![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
84 Cards in this Set
- Front
- Back
|
What’s the overall ATP and NADH yield in glycolysis—from glucose to pyruvate?
|
2 ATP/ NADH
|
|
|
What’s the overall yield of ATP and NADH in the reactions from glucose to lactate?
|
2 ATP
|
|
|
When is pyruvate converted to lactate? why?
|
When we go aerobic; to regenerate NAD+
|
|
|
What does the pentose phosphate path do?
|
Make NADPH and Ribose
|
|
|
What molecule feeds into the citric acid cycle?
|
Acetyl CoA
|
|
|
glucose --> glucose-6-p
|
Irreversible, phosphoralation, uses ATP
|
|
|
glucose-6-p --> fructose-6-p
|
isomerization
|
|
|
fructose-6-p --> fructose-1, 6-bisP
|
Irreversible, phosphoralation, uses ATP
|
|
|
fructose-1, 6-bisP --> glyceraldehyde- 3p + dihydroxyacetone- P
|
aldol cleavage
|
|
|
dihydroxyacetone- P -->glyceraldehyde-3P
|
isomerization
|
|
|
glyceraldehyde- 3P --> 1,3 bisP- glycerate
|
oxidation, phosphorylation uses no ATP because the oxidation provides the energy, this is also the NADH making step.. glycerate is a carboxylic acid
|
|
|
1,3 bisP- glycerate -->3P-glycerate
|
substrate level phosphorylation, makes ATP
|
|
|
3P-glycerate --> 2P-glycerate
|
isomerization
|
|
|
2P-glycerate --> PEP
|
dehydration
|
|
|
PEP --> pyruvate
|
ATP forming, substrate level phosphorylation
|
|
|
what are 2 molecules that are allosteric effectors for glycolysis reactions?
|
ATP- inhibit glycolysis, AMP- stimulate
|
|
|
in yeast, under anaerobic conditions pyruvate -->? why does this reaction occur?
|
pyruvate goes to acid aldehyde, which goes to ethanol; regenerate NAD+
|
|
|
what is gluconeogenesis?
|
making glucose from non carbohydrate precursors
|
|
|
what is the net reaction of gluconeogenesis?
|
starting with pyruvate.. 2 pyruvate, 2 NADH, 6 nucleoside triphosphates
|
|
|
what is the cori cycle?
|
when exercising muscle goes anaerobic, you make lactate, the lactate goes to the liver where it makes glucose from it and sends it back to the muscle.
|
|
|
what is glycogenolysis?
|
breakdown of glycogen
|
|
|
what is the product of glycogenolysis?
|
immediate product is glucose- 1P, has alpha 1-4 branches, phosphate attack on the 1 carbon.
|
|
|
hormones this pathway and how? what hormone deactivates?
|
glucagon when in low sugar conditions, and epinephrine. insulin to deactive the path
|
|
|
glycogenesis?
|
making glycogen
|
|
|
when glucose is oxidized or reduced energy is? (delta G is below zero)
|
released
|
|
|
NADH is a high or low energy molecule?
|
High
|
|
|
what is oxidation?
|
removing hydrogen atoms or electrons..
|
|
|
what pathways are required to convert glucose --> carbon dioxide?
|
glycolysis.. takes glucose to pyruvate then one reaction to make it acetyl CoA, then the citric acid cycle..
|
|
|
hormones this pathway and how? what hormone deactivates?
|
glucagon when in low sugar conditions, and epinephrine. insulin to deactive the path
|
|
|
what reactive species is added to the growing chain?
|
glucose --> glucose- 6-P -->glucose- 1P--> UDP glucose added to form an alpha 1-4 branch
|
|
|
what hormones activate or deactivate the glycogenesis path?
|
insulin activates, glucagon deactivates
|
|
|
what processes convert NADH or FADH2 --> ATP?
|
oxidative phosphorylation makes them and uses the energy from those to convert the energy into ATP
|
|
|
describe the structure of mitochondria
|
have outer porous membrane, intermembrane space, inner membrane that is very permeable and a matrix inside there.
|
|
|
what reaction does pyruvate undergo to feed the citric acid cycle?
|
pyruvate + CoA --> coenzyme A, releases a CO2 and makes NADH
|
|
|
Where does Glycolysis occur?
|
cytoplasm of mitochondria
|
|
|
Where does the Citric Acid Cycle occur?
|
matrix of mitochondria
|
|
|
Where does Oxidative phosphorylation occur?
|
in the inner membrane of the mitochondria
|
|
|
what is the net result of "1 turn" of the citric acid cycle?
|
3 NADH, 1 FADH2, 1 GTP or ATP, 2 CO2 if counting those..
|
|
|
Describe the steps of the citric acid cycle.
|
Acetyl CoA and OAA condenses to make citrate. citrate isomerizes to iso- citrate. iso-citrate goes to alpha ketogluterate. if given the formula could determine it was an isomerization reaction, could you count carbons and see that it was an oxidative carboxylation where NADH got made. alpha ketogluterate goes to succinyl CoA goes to succinate. succinate goes to fumerate. fumerate to malate. malate back to oxyloacitate (OAA)
|
|
|
sequence of molecules for the TCA cycle (names only)
|
alpha ketogluterate goes to succinyl CoA. Succinyl CoA goes to succinate. succinate goes to fumerate. fumerate to malate. malate back to oxyloacitate (OAA)
|
|
|
describe oxidative phosphorylation. what does electron transport have to do with it?
|
electrons removed from NADH or FADH2. transferred through a series of electron carriers. while the electrons are carried or flow, protons are simultaneously pumped to the inter membrane space. so we essentially get an H+ ( proton) gradient. while the protons flow back through the ATP synthase we get ATP formed. e transport and proton pump go together.
|
|
|
what is the end receptor for the electrons?
|
oxygen. turns into water.
|
|
|
What electron carriers are in Complex I, III, IV, II? what are the mobile carriers between complex I-III and III-IV?
|
NADH-->complex I--> CoQ--> complex III--> cytocrome C-->complex IV--> oxygen//// FADH2-->complex 2-->CoQ-->complex III-->cytochrome C--> complex IV--> oxygen. the mobile carriers are NADH and CoQ or ubiquinome for I and II; cytochromes for III and IV.
|
|
|
how many sites are there for proton pumping?
|
3
|
|
|
In general, how does ATP synthase work? what are the F1 or F0 subunits? what does O, L, T mean?
|
have a proton gradient (protons in the inter membrane space), protons jump on little helical proteins in the F0 subunit and whirl around in this subunit being ejected into the matrix. while the F0 proteins are turning, this turns the beta subunits forcing them into these different conformations, (open- where product is released) (loose- where ATP is bound), (tight- where ATP is synthesized )
|
|
|
Approximately how many ATP are formed from 1 NADH?
|
2 1/2- 3, use 2 1/2
|
|
|
Approximately how many ATP are formed from one 1 FADH2?
|
1 1/2- 3, use 1 1/2
|
|
|
Typically at which position are phosphoryl groups attached?
|
5 prime phosphates or 3 prime phosphates.
|
|
|
one acetyl CoA --> ?ATP
|
3 NADH, 1 GTP, 1 FADH2 = 10 ATP
|
|
|
one glucose ---> ?ATP (glycolysis)
|
30 ATP; 2 NADH, 2 ATP...the 2 NADH, we lose energy.. so 2 FADH2= 3 ATP.. pyruvate --> NADH x 2 inside the matrix= 2 x 2.5 = 5 ATP.. 2 acetyl CoA.. 3 ATP x 5 ATP x 2 ATP = 30 ATP
|
|
|
What bases are purines?
|
adenine and guanine
|
|
|
what bases are pyrimidines?
|
thymine, uracil, cytosine
|
|
|
Which base is typically found only in RNA?
|
uracil
|
|
|
How is the base attached to the sugar?
|
beta n gylcosidic bond
|
|
|
Is a nucleotide a phosphomonoester or a diester?
|
phosphomonoester
|
|
|
what sugar is found in nucleotides that make up DNA?
|
2' deoxyribose
|
|
|
Describe the structure of B-DNA backbone?
|
sugar phosphate
|
|
|
where are the bases found in B-DNA?
|
bases are on the inside, perpendicular to the axis
|
|
|
are the strands parallel in B-DNA?
|
anti parallel
|
|
|
what are the 2 "grooves" called in B-DNA?
|
major and the minor
|
|
|
which bases pair in B-DNA? why?
|
C & G, A & T.. pair because of Hydrogen bonding
|
|
|
What is another stabilizing force in B-DNA?
|
hydrophobic, because of bases stacking on top of one another
|
|
|
is B-DNA a double helix?
|
yes
|
|
|
what happens if DNA denatures? can DNA renature?
|
two strands separate, typically when heated. yes they can re-nature if cooled gently
|
|
|
describe the structure of B-DNA.
|
double stranded, polymer of 2' deoxyribonucleotides, with 3' to 5' phosphodiester bonds, with antiparallel strands that wind around one another to make the double helix.
|
|
|
what is a nucleosome?
|
approximately 200 base pairs that are wrapped around histones. fairly basic unit of DNA stucture
|
|
|
describe differences between RNA and DNA.
|
RNA typically is not double stranded, not in a helix, more unstable, has ribose, and Uracil replaces Thymine.
|
|
|
what is the central dogma?
|
information travels from DNA --> RNA --> protein
|
|
|
what is replication?
|
duplicating DNA
|
|
|
what is transcription?
|
forming RNA from DNA templates
|
|
|
what is translation? reverse translation?
|
making protein from RNA template; vice versa
|
|
|
describe messenger RNA (mRNA)
|
has introns that are removed, is involved in protein synthesis, is a copy of part of the DNA
|
|
|
describe transfer RNA (tRNA)
|
contains an anti-codon, forms regions of double strands, is involved in protein synthesis, is a copy of part of the DNA, carries amino acids.
|
|
|
describe ribosomal RNA (rRNA)
|
found in the 60S particle, involved in protein synthesis, is a copy of part of the DNA
|
|
|
draw a phosphodiester, R is the ester group
|
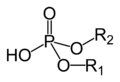
|
|
|
describe small nuclear RNA
|
processes heterogeneous nuclear RNA, is involved in protein synthesis, is a copy of part of the DNA
|
|
|
describe micro RNA
|
regulates some of transcription, involved in protein synthesis, is a copy of part of the DNA.
|
|
|
DNA polymerase III
|
bonds nucleotides
|
|
|
DNA polymerase I
|
removes primer
|
|
|
Primase
|
makes RNA primer
|
|
|
Ligase
|
connects Okazaki fragments
|
|
|
helicase, sometimes gyrase
|
help unwind helix
|
|
|
SSB- single stranded binding
|
protects single stranded DNA
|
|
|
draw 2' -deoxyribose
|
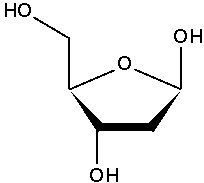
|

