![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
402 Cards in this Set
- Front
- Back
- 3rd side (hint)
|
System (Bioenergetics)
|
Restricted portion of the universe being studied.
State of system defined by variables. Study changes by intial->final states. |
|
|
|
Calorie (cal)
|
Amt of E to heat 1g of water 1 degree Celcius at 1 atm.
Joule (J)=0.239cal/mole |
|
|
|
dE equation
|
E prod's. - E react's.
|
|
|
|
neg. dG
|
Exergonic
Spontaneous |
|
|
|
pos. dG
|
Endergonic
Nonspontaneous |
|
|
|
Entropy (s)
|
A measure of randomness or disorder.
ds = change in entropy w/ rxn. higher entropy=more disorder |
|
|
|
Enthalpy (H)
|
Heat content
dH can approximate dE dH = H prod's. - H react's. |
|
|
|
neg dH
|
exothermic - heat released
|
|
|
|
pos dH
|
endothermic - heat absorbed
|
|
|
|
Irreversible Inhibition
|
Inhibitor binds covalently to active site.
|
|
|
|
Michaelis-Menton kinetics
|
Rate increases w/ increase in substrate conc.
With each increase-smaller increases in rate. Can alter rate by increasing [E]. |
|
|
|
Enzyme Kinetics
|
Rate of rxn.
Enzyme activity depends on conc. of substrates, products, and inhibitors. Measured in terms of initial rxn rate - before substrate decreases and product accumulation is small. |
|
|
|
Active site
|
Usually groove or pocket.
Part of 3 prime configuration. |
|
|
|
Catalysts
|
Inc. rate of rxn by lowering activation E.
Forms transient complexes - facilitates interactions b/t reactants. Changes ONLY rate at which equilibrium is reached. |
|
|
|
Activation Energy
|
Specific to every rxn.
Minimun E amt reactants need to be converted to product. |
|
|
|
Protein Secondary Structure
|
Regions of polypeptide coil into a helix.
2 structural patterns: alpha helix and beta sheet. |
|
|
|
Alpha helix
|
A form of secondary structure.
Spiral in shape, consisting of a backbone of peptide bonds w/ specific R groups of a.a. residues jutting out from it. Each turn of the coil stabilized by H2 bonds b/t the CO and NH groups of one peptide bond and those of the peptide bonds 4 a.a. away in each direction. |
|
|
|
Beta sheet
|
Protein secondary structure.
An extended sheetlike conformation w/ successive atoms in the polypeptide chain located at the peaks and troughs of the pleats. R groups of the a.a. jut out on alternating sides of the sheet at each fold. Stabilized by H2 bonds b/t the CO and NH groups of peptide bonds in adjacent polypeptide regions. |
|
|
|
Protein Primary structure
|
Sequence of a.a. linked together by peptide bonds forming a polypeptide (covalent bonds).
|
|
|
|
Why is carbon so important to biological molecules?
|
Specific bonding properties.
"valence of 4" prop. - can bond w/ each other. Forms covalent bonds. Stable organic cmpds have 4 covalent bonds for every C atom, giving C-containing molecules great structural and functional diversity. |
|
|
|
Bond Energy
|
Amt of E required to break 1 mole (6x10^23) of a bond.
Expresses stablility. |
|
|
|
Why is the polarity of water so important?
|
It accounts for its cohesiveness - Forms H2 bonds.
Temp. stabilizing capacity - due to its extensive H2 bonding, buffers soln's against large temp. changes. Excellent solvent - can form spheres of hydration. |
|
|
|
Vacuole
|
Storage vescicle
May hold food molecules. In plants - maintain turgor pressure. |
|
|
|
Ribosome
|
Protein synthesis.
Not an organelle. 2 subunits |
|
|
|
Microtubules
|
Organize cytoplasm (cell shape).
Cilia and flagella. Largest; mitotic spindle fibers. Hollow cylinders of tubulin subunits. |
|
|
|
Microfilaments
|
Muscle fibriles (locomotion).
Smallest. Actin polymers (G-actin). |
|
|
|
Peroxisome
|
Generates and degrades H2O2.
In animals, found in liver and kidneys - detox and catabolize, breakdown of fatty acids. |
H2O2 -> H2O + O2
|
|
|
Secretory vescicles
|
Molecules for export.
Synthesized in rough ER. Packaged in Golgi. Export via exocytosis. |
|
|
|
Lysosome
|
Vescicle w/ hydrolases.
Recycling. Enzymes synthesized on rough ER and packaged in Golgi. |
|
|
|
Golgi Complex
|
Stack of flattened vesicles.
Processes and packages secreted proteins - accepts vescicles from rough ER. Synthesizes polysaccharides. |
|
|
|
Endoplasmic Reticulum (ER)
|
Membrane Network - interconnected cisternae - tubular membranes.
Continuous w/ outer nuclear membrane. Rough ER b/c of ribosomes. Smooth ER - lipid synthesis, detoxification enzymes. |
|
|
|
Chloroplast
|
Photosynthesis in plants and algae.
Numerous in plants. Dark rxns - cell makes sugars. Double Membrane - also 3rd membrane = thylakoid sacs. Own DNA and ribosomes. |
|
|
|
Makes ribosomal RNA?
|
Nucleolus
|
|
|
|
Bacterial genetic material
|
Circular DNA molecule.
Some proteins. |
|
|
|
Nucleoid
|
In prokaryotes.
Folded genetic material. Usually attached to cell membrane. |
|
|
|
Protein functions
|
Enzymes - catalyst for rxns.
Structure - support and shape. Motility - contraction and movement (actin and myosin). Regulatory - control, coordination of cell processes. |
|
|
|
Amino Acids (a.a)
|
Monomer of proteins.
20. All have central carbon. Side chains vary in biochemistry. |
|
|
|
Phospholipid structure
|
Polar head - hydrophilic.
2 non-polar hydrocarbon tails - hydrophobic. Polarity of the hydrophilic head is due to a neg. charged phosphate group often linked to a pos. charged group (amino group). |
|
|
|
Amphiphatic molecule
|
Have both hydrophilic and hydrophobic regions.
Membrane phospholipids are amphipathic. |
|
|
|
Phospholipid structure
|
Polar head - hydrophilic.
2 non-polar hydrocarbon tails - hydrophobic. Polarity of the hydrophilic head is due to a neg. charged phosphate group often linked to a pos. charged group (amino group). |
|
|
|
Amphiphatic molecule
|
Have both hydrophilic and hydrophobic regions.
Membrane phospholipids are amphipathic. |
|
|
|
Polysaccharides
|
Polymers of sugar - usually 1-2 types per polymer.
Used for storage and structure. Starch, glycogen, cellulose. |
|
|
|
Monosaccharide
|
Monomer of polysaccharides
Usually 3-7 C's. General formula Cn H2n On Glucose most common. |
|
|
|
Most common monosaccharide in biosphere/cells?
|
Glucose
|
|
|
|
Nucleic acids
|
RNA, DNA
Storage, transmission, expression of genetic info. Differ by sugar molecule and one base. |
|
|
|
Nucleic acid monomer
|
Nucleotides
|
|
|
|
Lipids function
|
Energy storage
Membrane structure Other specific functions |
|
|
|
Steroid structure
|
4 fused rings
Differ by # and position of double bonds |
|
|
|
Steroids
|
Only in eukaryotic cells
Cholesterol most common |
|
|
|
Cholesterol
|
Most common steroid
Starting pt in synthesis of sex hormones - glucocorticoids and mineralocorticoids. Found in cell membranes |
|
|
|
What allows proteins to shift conformations?
|
Weak interactions such as H2 bonds.
|
|
|
|
Peptide bond
|
A covalent C-N bond linking 2 a.a.
Formed by dehydration. |
|
|
|
What is a dimer?
|
A multimeric protein composed of 2 polypeptides
|
|
|
|
Starch
|
Storage polysaccharide found in plant cells.
|
|
|
|
Glycogen
|
Storage polysaccharide found in animal cells, also bacteria
|
|
|
|
Energy
|
The capacity to cause specific physical or chemical changes.
The capacity to do work. |
|
|
|
Cellular activities which give rise to specific physical and chem. changes (6).
|
Synthesis
Mechanical - Change in location Concentration Electrical Heat - by product of rxns Bioluminescence - light |
|
|
|
Synthesis
|
Biosynthesis
Change in chemical bonding Formation of new chem. bonds. Maintaining chem. and cellular structures. Photosynthesis |
|
|
|
Mechanical work
|
Positioning/orientation
Change in location Use of cilia and flagella. Muscle contraction |
|
|
|
Concentration work
|
Changes in chem. concentration across membrane.
Movement of molecules across a membrane against a concentration gradient. Accumulate or removal of molecules. |
|
|
|
Electrical work
|
Movement of ions across a membrane against a electrochemical gradient.
Maintaining electrical potential. Nerve impulse transmission. ATP production. |
|
|
|
Heat
|
An incr. in temp. that is useful for warm blooded animals.
Regulation of body temp. Enzymes function at 37 C in our bodies. |
|
|
|
Bioluminescence
|
Production of light.
Generated by rxn of ATP w/ specific luminescent cmpds. |
|
|
|
Phototrophs
|
Can obtain E for cellular needs by capturing light E from the sun using specific pigment systems.
Transform this E to chem. E and store the E in the form of ATP. |
|
|
|
Chemotrophs
|
Rely on oxidation of chem. bonds in organic or inorganic (chemoautotrophs) molecules for E.
|
|
|
|
Oxidation rxns release _____?
|
Energy
|
|
|
|
DIC treatment?
|
**Identify and treat the underlying disease!
Also transfusion (especially fresh frozen plasma) |
|
|
|
Spontaneous heat transfer
|
Always occurs from the hotter place to the colder place.
|
|
|
|
First Law of Thermodynamics
|
The law of conservation of E.
In every physical or chem. change, the total amt of E in the universe remains constant, although the form of the E may change. E cant be created or destroyed |
|
|
|
Internal Energy
|
The total E stored w/in a system.
|
|
|
|
Enthalpy
|
Heat content
Represented by H |
|
|
|
Second Law of Thermodynamics
|
Law of thermodynamic spontaneity.
In every physical or chem. change, the universe always tends toward greater disorder or randomness (entropy). Allows us to predict in what direction a rxn will proceed under specified conditions. |
|
|
|
Thermodynamic spontaneity
|
A measure of whether a rxn or process can go, but it says nothing a/b whether it will go.
Directionality important |
|
|
|
Two alternative means of assessing thermodynamic spontaneity.
|
Entropy
Free energy |
|
|
|
Entropy (S)
|
A measure of randomness or disorder.
Can't quantify directly, but can get some feel for it. Increases: Melting, evaporation of gasoline, burning paper |
|
|
|
Link b/t spontaneous events and entropy changes
|
Whenever a process occurs spontaneously in nature, entropy of the universe increases.
dS is positive for all spontaneous rxns/events. |
|
|
|
Free energy (G)
|
Measure of spontaneity for a system alone, not including the universe.
Related to enthalpy and entropy |
|
|
|
dH is positive when?
|
Endothermic rxns
|
|
|
|
dH is negative when?
|
Exothermic rxns
|
|
|
|
Second Law of Thermodynamics and Free Energy
|
All processes that occur spontaneously result in a decrease in the free energy (dG) content of the system.
An exergonic rxn. |
|
|
|
Exergonic
|
An energy-yielding rxn.
Decrease in free energy (G). |
|
|
|
Endergonic
|
An energy-requiring rxn.
Increase in free energy (G). |
|
|
|
Spontaneity
|
Tells us only that a rxn can go.
Says nothing a/b if it will go. A rxn can have a -dG and no proceed to any measurable extent. |
|
|
|
Keq
|
Equilibirium constant.
Ratio of product conc. to reactant conc. at equilibrium. |
|
|
|
What drives every rxn?
|
Tendency to equilibrium
|
|
|
|
The amt of free E available from a chem. rxn depends on what?
|
How far the components are from equilibrium.
|
|
|
|
dG calculates what?
|
Energy needed to reach equilibrium.
|
|
|
|
K'eq > 1.0
|
dG' neg.
Rxn will proceed to rt under standard conditions. Products predominate over reactants at eq. |
|
|
|
K'eq < 1.0
|
dG' pos.
Rxn will proceed to lft under standard conditions. Reactants predominate over products at eq. |
|
|
|
2 ways to incr. the proportion of molecules w/ sufficient E to overcome Ea.
|
Incr. avg E content of all molecules - Input of heat
Lower Ea requirement - catalyst |
|
|
|
Catalysts
|
Enhances the rate of rxn by lowering Ea.
Ensures that a higher proportion of the molecules possess suffiecient E to undergo rxn w/out the input of heat. Do NOT change position of eq. |
|
|
|
a.a. common in active sites
|
Cysteine
Histidine Serine Aspartate Glutamate Lysine |
|
|
|
Prosthetic groups
|
Specific non protein components sometimes on an active site.
Usually either metal ions or coenzymes. Frequently func. as electron acceptors (a.a. not good acceptors) |
|
|
|
Coenzymes
|
Small organic molecules derived from vitamins.
|
|
|
|
3 mechanisms of substrate activation
|
Bond distortion
Proton transfer Electron transfer |
|
|
|
Bond distortion (substrate activation)
|
Change in enzyme conformation by initial substrate binding to the active site.
Distorts 1 or more bonds. Makes more susceptible to catalytic attack. |
|
|
|
Proton transfer (substrate activation)
|
Enzyme may accept or donate protons.
Inc. chem. reactivity of the substrate. Via charge a.a. in active site. |
|
|
|
Electron transfer (substrate activation)
|
Enzymes may accept or donate electrons.
Forms temporary covalent bond b/t enzyme and substrate |
|
|
|
The catalytic event
|
Substrate collides w/ active site forming temporary bond.
Changing in enzyme conformation for better fit. Products formed and released. Active site available for another substrate molecule. |
|
|
|
Initial rxn velocity (v) depends on what?
|
Substrate concentration
|
|
|
|
Maximum velocity (Vmax)
|
As substrate conc. becomes large, value of v reaches a max.
Depends on # of enzymes, can only be incr. by adding more enzyme. |
|
|
|
Saturation
|
Inablility of incr. substrate conc. to incr. rxn velocity.
|
|
|
|
Saturation
|
All available enzyme molecules are operating at max. capacity.
|
|
|
|
Km
|
Concentration of substrate that gives exactly half the max. velocity.
|
|
|
|
Low Km
|
The lower the Km value for a given enzyme and substrate, the lower the substrate conc. range in which the enzyme is effective.
|
|
|
|
Enzyme regulation
|
Incr. product decr. rxn rate.
2 mechanisms: allosteric regulation and covalent modification |
|
|
|
peu importe
|
never mind, it does not matter
|
|
|
|
Allosteric enzymes
|
Have 2 forms
High affinity for substrate or little to none. Forms are interconvertible. |
|
|
|
Allosteric effector
|
Small organic molecule that regulates the activity of an enzyme for which it is neither the substrate nor immediate product.
|
|
|
|
Allosteric effector mechanism
|
Binds to one of the two forms of an allosteric enzyme, stabilizing it.
Binds to allosteric site which is distinct from active site. May be inhibitor or activator. |
|
|
|
Cooperativity
|
Property of allosteric enzymes.
Positive or negative As the enzyme binds substrate, the enzyme changes conformation which affect the affinity of remaining substrate sites. |
|
|
|
Covalent modification
|
Add or remove chem. groups
Proteolytic cleavage Phosphorylation/dephosphorylation |
|
|
|
Neutral glycolipids
|
Cerebrosides
|
|
|
|
Neg. charged glycolipid
|
Ganglioside
|
|
|
|
Gangliosides
|
A glycolipid
Func. as antigens recognized by antibodies in immune rxns. Blood typing |
|
|
|
Important to brain and nerve cells
|
Glycolipids (cerebrosides, gangliosides)
|
|
|
|
Common membrane saturated F.A.
|
Palmitate
Stearate |
|
|
|
Common membrane unsaturated F.A.
|
Oleate
Linoleate |
|
|
|
All unsaturated F.A. in membranes are in what conformation?
|
cis
Sharp bend, or kink, in the hydrocarbon chain at every double bond. Do not pack tightly as a result. |
|
|
|
Flippases
|
Smooth ER
Phospholipid translocator Catalyze the flip-flop of membrane lipids from one monolayer to the other. |
|
|
|
Tm
|
Membrane transition temp
Membrane must be above Tm value, or in its fluid state, to func. properly |
|
|
|
Low Tm
|
More fluid
Tm decreases w/ incr. in double bonds, or unsaturation. |
|
|
|
Cholesterol and Tm
|
Decr. membrane fluidity at temp above the Tm
Incr. fluidity at temp. below the Tm. |
|
|
|
Homeoviscous adaptation
|
Regulation of membrane fluidity.
Important for org. that control body temp. Alteration of membr FAs w/ change in temp Temp drop act’n enzyme cleave C’s from FA’s shorter chains lower Tm more fluid Temp drop incr’d desaturase enzyme intro db into FA’s decr’d sat’n lower Tm more fluid |
|
|
|
Homeoviscous adaptation and O2
|
Desaturase also responds to O2 changes with temp
O2 is substr of desaturase Temp drop incr’d O2 incr’d desaturase activity intro db into FA’s decr’d sat’n lower Tm more fluid Impt to plants, hibernating animals |
|
|
|
Membrane protein categories
|
Integral
Peripheral Lipid anchored |
|
|
|
Integral membrane proteins
|
Amphipathic
Hydrophobic regions embedded w/in the memb. Difficult to remove |
|
|
|
Molecules transported via simple diffusion
|
Small nonpolar molecules
Oxygen CO2, ethanol |
|
|
|
Transport proteins
|
Integral memb. proteins that recognize substances w/ great specificity and speed their movement across the memb.
|
|
|
|
Faciliated diffusion
|
Transport proteins move solutes down their free E gradient in the direction of thermodynamic equil.
Also passive transport |
|
|
|
The movement of an ion is determined by its what?
|
electrochemical potential.
|
|
|
|
Electrochemical potential
|
The sum of its conc. gradient and the charge gradient across the memb.
Faciliated diff - exergonic movement Active transport - endergonic movement. |
|
|
|
Neg. Vm
|
Most cells have a neg. membrane potential (Vm).
Excess of neg. solutes inside the cell. |
|
|
|
Osmosis
|
Water tends to move from regions of lower solute conc. (higher free E) to regions of higher solute conc. (lower free E).
|
|
|
|
Direct A.T.
|
Accumulation of solute molecules or ions on one side of the memb. is coupled directly to an exergonic chem rxn, most commonly the hydrolysis of ATP.
|
|
|
|
Indirect A.T.
|
Simultaneous transport of 2 solutes.
One solute drives the unfavorable movement of other solute up its gradient. Symport or antiport. |
|
|
|
V type ATPase
|
H+ pumped into vesicles, Golgi
Keeps pH low activating hydrolytic enzymes |
|
|
|
What organelles compose the endomembrane system of a eukaryotic cell?
|
Endoplasmic reticulum, Golgi complex, endosomes, lysosome (but NOT peroxisomes).
|
|
|
|
Endoplasmic Reticulum (ER)
|
A cont. network of flattened sacs, tubules, and associated vesicles that stretch throughout the cytoplasm.
50-90% total mammalian membrane. Important for plasma memb. synthesis. Synthesis of proteins for export from the cell. Biosynthesis of lipids. |
|
|
|
Rough ER
|
Ribosomes on cytosolic side.
Subdomain = transitional elements - shuttle lipids and proteins from the ER to Golgi. |
|
|
|
Transitional elements
|
Subdomain of rough ER.
Important role in formation of transitional vesicles - shuttle lipids and proteins from ER to Golgi complex. Resemble smooth ER. |
|
|
|
Are the rough ER and smooth ER seperate organelles?
|
No.
Their lumenal spaces are cont. Material can travel b/t the rough and smooth ER w/o vesicles. |
|
|
|
ER cisternae
|
Membrane-bound sacs w/in the ER.
Enclose a space called the ER lumen. |
|
|
|
ER lumen
|
Space enclosed by ER cisternae.
Of the total memb. in a mammalian cell, up to 50-90% surrounds the ER lumen |
|
|
|
Rough ER functions
|
Ribosomes attached to cytosolic side synth. membr-bound and soluble proteins for the endomembrane system.
Initial steps of addition of carbohydrate groups to glycoproteins. Folding of polypeptides, removal of misfolded polypeptides. |
|
|
|
Smooth ER functions
|
Drug detoxification
Steroid biosynthesis Carbohydrate metabolism Calcium storage |
|
|
|
Drug detoxification
|
Smooth ER
Rxn common to most pathways for drug detoxification and steroid biosynth. is hydroxylation. Catalyzed by cytochromes P450 proteins. Addition of hydroxyl groups to organic acceptor molecules making them more soluble and easier to excrete from the body. |
|
|
|
Hydroxylation
|
Drug detox and steroid biosynth. in smooth ER.
Enzyme catalyzed addition of hydroxyl groups makes more soluble and easier to excrete. Catalyzed by cytochrome P450 proteins. e- transport system transfers e-'s from either reduced coenzymes NADPH or NADH to a cytochrome P450 protein. This reduced P450 donates e- to O2. |
|
|
|
Carbohydrate metabolism
|
Smooth ER function
Lots of glu.-6-phosphatase in hepatocyte smooth ER memb. Liver important to maintaing blood glu. levels. Enzymatic breakdown of stored glycogen |
|
|
|
Carbohydrate metabolism rxn
|
glu.-6-phosphate + H2O --> glu. + Pi
|
|
|
|
Carbohydrate metabolism mechanism
|
When glu. needed by the body, liver glycogen is broken down by phosphorolysis producing glu.-6-phosphate.
Memb. impermeable to phosphorylated sugars so must be converted to free glu. by glu.-6-phosphatase. Free glu. leaves the liver cell via GLUT-2. Glu.-6-phosphatase activity present in liver, kidney, intestine, but NOT muscle or brain cells - retain glu.-6-phosphate for their own E needs. |
|
|
|
Specializes in calcium storage
|
sarcoplasmic reticulum found in muscle cells.
type of smooth er |
|
|
|
steroid biosynthesis (p.p)
|
smooth er
many pathway enzymes are p450's. much smooth er in adrenal, leydig cells, liver, ovary, plastids in plants |
|
|
|
carbohydrate metabolism (p.p)
|
much glu.-6-phosphatase in hepatocyte smooth er memb.
liver impt to maintaining blood glu. levels. GLUT2 glu. transporters |
|
|
|
steroid biosynthesis (book)
|
smooth er site of biosynth. of cholesterol and steroid hormones: cortisol, testosterone, estrogen.
Hydroxymethylglutaryl-CoA reductase (HMG-CoA reductase) is committed step in cholesterol biosynth. present in large amts in liver cells. this enzyme targeted for inhibition by statin drugs. p-450 monooxygenases impt in synth of cholesterol and its conversion to steroid hormones by hydroxylation |
|
|
|
HMG-CoA reductase
|
hydroxymethylglutaryl-CoA reductase
committed step of steroid biosynthesis. targeted for inhibition by statin drugs. |
|
|
|
Calcium storage (p.p)
|
sarcoplasmic reticulum - specialized smooth er in muscle cells.
much ca-binding protein in lumen atp dependent ca-ATPases pump ca into er - signals from neurotransmitter binding |
|
|
|
Most membr. lipids and proteins synthesized where.
|
endoplasmic reticulum
|
|
|
|
Membrane biosynth (p.p)
|
most membr lipids and proteins synth. in er.
lipids produced and incorporated into inside layer of bilayer phospholipid translocators (flippases) catalyze transverse diffusion. particular enzymes for particular phospholipids - membr. asymmetry - cont. when er vesicles merge w/ plasma membr. |
|
|
|
Membrane biosynthesis (book)
|
er is primary source for phospholipids and cholesterol.
biosynth and incorporation of membr phospholipid molecules restricted to monolayer of er membr facing the cytosol. cellular membr are phospholipid bilayers - so phospholipid translocators, flippases, catalyze translocation of phospholipids thru er membr. |
|
|
|
phospholipid translocators
|
also flippases
specific so type of phospholipid molecules transferred across a membr depends on translocator present - contributes to membr asymmetry. |
|
|
|
cytosolic phosphlipid exchange proteins
|
movement of phospholipids from er to mitochondria, chloroplast, or peroxisome poses a problem - these organelles do not grow by fusion w/ er derived vesicles (endomembr organelles do)
so these proteins convey phospholipid molecules from er membr to outer mitochondrial membr. specific also contribute to movement to plasma membr. |
|
|
|
golgi complex (p.p.)
|
linked w/ er
membr and protein trafficking er glycoproteins further processed er glycoproteins, membr lipids sorted and packaged for export flattened, stacked sacs (cisternae) surrounded by transport vesicles. |
|
|
|
golgi complex (book)
|
endomembr organelle
central role in membr and protein trafficking in eukaryotic cells series of flattened membr bounded cisternae - disc shaped sacs that are stacked together (3-8 sacs) intracisternal space is golgi's lumen and part of endomembr system's network of internal spaces. both er and golgi surrounded by numerous transport vesicles |
|
|
|
golgi complex faces
|
each golgi stack has 2 distinct sides, or faces.
cis oriented toward er (CGN) - vesicles w/ newly synth. lipids and proteins from er fuse w/ CGN membr. trans (TGN) - proteins and lipids leave in transport vesicles that cont. bud from tips of TGN cisternae, carry lipids and proteins to secretory granules, endosomes, lysosomes, plasma membr. b/t CGN and TGN - medial cisternae where protein processing occurs. |
|
|
|
medial cisternae of golgi complex
|
b/t cis golgi network and trans golgi network
much of protein processing occurs here. |
|
|
|
medial cisternae, CGN,and TGN of golgi complex
|
biochemically and fucntionally distinct.
each compartment contains specific receptor proteins and enzymes necessary for specific steps in protein and membr processing biochemical polarity |
|
|
|
stationary cisternae model (p.p)
|
explains movement of lipids and proteins from cgn to tgn via medial cisternae of golgi.
shuttle vesicles move b/t areas vesicles bud from cisterna, fuse w/ next cisterna, cis tot rans - molecules may remain in lumen |
|
|
|
stationary cisternae model (book)
|
each compartment of golgi stack is a stable structure
trafficking b/t cisternae mediated by shuttle vesicles that bud from cisterna and fuse w/ next cisterna in cis to trans sequence. proteins destined for TGN are carried forward by shuttle vesicles. mol. that belong in er and golgi compartments are actively retained or retrieved. |
|
|
|
cisternal maturation model (book)
|
golgi cisternae are transient compartments that gradually change from CGN cisternae thru medial cisternae to TGN cisternae.
transition vesicles from er converge to form CGN cis cisterna transformed to med. cisterna then to trans cisterna as additional enzymes acquired. |
|
|
|
two models that depict flow of lipids and proteins thru golgi complex
|
cisternal maturation model
stationary cisternae model which model may depend on cell type both models: membr lipids and proteins move er - golgi - plasma membr. both models: TGN forms transport vesicles or secretory granules w/ cargo targeted for various destination beyond the golgi. |
|
|
|
anterograde transport
|
the movement of material from er thru golgi complex toward plasma membr.
|
|
|
|
retrograde transport
|
flow of vesicles from golgi cisternae back toward the er
func. to balance the flow of lipids toward plama membr and to ensure a supply of components for forming new vesicles - cell recycles lipids and proteins no longer needed during the late stages of anterograde transport |
|
|
|
protein glycosylation (p.p)
|
addition of CH chains to prot's in er, golgi
N-linked via asparagine O linked via serine, threonine |
|
|
|
protein glycosylation (book)
|
addition of carbohydrate side chains to specific a.a residues of prot's forming glycoproteins.
N-linked and O-linked glycoslyation |
|
|
|
N-linked glycosylation (p.p)
|
cytosolic er membr surface
dolichol phosphate inserted into er membr. 2 GlcNAc added to dolichol PO4 Core oligosaccharide of 99 monosaccharides Flippase moves growing CH chain to lumen more mannose added, glu added - shuttled by dolichol phosphate core oligosacch transferred to asp modification - glucosidases, mannosidases first Glc signals interaction of glycoprot w/ er prot's proper folding sensed by er glucosyl transferase terninal glycosylations in golgi |
|
|
|
N-linked glycosylation (book)
|
dolichol phosphate inserted into er membr - core glycosylation
carb. groups added to po4 group of dolichol po4. first 2 groups added are N-acetylglucosamine (GlcNAc) Next, 5 mannose groups added. Translocation of growing core oligosaccharide from cytosol to er lumen by flippase. inside er lumen, dolichol po4 carriers bring mannose and glu units that are added to specific positions completed core oligosaccharide is then transferred from dolichol to asparagine residue of recipient prot - catalyzed by oliosaccharyl transferase. 3 glu units and on mannose removed by glucosidases and mannosidases. usually core oligosacch is added to prot as polypeptide is being synth by ribosome bound to er membr. - cotranslational glycosylation calnexin and calreticulin promote disulfide bond formation forming ERp57. further processing in golgi - prot move from cis thru medial to trans face of golgi stack - terminal glycosylations. terminal glycosylation includes removal of a few carb units of core oligosacch. |
|
|
|
initial steps of n glycosylation take place where
|
cytosolic side of er membr.
|
|
|
|
glycosylation (book 12-6)
|
biosynth of core oligosacch for n linked glycosylation of certain asparagine residues
initial processing of core oligosacch. identification and removal of misfolded proteins. attachment of n-acetylgalactosamine to serine or threonine. 1st step of phosphorylationo f lysosomal prot's removal of mannose 2nd step of phosphorylation of lysosomal prot's removal of mannose and attachment of n-acetylglucosamine addition of galactose and sialic acid addition of sialic acid and attachment of sulfate to tyrosine |
|
|
|
protein trafficking (p.p)
|
prot's from er must go to correct site, stay there or be excreted - targeting tags on prot's, tags on lipids of vesicle membr's
sorting - early in er, cgn, med. cisternae, later sorting in tgn (packaging) prot's impt to remain in er prot's impt to remain in golgi may have tags, may form large complexes (can't escape) |
|
|
|
protein trafficking (book)
|
once a prot reaches an organelles where it is to remain, must be mechanism to prevent it from leaving.
each prot contains a specific tag - a.a sequence, oligosacch. side chain, hydrophobic domain memb lipids may also be tagged to help vesicles reach proper destination - attach po4 groups to 3, 4, or 5 of memb phosphatidylinositol (PI) mol by specific kinase |
|
|
|
Overview of protein trafficking
|
Prot's synth by ribosomes attached to cytosolic surface of rough er - initial glycosylation occur in er lumen
transition vesicles carry newly synth lipids and glycosylated prot's to CGN Lipids and prot's move thru cisternae to TGN At TGN, ves. bud of to form secretory ves or endosomes secretory ves. move to plasma memb prot's taken into cell by endocytosis form endocytic ves that fuse w/ early endosomes. components not destined for digestion after endocytosis are recycles to plasma memb. endosomes w/ material for digestion mature to form late endosomes then lysosomes retrograde traffic returns prot's to earlier compartments |
|
|
|
Protein glycosylation
|
addition of CH side chains to specific a.a. residues of prot's
strict dependence on each step in previous modification |
|
|
|
N-linked glycosylation
|
via asparagine
|
|
|
|
O-linked glycosylation
|
via serine, threonine
|
|
|
|
oligosaccharyl transferase
|
catalyzes the transfer of a complete core oligosaccharide to asparagine of recipient prot.
|
|
|
|
catalyzes the transfer of a complete core oligosaccharide to asparagine of recipient prot
|
oligosaccharyl transferase
|
|
|
|
how many glu. and how many mannose removed when core oligo is being trimmed and modified? what enzymes perform this func?
|
3 glu
1 mannose glucosidases and mannosidases |
|
|
|
when is core oligosaccharide added to a prot?
|
when the polypeptide is being synthesized by a ribosome bound to the er membr.
cotranslational glycosylation |
|
|
|
cotranslational glycosylation
|
core oligo usually added to prot as the polypeptide is being synthesized by a ribosome bound to the er membr.
promotes proper prot folding |
|
|
|
what does cotranslational glycosylation help do
|
promote proper prot folding
|
|
|
|
proper folding of a newly synth glycoprot is sensed by what enzyme?
|
er glucosyl transferase
|
|
|
|
ERAD
|
er-associated degradation - if a improperly folded glycoprot is not corrected
|
|
|
|
if glycoprot not properly folded, what can happen to correct it?
|
add glu
|
|
|
|
promotes disulfide bond formation
|
calnexin
calreticulin |
|
|
|
catalyzes disulfide bond formation
|
ERp57
|
|
|
|
ERp57
|
catalyzes disulfide bond formation
|
|
|
|
terminal glycosylations occur where?
|
golgi
|
|
|
|
microtubules (MT)
|
largest structures
2 types: cytoplasmic, axonemal |
|
|
|
2 types of microtubules
|
cytoplasmic
axonemal |
|
|
|
cytoplasmic microtubules
|
maintain polarized shape
orientation of cellulose microfibrils in plants form mitotic and meiotic spindles |
|
|
|
axonemal microtubules
|
highly organized
found in cilia, flagella - struct's impt to cell movement |
|
|
|
microtubule structure
|
straight hollow cylinders
protofilaments organized longitudinally. 13 around lumen is most common |
|
|
|
basic subunit of protofilaments
|
heterodimer of tubulin prot
|
|
|
|
alpha/beta tubulins
|
have GTP binding site at N-terminus
domain at C-terminus which interacts with MAP's all tubulin dimers oriented in same direction - gives polarity |
|
|
|
why are tubulin dimers oriented in the same direction
|
gives polarity
|
|
|
|
how are microtubules formed?
|
reversible polymerization of tubulin dimers
|
|
|
|
microtubule formation
|
reversible polymerization of tubulin dimers
requires GTP, Mg++ aggregation of tubulin dimers into clusters = oligomers oligomers serve as nuclei from which new MT's can grow = nucleation then elongation - addition of subunits at either end. |
|
|
|
What is needed for microtubule formation?
|
GTP, Mg++
|
|
|
|
importance of GTP in microtubule formation
|
each tubulin heterodimer binds 2 GTP mol's.
forms GTP cap - protects MT from subunits coming off plus end, provides a tip to which further dimers can be added. hydrolysis of GTP by beta tubulin results in unstable tip |
|
|
|
in microtubule formation, what hydrolysis of GTP result in?
|
unstable tip which can lead to depolymerization
GTP - GDP - unstable tip - depolymerization |
|
|
|
what hydrolyzes GTP in MT formation?
|
beta tubulin
|
|
|
|
microtubule originate from where?
|
microtubule-organizing center (MTOC)
regulated, ordered construction |
|
|
|
microtubule organizing center (MTOC)
|
MT's originate from here
anchors one end of MT centrosome acts as MTOC during cell division (interphase) - near nucleus w/ 2 centrioles associated. |
|
|
|
centrioles
|
symmetrical - walls formed by 9 pairs of triplet MT's
Impt to formation of basal bodies basal bodies impt for formation of cilia, flagella NOT impt in plant cells |
|
|
|
most impt role of MTOC
|
its ability to nucleate and anchor MT's
|
|
|
|
how does MTOC limit number of MT"s
|
it has a limited number of nucleation and anchorage sites
|
|
|
|
microtubule associated prot's (MAP's)
|
bind at regular intervals along MT wall - allows for interaction w/ filaments and other cellular struct.
incr MT stability and affect MT bundle density impt in brain - MT bundels denser in axons neurofibrillary tangles |
|
|
|
neurofibrillary tangles
|
from dysfunction of MAP's
dense tangles of neurites can lead to alzheimers, palsy. |
|
|
|
microfilaments (MF's)
|
smallest of cytoskeletal filaments
impt to movement - cell migration, amoeboid movement produce cleavage furrows that divides the cytoplasm of animal cells help maintain cell shape and development cell cortex structural core of microvilli |
|
|
|
cell cortex
|
dense network of MF's
|
|
|
|
form structural core of microvilli
|
microfilaments
|
|
|
|
actin
|
once synthesized, folds into U shaped mol w/ central cavity that binds ATP or ADP.
|
|
|
|
individual actin mol's
|
G-actin (globular actin)
|
|
|
|
G-actin (globular actin)
|
individual actin mol's
G-actin polymerized to form MF's - now F-actin |
|
|
|
G-actin polymerizes into what?
|
microfilaments of F-actin
G-actin monomers polymerize reversibly |
|
|
|
why is polarity impt in MF's
|
allows for independent regulation of actin assembly and disassembly at either end of the filament.
|
|
|
|
microfilament structure
|
ends of growing MF have ATP-F-actin
but bulk of MF has ADP-F-actin |
|
|
|
receptor
|
cellular struct, usually a prot, that receives a signal from another cell
embedded in cell membr cytosolic or nuclear |
|
|
|
ligand
|
mol that binds cellular receptor
|
|
|
|
signal transduction
|
cell's translation of receptor-ligand interaction
changes in cell behavior or gene expression |
|
|
|
second messenger
|
cellular mol's produced when receptor/ligand bind
|
|
|
|
01: In what step does most eukaryotic regulation take place?
|
TRANSCRIPTION
|
|
|
|
cell growth is generally accompanied by:
|
cell division
|
|
|
|
02: If both an activator and repressor are working, which usually wins out?
|
the repressor
|
|
|
|
M phase
|
Point in cell cycle when the cell actually divides
involves 2 overlapping events in which the nucleus divides 1st and the cytoplasm 2nd |
|
|
|
03: Where does RNA Pol bind?
|
the promoter
|
|
|
|
Phase in cell cycle when the cell actually divides.
|
M phase
|
|
|
|
04: How is the eukaryotic regulatory region different from prokaryotes?
|
it is often very far (thousands of BP) away from the promoter
|
|
|
|
growth phase b/t divisions
|
interphase
|
|
|
|
05: How does a regulatory specify which gene it works on?
|
BOUNDARY ELEMENTS (aka INSULATORS): regions that define where a gene is. will have proteins bound to them
|
|
|
|
interphase
|
growth phase b/t cell divisions
most cellular contents are synthesized cont. during interphase = incr in cell mass |
|
|
|
06: Does transcription in eukaryotes usually require one regulatory element?
|
NO: usually needs lots to initiate transcription
|
|
|
|
S phase
|
specific portion of interphase
new nuclear DNA is synthesized |
|
|
|
07: What helps us learn about gene eukaryotic regulation? One example
|
REPORTER GENE:
a gene whose activity is easily measured. ie β Galactosidase: can add a protein which will change color depending on how much of it is available. if you put in a regulatory region of another gene that will instead act on your reporter, you can see where those genes act. |
|
|
|
seperates S phase from preceding M phase
|
G1 phase
|
|
|
|
08: What is the structure of eukaryotic regulatory proteins?
|
often DIMERS:
have separate BINDING and ACTIVATION domains |
|
|
|
seperates the end of S phase from the onset of the next M phase.
|
G2 phase
|
|
|
|
09: How can we have variation in eukaryotic regulatory proteins?
|
HOMODIMERS: same dimers.
HETERODIMERS: different dimers together, gives larger possibilities for variation |
|
|
|
overall length of cell cycle
|
generation time
|
|
|
|
10: Where do eukaryotic regulatory proteins bind?
|
ACTIVATING SEQUENCES: can be upstream or downstream. can have multiple binding sites
|
|
|
|
nuclear DNA doubles in what phase?
|
S phase of interphase
|
|
|
|
11: what did we learn from the DOMAIN SWAP EXPERIMENT?
|
It showed us that both the activating and binding regions of the regulatory protein are needed.
Activating regions can bind to other binding regions to activate the binding region's gene. |
|
|
|
phase in which a "decision" is made as to whether the cell is to divide again
|
G1 phase
|
|
|
|
12: What are some binding motifs?
|
HOMEODOMAIN: single subunit...esp helix-turn-helix
ZINC FINGERS: charged, will interact w/ negative DNA backbone LEUCINE ZIPPER: dimer. each part could have different binding domains. HELIX-LOOP-HELIX |
|
|
|
cells that become arrested in G1 phase, awaiting a signal that will trigger reentry into the cell cyle
|
G0 phase
|
|
|
|
13: What is a common mechanism for activating proteins? What is the significance?
|
activator brings in MEDIATOR: holding onto RNA Pol via many NON-SPECIFIC SEQUENCES...means that it can work with several different activators.
mediators are often positively charged |
|
|
|
G0 phase
|
cells that become arrested in G1 phase, awaiting a signal that will trigger reentry into the cell cycle and a committment to divide.
|
|
|
|
14: Is there a holoenzyme for eukaryotic activation?
|
it is debatable, we think it only forms in vitro.
|
|
|
|
terminal differentiation
|
cells exit from cell cycle entirely, destined to never divide again.
most nerve cells are in this state. |
|
|
|
15: What is one way the mediator can act?
|
can push the LexA protein (bound to Rna Pol) onto lexA site on DNA
|
|
|
|
semiconservative replication
|
half of parent mol is retained by each daughter mol
|
|
|
|
16: What is a very important overall eukaryotic activation/repression mechanism? How does it act?
|
NUCLEOSOME REMODELING:
activator protein binds, brings in acetyl transferase, expands chromatin to 10nm fiber OR: activator recruits chromatin remodeling complex, moves DNA around histones |
|
|
|
replication of circular DNA
|
begins at a single origin
proceeds bidirectionally around the circle w/ 2 replication forks moving in opp. directions. process generates intermediates that resemble theta = theta replication membr growth b/t attachment sites of 2 replicating copies move daughter chromosomes toward opp sides of cell. |
|
|
|
17: How do boundary elements work?
|
(aka insulator region)
they act like a FENCE: stops enhancer from turning on a promoter of the wrong gene. it can also act as a boundary for CHROMATIN MODIFYING: will stop it from going further down the chromatin |
|
|
|
replicons
|
multiple replication units
DNA of eukaryotic chromosomes is initiated at these multiple sites. maybe thousands |
|
|
|
18: what is an LCR?
|
LOCUS CONTROL REGION: lets you turn on access to a number of genes.
ie β Globulin: you have several different types. LCR turns on access to all the genes, which can then be individually activated. like turning on the power to the entire building, then flipping switches in individual rooms. |
|
|
|
origin of replication
|
special DNA sequence at the center of each replicon.
where DNA synthesis is initiated by a mechanism involving several groups of initiator prot's. |
|
|
|
19: Are genes usually on or off?
|
OFF: most need an activator to turn them on since they have a poor promoter
|
|
|
|
ensures only one copy of each chromosome
|
licensing
|
|
|
|
20: what is the importance of SIGNAL INTEGRATION?
|
there are many different activators and repressors all acting at once.
the summation of their effects together is often much more powerful than summing them individually. you can also have one protein act as both a positive and negative signal, depending on concentration gradient. |
|
|
|
cyclin-dependent kinase (Cdk)
|
functions both in activating DNA synth. at licensed origins and in ensuring these same origins can't become licensed again.
|
|
|
|
21: basic mechanisms of cooperative binding(4)
|
1: A+B need each other to work as activators
2: A+B need to bind to a third molecule in between them to act as activators 3: A works, blocking B, but then brings in a chromatin modifier so that B can bind. 4: A binds, unwinding DNA so that B can bind. |
|
|
|
geminin
|
prot that inhibits relicensing.
blocks binding of MCM prot's to DNA. |
|
|
|
22: how is the HO gene controlled?
|
three molecules of SWI5 bind to sites
allows CHROMATIN REMODELERS to bind makes the SBF site available, 3 molecules SBF bind to activate gene |
|
|
|
Catalyzed DNA elongation
|
DNA polymerase
|
|
|
|
23: How is the human interferon protein regulated?
|
HMG proteins bend the DNA in a "U" shape so that other proteins can bind.
ENCHANCEOSOME complex forms with several proteins, all of which are needed to turn on the gene. this complex forms in many organisms |
|
|
|
joins DNA fragments
|
DNA ligase
|
|
|
|
24: What is COMBINATORIAL CONTROL?
|
one protein regulating many genes in combination with other proteins (ie CAP)
ie one gene needs activators 1, 2, 3, 4 another gene needs 3, 5, 6 |
|
|
|
strand synthesized as a cont. chain in the 5'-3' direction
|
leading strand
|
|
|
|
25: are most activators specific?
|
some are TISSUE SPECIFIC,
most are NOT gene specific |
|
|
|
lagging strand
|
must grow in the 3'-5' direction
formed as a series of short, discontinuous Okazaki fragments that are synth. 5'-3' b/c DNA polymerase can't add nucleotides in 3'-5' direction. fragments joined by DNA ligase to make a cont. new 3'-5' strand. |
|
|
|
26: What is an example of gene control in S. Cervisae (yeast)
|
mostly haploid, but when stressed will become diploid
MCM1 protein acts as an activator in certain cells, repressor in others, depending on the cell protein that it binds to. |
|
|
|
Primase
|
synth. RNA fragments a/b 10 bases long using DNA as a template.
can initiate RNA synth from scratch by joining 2 nucleotides together. |
|
|
|
27: ways repressors work (4)
|
COMPETITION: both trying to bind to overlapping sites
INHIBITION: both binds, but repressor occludes the activator's activating site DIRECT REPRESSION: repressor interacts w/ mediator to stop transcription INDIRECT REPRESSION: interacts w/ deacetylase to transform chromatin to 30nm fiber |
|
|
|
Primers
|
RNA primers needed to initiate DNA synth.
synth'd by primase |
|
|
|
28: What is an example of yeast genetic repression?
|
Gal1 gene:
Mig1 binds to Mig1 site. Tup1 binds to Mig1 protein, turns off gene. overrides UAS which promotes gene |
|
|
|
DNA helicase
|
prot responsible for unwinding DNA.
uses E derived from ATP hydrolysis |
|
|
|
29: What is a second Yeast repressor?
|
Gal80, occludes Gal4 activator dimer.
binds to activator when no galactose. |
|
|
|
topoisomerases
|
relieves supercoiling
DNA gyrase=type II topoisomerase (cuts both DNA strands) |
|
|
|
30: How can we silence genes at the chromatin level?
|
esp a TELOMERE:
RAP protein binds to telomere SIR2 binds to it, then SIR 3, 4, which bind to chromatin across it keeps telomeres condensed always. it becomes HETEROCHROMATIN |
|
|
|
single-stranded DNA binding prots (SSB)
|
keep DNA strand separated
|
|
|
|
31: What is a long term way to silence genes?
|
DNA METHYLATION: blocks activation in the long term.
proteins bind methylated DNA which blocks transcription oven more. |
|
|
|
Central Dogma of Molecular Biology
|
the concept of directional flow of genetic information
Francis Crick |
|
|
|
32: What is another way DNA methylation could act?
|
could block a BOUNDARY ELEMENT SITE: then an enhancer will act across it and turn on the wrong gene.
|
|
|
|
Transcription
|
synth. of an RNA mol using DNA as a template
|
|
|
|
33: when do you use the CHIP ASSAY?
|
to amplify sections of DNA that a certain protein binds to
can do it without knowing the sequences |
|
|
|
mRNA
|
RNA that is translated into a prot
|
|
|
|
34: How does the CHIP assay work?
|
purify DNA (or chromatin) and shear it into fragments. can use entire chromosome
then add a binding protein, and use an antibody against it. antibody will make a large chain from the protein, which will precipitate in solution. skim rest off, centrifuge to form a pellet. you can remove the proteins and antibodies, and clone or PCR the DNA fragments to have pieces of all the genes that the binding protein binds to. |
|
|
|
RNA that is translated into a protein:
|
mRNA
|
|
|
|
Exceptions to Central Dogma
|
Retroviruses - "backward" flow of genetic info, Ex. HIV
RNA viruses - RNA genome is template for RNA synth., Ex. influenza, hepatitis C |
|
|
|
The genetic code is dependant upon:
|
the relationship b/t:
DNA nucleotide sequence. linear order of a.a.'s in prot. mol. |
|
|
|
triplet code
|
3 base pairs in double-stranded DNA required to specify each a.a. in a polypeptide.
|
|
|
|
gene amplification
|
multiple copies of same gene.
by repl. of DNA in specific chromosomal region |
|
|
|
Genomic control mechanisms
|
gene amplification
gene deletion gene rearrangement DNA methylation |
|
|
|
5 main levels gene expression is regulated:
|
the genome
transcription RNA processing and export from nucleus to cytoplasm. translation post-translational events |
|
|
|
why can gene amplification be regarded as an example of genomic control?
|
it is a regulatory change in the make-up of structural organization of the genome.
|
|
|
|
genomic control by gene amplification
|
creates mult. copies of same gene.
|
|
|
|
gene deletion
|
a means of genomic control
some cells delete genes whose products are not required. occurs in RBC's - discard entire nuclei after sufficient hemoglobin mRNA has been made. |
|
|
|
DNA rearrangement
|
movement of DNA segments from one location to another w/in the genome.
used by lymphocytes to prod. antibody mol's |
|
|
|
antibodies
|
prot's composed of 2 polypeptide subunits - heavy chain and light chain
|
|
|
|
lymphocyte DNA rearrangement
|
lymphocytes use small # of DNA segments and rearrange them in various combinations
V, J, D, C DNA segments |
|
|
|
4 DNA sequences used in lymphocyte rearrangement
|
V, J, D, C segments
|
|
|
|
DNase 1
|
an endonuclease from pancreas.
degrades transcriptionally active DNA in chromatin - evidence that DNA is uncoiled. |
|
|
|
DNA methylation
|
genomic control
addition of methyl groups to selected cytosine bases in DNA usually at promoter region (5'). |
|
|
|
DNA methylation at promoter regions can have what effects?
|
can block acces of prot's required for transcription activation.
can serve as binding sites for prot's that condense chromatin to an inactive configuration - no gene expression |
|
|
|
histones H3, H4
|
acetylation of these histones is associated w/ gene activation
|
|
|
|
Transcriptionally active chromatin often lacks what histone and why:
|
histone H1 - keeps chromosome uncoiled.
|
|
|
|
Flow of genetic information
|

|
|
|
|
The nucleus
|
Location of
Chromosomes Replication Transcription Double-membrane structure Nuclear envelope Perinuclear space between inner, outer membranes |
|
|
|
outer nuclear membrane
|
Continuous w/ ER
Perinuclear space continuous w/ ER lumen Ribosomes Cytoskeleton anchoring proteins bind actin, intermediate filaments Movement Nucleoplasmic reticulum – projections Contacts inner membrane |
|
|
|
nuclear pores
|
Connect cytosol w/ nucleoplasm
Specialized channels 3000-4000 Fuse inner/outer membranes |
|
|
|
nuclear pore complex
|
Impt to transport
Need enzymes, proteins, precursors for replication/ transcription Move mRNA, ribosomal subunits to cytosol |
|
|
|
nucleoporin protein complexes
|
Octagonal symmetry
Protrude into cytosol and nucleoplasm Wheel-like structure <30,000 MW mol’s move between “spokes of wheel” Center = “transporter” |
|
|
|
transport through nuclear pores
|
Active
Energy Specific binding to pore complex proteins Nuclear localization signals (NLS) on protein moving into nucleus 8-10 aa sequences (incl proline, lysine, arginine) Recognition Nuclear export signals (NES) on mol’s moving out of nucleus Aa sequences |
|
|
|
importin
|
binds NLS
Shuttles/chaperones protein through center of NPC Releases protein as assoc’s w/ GTP-binding protein (Ran-GTP) in nucleoplasm Importin-GTP-binding prot complex shuttled back through NPC cytosol Recycled |
|
|
|
exportin
|
similar protein needed to export mol’s out of nucleus
Impt mostly for RNA movement Also shuttles through center of NPC Uses same GTP-binding protein (Ran-GTP) |
|
|
|
import/export cycle (image)
|
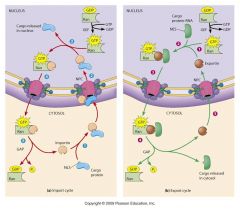
|
|
|
|
Role of Ran-GTP in import/export cycle
|
Ran-GTP concent gradient impt: [Ran-GTP] higher inside nucleus than in cytosol
Guanine nucleotide Exchange Factor in nucleus Encourages incr’d Ran-GTP over Ran-GDP GTPase Activated Protein in cytosol Encourages hydrolysis Ran-GTP Ran-GDP Equilibrium w/ high [Ran-GTP] in nucleus Promotes release of Importin from imported prot’s inside nucleus Promotes binding of Exportin to RNA to be exported from nucleus |
|
|
|
nuclear matrix
|
Protein network shapes nucleus
Organizing skeleton for nucleic acids (?) May anchor DNA/RNA during synthesis |
|
|
|
nuclear lamina
|
Fibers lining inner membrane
Intermediate filaments (lamins) Mutations give rise to diseases Nuclear shape distortions |
|
|
|
nucleolus
|
Ribosome factory
One or several 1000’s Varied sizes, shapes; usually rel large Size dependent on activity prot synthesis Contains DNA w/ genes coding for rRNA DNA Nucleolus Organizer Regions Higher organisms have 100s-1000s NOR copies Contains fibriles DNA being transcribed rRNA Contains granules rRNA packaging w/ proteins ribosomal subunits Many in cells w/ high prot synth activity Some additional activities Nuclear transport Chem mod’n RNA Cell division control |
|
|
|
DNA replication
|
Accuracy duplicate copies to daughter cells
Purine/pyrimidine pairing fits physical dimensions Strands held by H-bonds Allows separation Complementarity allows strand sequence determined by opposite strand Template Right handed helix (B-DNA) Left-handed helix can form (Z-DNA) Antiparallel strands |
|
|
|
double helix model
|
Watson/Crick, 1952
Repeating phosphate/deoxyribose units Nitrogenous base attached to each sugar A/T, G/C could hydrogen bond Franklin’s X-ray diffraction showed repeating structures “Circular staircase” |
|
|
|
major groove
|
exposes H, O, N (all can form H bonds) and T’s methyl
Regulatory proteins can bind |
|
|
|
supercoiling
|
Studied in closed, circular DNA
Bacteria, viruses, mitochondria, chloroplasts Linear DNA when anchored, cannot freely rotate Compacted chromosomes During replication, transcription Impt to interaction w/ other molecules |
|
|
|
topisomerase
|
Topoisomerases – enzymes catalyzes supercoiled relaxed DNA forms
Type I introduce ss breaks, Type II ds breaks DNA strands pass through break unwinding Type II requires ATP Resealing also catalyzed by topoisomerase |
|
|
|
DNA gyrase
|
DNA gyrase – bacterial
Can relieve supercoiling AND induce supercoiling Impt to replication |
|
|
|
DNA denaturation
|
Strand separation
H-bonds relatively weak, rel easily broken w/ incr’d heat, pH What holds each strand together?? Use UV absorbance to monitor strand sep’n Ss DNA has higher absorbance Tm = DNA melting temp = temp @ which ½ abs change found |
|
|
|
DNA melting temperature
|
Tm depends on base composition
3 H-bonds bind GC base pairs 2 H-bonds bind AT base pairs Which base pair requires more heat to separate? Incr directly proportional to # GC bp’s Proper base pairing also incr’d Tm |
|
|
|
DNA renaturation
|
Renaturation w/ lowering temp
Use UV abs to determine hybridization ability Tells complementarity of strands Can use probes to locate genes DNA-DNA DNA-RNA RNA-RNA |
|
|
|
genome
|
DNA w/ one complete copy of all genetic info of that organism
Prokaryotes, viruses – one or few linear or circular DNA mol(s) Eukaryotes – nuclear, mitochondrial and chloroplast genomes Mitoch, chloroplast – single, circular (usually) Nuclear – multiple DNA mol’s |
|
|
|
bioinformatics
|
Impt to locating genes w/in genome, determining which proteins genes code for
Much info needs computer analyses Book: <2% human genome codes for proteins; unclear function of rest |
|
|
|
transcriptomes
|
Transcriptomes – entire set of RNA mol’s prod’d by genome
Products of transcription DNA microarrays use cDNA from RNA, then DNA-DNA hybridization |
|
|
|
proteomes
|
Proteomes – all proteins produced by a genome
More complex than genome Book: human 25,000 genes 100,000’s proteins Alternative splicing Biochem modifications Protein microarrays used to study chem properties of proteins |
|
|
|
human genome
|
Human genome project studied genomes of 10 people
All humans’ genomes 99.7% identical 0.3% variable ~ 3.2 Gb SNPs CNVs |
|
|
|
single nucleotide polymorphisms
|
Single Nucleotide Polymorphisms (SNPs) – single base changes w/in genome
~1x107 SNPs explain variability Some understood due to known diseases/ dysfunctions Most not in coding regions Many related to each other When on same chromosome, inherited in sections (haplotypes) So easier to locate several at once |
|
|
|
copy number variations
|
Copy Number Variations (CNV’s)
Also impt to variability in genome DNA rearrangements, deletion, duplications Involve 100’s of Gb’s as segments of Kb’s Useful medical info, but difficult ethically |
|
|
|
tandem repeated DNA
|
Multiple copies arranged next to each other
10-15% mammalian genome Varies in length of repeat, # times repeated in succession Length 1-2000 bp; usually simple sequence (<10 bp) “Satellite” DNA Much of centromeres, telomeres Book: human telomeres 250-1500 copies TTAGGG; highly conserved among vertebrates Small satellite DNA used for DNA fingerprinting Excessive repeats found in diseases |
|
|
|
interspersed repeated DNA
|
Scattered throughout genome
100’s-1000’s bps length per unit x 100’s-1000’s of copies Transposons (transposable elements) move around Some w/ genes coding for enzymes that copying, insert these sequences elsewhere Believed impt for evolutionary adaptation May be impt in gene regulation |
|
|
|
bacterial chromosome
|
Commonly single, circular molecule
Nucleoid region Anchored to RNA, protein DNA wrapped around basic proteins bead-like packets Packets arranged in loops held by proteins Loops supercoiled in nucleoid |
|
|
|
bacterial plasmids
|
Small, circular DNA
Mostly nonessential genes + genes for plasmid repl’n Conjugation Resistance to antibiotics Secr’n antibiotics Secr’n toxic proteins Metabolic rxns Supercoiled |
|
|
|
linker DNA
|
~146 bp
Histone H1 assoc’d w/ linker |
|
|
|
looped domains
|
30-nm fibers folded
Protein scaffold assists Active DNA less tightly packed, more accessible in loops |
|
|
|
euchromatin
|
transcriptionally active DNA (less packed)
Includes most DNA in metabolically active cells Compaction ~750x |
|
|
|
heterochromatin
|
tightly packed (~15,000x compaction)
Not active transcriptionally All chromatin in dividing cells is heterochromatin |
|
|
|
organellar DNA
|
Mitochondria, chloroplasts
Also have enzymes, proteins for repl’n, transcription, translation Semiautonomous (nuclear genome needed) Symbiotic origin? No histones Circular, relatively small >90% apparently noncoding |
|
|
|
cell growth
|
Macromolecule synthesis
Increased cell volume decr’d surface/vol ratio ineffective exchange ability Cell division accompanies cell growth Increases cell # in organism OR Replaces dead/injured cells Daughter cells are genetic duplicates of parent REMEMBER: somatic cells |
|
|
|
M phase of cell growth
|
Mitosis – nuclear division
Cytokinesis – cytoplasmic division Chromatin condensed chromosomes DNA has replicated, so 2 duplicate chromosomes (sister chromatids) Stay together til cell divides Nuclear envelope dissolves Mitotic spindles guide chromosomes to poles What type of structure are mitotic spindles?? |
|
|
|
interphase
|
Rest of cell cycle; cell growth
Continuous synthesis of macromolecules S phase – nuclear DNA doubles 6-8 hours G1, G2 phases – time gaps G1 8-10 hours G2 4-6 hours G0 – temporary arrest from cell growth Await signals to reenter cell cycle (M phase 30-45 mins) |
|
|
|
eukaryotic replication
|
more complex
Larger genome; takes longer Nucleosomes must be unwound Multiple origin sites multiple replicons Center = origin of replication AT rich (why??) Initiator proteins bind DNA, unwind helix Prereplication complex “licensing” repl’n Two replication forks synth DNA in opp directions replication bubble Bubbles merge |
|
|
|
licensing
|
ensures only one copy of each chromosome
Occurs through initiator proteins (particularly MCM proteins) Requires all proteins be present MCM proteins prevented from re-binding same origination site Cyclin-dependent kinase (Cdk) prod’d at beginning of S phase Phosphorylates intiator proteins inhib’n of those proteins from re-binding Geminin made during S-phase Blocks binding of MCM prot’s to DNA |
|
|
|
DNA polymerase
|
Catalyzes DNA elongation
Energy released as NTP’s inc’d (dNMPn + dNTP dNMPn+1 + PPi) Nucleotides added to 3’ –OH end of chain as phosphodiester bond between PO4 at 5’ C of dNTP and 3’ –OH of chain Growth is 5’ 3’ |
|
|
|
primers
|
RNA found to be needed to initiate DNA synthesis
DNA polymerase can only add dNTP’s to established chain (can’t start new chain) Short RNA segments found @ initiation site Primase synthesizes Uses DNA as template Primosome in E. coli – primase + 6 proteins Recognize origination site, unwind helix |
|
|
|
primase
|
Primase in eukaryotes closely assoc’d w/ DNA polymerase
Leading strand needs 1 primer Lagging strand needs 1 primer for each Okazaki fragment Polymerase adds nucleotides til reaches next primer Primers later cleaved, dNTP’s fill in E. coli – DNA polymerase does both Ligase connects sections Best method for minimizing errors |
|
|
|
“Accessory” Enzymes Needed for Replication
|
DNA helicases
Unwinding; H-bonds broken Where are the H-bonds?? Unwinding supercoiling ATP nec E. coli – part of primosome Topoisomerases Relieves supercoils DNA gyrase – impt in E. coli; Type II Single-strand Binding Protein (SSB) Keep DNA strands separated |
|
|
|
DNA helicase
|
Unwinding; H-bonds broken
Where are the H-bonds?? Unwinding supercoiling ATP nec E. coli – part of primosome |
|
|
|
single stranded binding prot
|
Keep DNA strands separated
|
|
|
|
telomeres
|
Short repeating units at ends of each linear chromosome
TTAGGG Humans 100-1500 copies Noncoding No coding info lost w/ repl’n Telomerase catalyzes add’l copies of telomeres Telomeres protected by telomere capping proteins Bind 3’ end of DNA Loops back, base pairs w/ opp DNA strand |
|
|
|
telomerase
|
Telomerase found in germ cells, actively proliferating cells
Other cells don’t have telomerase Telomere shortening DNA coding region shortening Signals apoptosis May be impt to degenerative diseases w/ aging Telomerase dysfunction leads to similar diseases, premature aging Telomerase found in cancer cells Highly proliferative |
|
|
|
cell cycle regulation
|
General cell cycle progression over ~24 h
G1, S, G2, M Varied overall length, stage length, mitosis/cytokinesis pairing Regulated to meet cell, organism needs Cancer cell research |
|
|
|
cell cycle length
|
Stem cells, sperm cells constantly divide
~8 hours Where are adult stem cells?? Some cells never divide (example??) Liver cells, lymphocytes divide only when stimulated Most time variation is in G1 Non-dividing cells G0 Rapidly dividing may skip G1 |
|
|
|
speed G1 phase
|
Increase number of replicons shortened time for each synthesis event
What is a replicon?? Large eggs w/ much cytoplasm little time needed to replicate cytoplasm contents |
|
|
|
TOR
|
prot kinase
|
|
|
|
decision points in cell cycle
|
Cell must check for
Appropriate sequence, time for each mechanism Completion of phase before next is started Environmental conditions or signals Cell uses biochemicals at particular stages Late G1, late G2, during M (metaphase/ anaphase) |
|
|
|
decision points in cell cycle graph
|
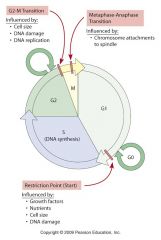
|
|
|
|
late G1 control point
|
Time varies among cell types/species
G1 inhibited to S w/ decr’d nutrients, space or w/ inhibitors Restriction point Presence of growth factors impt S phase (committed), OR G0 Need signal to G1 |
|
|
|
late G2 control point
|
Arrest here possible for long periods
Less common than at G1 Important factors Cell size DNA viability Sufficient DNA |
|
|
|
metaphase/anaphase control point
|
Movement of 2 chromosome sets 2 poles
Spindle attachment crucial May lag for “catch-up” |
|
|
|
cyclin-dependent kinsases (Cdk's)
|
Cytoplasmic molecules
Kinases commonly regulate protein activity Phosphatases must also be present (why??) Cyclins Proteins that must be present Activate kinases that regulate other proteins impt to cell cycle progression So kinases are Cdk’s Get Cdk-cyclin complexes Cyclin” refers to “cycle” Concentration in cell incr’s, decr’s at different stages of cell cycle G1 cyclins bind G1 Cdks Mitotic cyclins bind mitotic Cdks S cyclins impt for DNA repl’n Mitotic cyclin/Cdk complex |
|
|
|
mitotic cyclin/Cdk complex
|
Maturation (Mitosis) Promoting Factor (MPF)
Triggers onset of mitosis Cyclin/Cdk’s must themselves be regulated to function at appropriate time Mitotic Cdk always present during all cycle stages BUT kinase activity only when bound to mitotic cyclin Mitotic cyclin concentration low, increasing over G1, S, G2 until critical concentration at end G2 Now Mitotic Cyclin/Cdk complex mitosis |
|
|
|
activation of Cdk
|
For activity, Cdk must be specifically phosphorylated
Inhibiting kinases add 2 PO4 grps Activating kinase adds 1 PO4 grp Phosphatase removes 2 inhibiting PO4 grps Now get positive feedback loop Activated mitotic Cdk stimulates phosphatase Get more PO4 removal more activated mitotic Cdk more stimulated phosphatase Overall, more rapid activation Now protein kinase subunit active to trigger mitosis |
|
|
|
Nuclear envelope effect
|
Phosph’n lamin proteins of nuclear lamina, and other prot’s assoc’d w/ envelope
depolymerization of lamins breakdown of nuclear lamina nuclear envelope destabilized |
|
|
|
condensation of chromatin fibers
|
Activated by phosphorylation of condensin
condensin binding to DNA supercoiling compact chromosomes Assembly of mitotic spindle Phosphorylation of microtubule-assoc’d proteins |
|
|
|
active mitotic-cyclin-Cdk activates:
|
nuclear envelope breakdown, chromosome condensation, mitotic spindle formation, targeted protein degradation
|
|
|
|
G1 Cyclin/Cdks
|
Regulates progression to S phase
Nutrients Growth factors Cell size Progression commitment to cell division Signals activation G1 Cdk/Cyclin phosph’n/activation proteins Rb protein binds, inhibits E2F transcription factor E2F inactivated Genes not transcribed Genes code for proteins needed for DNA replication initiation Cells don’t replicate DNA, don’t divide Growth factor signal 2nd messenger pathway G1 Cdk/Cyclin activation Phosph’n Rb protein Releases E2F Genes transcribed DNA repl’n, cell division |
|
|
|
mitotic spindle checkpoint
|
Can halt cell cycle if improper conditions
Mitotic spindle checkpoint Check kinetochore attachment to microtubules If not attached, signal inhibits anaphase-promoting complex Cohesion proteins not degraded Sister chromatids held together |
|
|
|
DNA replication checkpoint
|
Checks for complete DNA synthesis before mitosis
Proteins impt to repl’n inhibit phosphatase no final dephosphorylation Cdk-Cyclin Cdk-Cyclin not activated No progression beyond G2 phase |
|
|
|
DNA damage checkpoints
|
Checks for viability of DNA before cell division
Altered DNA activation ATM protein kinase Phosphorylation p53 protein Protection from degradation p53 builds up in cell Activation two pathways Pathway 1 codes for p21 protein Inhibits Cdk-Cyclin activity Halts cell cycle Pathway 2 codes for Puma protein Binds apoptosis-inhibiting proteins Apoptosis signalled |
|
|
|
Cell cycle control
|
Autonomous clock
Fixed cycle repeats Due to rhythmic synthesis, degradation of cyclins that bind Cdks Cyclin/Cdk complex activate cell cycle progression External, internal cues interrupt, regulate cycle when needed Additional proteins influence Cdk’s, cyclins Phosphatases, kinases |
|
|
|
stimulatory growth factors
|
Some very specific, some signal many cell types
Only cells w/ receptors for particular factor will respond Impt during embryo development, tissue damage/wound repair Platelet-Derived Growth Factor (PDGF), Epidermal Growth Factor (EGF) |
|
|
|
growth factors commonly bind what type of receptor?
|
Growth factors commonly bind tyrosine kinase receptors
Binding receptor autophosphorylation phosphorylation of second messengers signal transduction cascade |
|
|
|
signal transdution cascade
|
Signal transduction cascade: phosphorylation events include
Raf MEK MAPK’s Mitogen Activated Protein Kinases |
|
|
|
inhibitory growth factors
|
Ex: Transforming Growth Factor b (TGFb)
May sometimes be stimulatory (cell dependent) Antagonizes EGF, PDGF stimulating effects Work through yet another cascade Smad proteins nucleus regulation of gene expression Activates gene expression coding for proteins that inhibit cell division |
|
|
|
the genetic code
|
Set of rules governing gene expression
Dependent upon relationship between DNA nucleotide sequence and Linear order of amino acids in protein molecule Genetic Code: A Triplet Code Deciphering this “code”-- major scientific breakthrough When synthesizing the polypeptide chain, mRNA is read as 3 base units—codons Each amino acid (aa) coded by one or more triplet codes |
|
|
|
properties of the genetic code
|
Unambiguous
Degenerate Nonoverlapping Almost universal |
|
|
|
code vs codon
|
CODE = genetic code expressed in DNA language
Bases A, T, C, G CODES are written 3' to 5' as they appear in the DNA template strand CODON = genetic code expressed in mRNA language Bases A, U, C, G CODONS are complementary and antiparallel to genetic code of DNA from the template strand Actual coding units read by protein synthesis mechanism |
|
|
|
bacterial RNA polymerase
|
Very well characterized (480,000 Da)
Multiple subunits: Core enzyme (2α + 1β + 1β’ subunits) + 1 sigma factor “Core" + sigma = holoenzyme Sigma ensures that RNA synthesis initiates at right place on DNA (determines correct reading frame; core only will start at random) Sigma factor readily dissociates |
|
|
|
Transcription Factor Binding of RNA Pol II
|
6 general transcription factors (TF) required for RNA Polymerase II to bind
(Nomenclature: (e.g TFIID) Roman numeral – Polymerase type (II) Letter – specific TF protein (D)) Before Pol II can initiate transcription, (1) ATP-dependent phosphorylation of Pol II (2) Pre-initiation complex needs to be released For intact cellular DNA: requires participation of other regulatory proteins that open chromatin for initiation complex to bind |
|
|
|
code vs codon
|
CODE = genetic code expressed in DNA language
Bases A, T, C, G CODES are written 3' to 5' as they appear in the DNA template strand CODON = genetic code expressed in mRNA language Bases A, U, C, G CODONS are complementary and antiparallel to genetic code of DNA from the template strand Actual coding units read by protein synthesis mechanism |
|
|
|
bacterial RNA polymerase
|
Very well characterized (480,000 Da)
Multiple subunits: Core enzyme (2α + 1β + 1β’ subunits) + 1 sigma factor “Core" + sigma = holoenzyme Sigma ensures that RNA synthesis initiates at right place on DNA (determines correct reading frame; core only will start at random) Sigma factor readily dissociates |
|
|
|
Transcription Factor Binding of RNA Pol II
|
6 general transcription factors (TF) required for RNA Polymerase II to bind
(Nomenclature: (e.g TFIID) Roman numeral – Polymerase type (II) Letter – specific TF protein (D)) Before Pol II can initiate transcription, (1) ATP-dependent phosphorylation of Pol II (2) Pre-initiation complex needs to be released For intact cellular DNA: requires participation of other regulatory proteins that open chromatin for initiation complex to bind |
|

