixation (when N2 is reduced to Ammonia NH4) |
Azotobacter: free living in the soil Rhizobium: in symbiosis with plants (establish in the roots of plants, then feeds the plant ammonia in return of a good home to chill) |
|
|
|
Problems with acquiring nutrients for microbes |
-Rapid growth of microbes present acquiring nutrients (competition) -Food must enter..... (at high rates and across membranes) -selective fashion (must be able to identify nutrient vs toxic substance) -fights against concentration gradient |
|
|
|
Ways microbes attain nutrients hpic |
Diffusion: -no energy required -requires gradient (higher to lower) -two examples: passive and facilitated (* facilitated uses mobile carriers called permeases) |
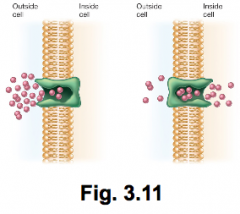
|
|
|
Rates of transport differences between diffusion methods |
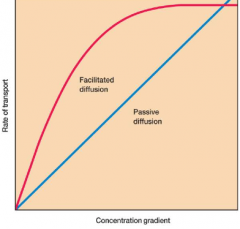
|
|
|
|
Primary active Transport (ABC transporters) hpic |
- found and identified in all domains of life -requires ATP to function -2 types: Uptake ABC transporters (moving nutrients in the cell) and Efflux ABC transporters (multidrug drug pumps move things from inside the cell to outside) |
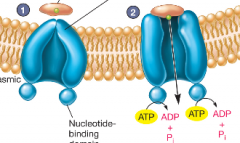
1) after binding solute, the protein moves to ABC-T 2) protein attaches to transporter and releases solute driven by ATP into cell |
|
|
Secondary active transport (hpic) |
- Uses potential energy of ion gradient -ex: electron transport across membrane generates proton (H+) gradient (this is the same energy that causes flagellum movement) -Symport and antiport |
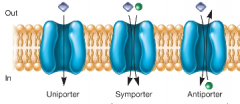
|
|
|
Active transport (group translocation) hpic |
-Phosphotransferase system (PTS) is an example in all bacteria -Nutrient chemically altered during transport -Energy for this alteration comes from phosphoenopyruvate (PEP) that attaches Phosphorus to sugars |
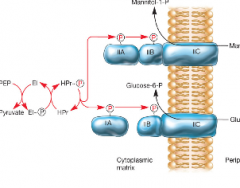
|
|
|
Iron uptake (absorption of iron in microbes) |
-problem( all microbes require iron, but there is little free Fe available) -Solution: microbes release siderophores to acquire Fe. They will bind up Fe and is transported into the cell by an ABC transporter |
|
|
|
Reproductive strategies (spore germination mechs) |
-Eukaryotic Microbes -Sexual and asexual
|
|
|
|
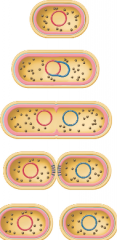
steps of binary fission:(hpic) |
1) DNA replicates 2) cell elongates and chromosomes segregate 3) septum forms (cleavage in the center) 4) cell divides finally |
|
|
|
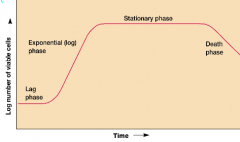
Growth curve (hpic) |
growth is studied by analyzing the growth curve of a microbial culture Lag phase: no growth of cells, currently synthesizing new components (Replenish, adapt) exponential(log) phase: balanced constant growth of microbial cells. (doubling # at constant interval) Stationary: they are merely survivingw/ no food death phase: slowly dying off due to no food |
|
|
|
Batch culture |
a closed vessel, a single batch of medium. Used to study and examine a growth curve |
|
|
|
Generation (doubling time) |
the rate of exponential growth: the time required for cells or population to divide - range: 7 min to 24 hours |
|
|
|
Sporulation: hnotes |
Under nutrient limiting condition, some microbes form stress-resistant and dormant spores. |
-some lived for thousands of years to only wake up due to nutrients ex: bacillus, clostridium |
|
|
measuring microbial growth (several methods) |
1) direct cell counting (counting chambers) 2) viable cell counts (plating, colony forming units) 3) Turbidity measurements (microbial cells scatter light striking them, more turbid = more cells = more light scattered, spectrophotometer used to observe) |
|
|
|
Extremophiles (hpic) microbes that live under extreme conditions |
-Thermophiles: ones that can survive in high - psychrophiles: can survive extreme cold - mesophiles: microbes that survive in middle termps (20-45 celcius) -osmophiles: live in high solute concentrations (halophiles) can live in very salty enviroments -acidophiles: live in acidic enviroments |
|
|
|
Physical requirements of temperature |
Hyperthermophiles: >80(C) Thermophiles : 40->80 (C) Mesophiles: 20->45 (C) Psychrophiles: 0->20 (C) |
|
|
|
How did thermophiles adapt to fluctuations of temperature? |
-Proteins are stabilized: increase H-bonds and covalent bonds, molecular chaperones that bind and refold dmged proteins. DNA stabilized: coat DNA with proteins to stop heat degeneration membrane stabilization: more saturated and branched |
|
|
|
Physical requirements: water related |
water is solvent in which molecules of life are dissolved. availability of water affects growth of all cells expressed as water activity, (Aw) higher [solute] = lower water activity (Aw) |
|
|
|
Osmophiles |
-grow at low Aw (water activity) and live in hypertonic environments [solute]outside >>[solute]inside Halophiles:live in high salt concentrations ex: staphylococcus aureus |
|
|
|
Changes in pH acidophiles and Agolophiles |
(find later) |
|
|
|
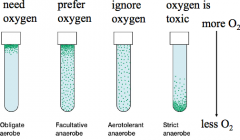
Microbes can grow in wide range of oxygen concentrations (hpic) |
Obligate aerobes: need oxygen facultative anaerobe: prefer oxygen Aerotolerant anarobe: ignore oxygen strict anaerobe: oxygen is toxic |
|
|
|
Basis of different oxygen sensitivities |
- oxygen reduced to toxic products (ROS) superoxide radical, hydrogen peroxide -aerobes have protective enzymes: superoxide dismutase O2 + O2 + 2H --> H2O2 + O2 H2O2 + H2O --> 2H20 + O2 |
|
|
|
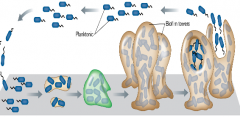
Biofilms - microbial community (mix of microbes) attached to surface, covered with polysaccharide matrix (hpic) |
1)biofilm is initiated with mobile planktonic cells, 2)then attachment to a surface 3) lose tails and colonization process begins after attaching to surface. (quorum sensing: cell density dependent signal) 4) mature biofilm mushroom and dispersal restarts the process
|
|

