![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
288 Cards in this Set
- Front
- Back
|
Define gene
|
Gross level - Inheritable units of information that specify phenotype (e.g. morphological characteristics)
Molecular level - a part of the DNA (polymer of nucleotides) of a chromosome that directs the synthesis of a specific polypeptide chain. |
|
|
Define Wild-Type
|
The normal, non-mutated version of a gene most common in nature.
|
|
|
Define chromosome
|
Thread-like, darkly staining bodies within the nucleus composed of DNA that carry the genetic information.
|
|
|
Define locus
|
The site of a gene on a chromosome
|
|
|
Define allele
|
Alternative form of a gene found at the same locus on homologous chromosomes (dominant/recessive)
|
|
|
Define homologous
|
Pair of chromosomes with the same gene sequence - same length, same centromere position etc.
|
|
|
Define genotype
|
The genetic constitution of an individual
|
|
|
Define phenotype
|
The set of observable characteristics of an individual resulting from the interaction of its genotype with the environment.
|
|
|
Define autosome
|
Any of the 22 non-sex chromosomes
|
|
|
Define homozygous
|
The presence of two identical alleles a a particular locus on homologous chromosomes
|
|
|
Define heterozygous
|
The state of having different alleles at a locus on homologous chromosomes
|
|
|
Define autosomal dominant
|
A gene on one of the non-sex chromosomes that manifests in the heterozygous state
|
|
|
What is a codon?
|
A three base pair sequence that codes for an amino acid
|
|
|
How many chromosomes are present in a human somatic cell? How are these paired?
|
46
23 pairs - 22 homologous chromosome pairs and one sex chromosome pair |
|
|
How many chromosomes are present in a human gamete?
|
23
|
|
|
Define karyotype
|
The number, size and shape of the chromosomes of an individual.
Also used to describe the photomicrograph of an individual's chromosomes arranged in a standard manner. |
|
|
Define a point mutation
|
A single base-pair change
|
|
|
What are the four types of point mutation?
|
Silent
Nonsense Missense Single nucleotide insertions or deletions (types of frameshift mutation) |
|
|
What is a transition substitution?
|
A substitution involving replacement by the same type of nucleotide - a pyrimidine for a pyrimidine (e.g. C for T) or a purine for a purine (e.g. A for G)
|
|
|
What is a transversion substitution?
|
Substitution of a purine for a pyrimidine or a pyrimidine for a purine.
|
|
|
Results of a silent mutation
|
The genetic code is degenerate (or redundant) - multiple codons code for the same amino acid (64 codons (4 bases, codons are triplets, therefore 4-cubed) for 20 amino acids).
In the case of silent mutations, the change of base does not change the amino acid sequence --> no effect on final protein (synonymous) |
|
|
Results of nonsense mutations
What dictates their severity? |
Change in base pair results in the affected triplet coding for a stop codon -prematurely signals to RNA polymerase to detach from the coding strand, thereby terminating transcription from the point of mutation onwards --> produces a truncated protein.
Depends on their location along the coding strand – the earlier the stop codon is inserted, the more affected the final protein product will be. |
|
|
Results of missense mutations
What dictates their severity? |
Change in the amino acid coded for by the triplet
Depends on the amino acid changed, its role in the protein's function and what it is changed to - can be conservative and non-conservative. |
|
|
Conservative substitution
|
Type of missense mutation where the single base-pair substitution results in the incorporation of a different amino acid however since the new amino acid is chemically similar to the original, the change has no functional effect.
|
|
|
Non-conservative mutation
|
Missense mutation incorporating an amino acid that differ greatly from the original, resulting in grossly altered and even loss of the final protein’s function.
|
|
|
Synonymous mutation
|
One which does not change the structure of the final protein product (silent mutation)
|
|
|
Non-synonymous mutation
|
Mutation that leads to an alteration in the encoded polypeptide
|
|
|
Results of single nucleotide insertions and deletions
|
Frame shift mutations - change the reading frame of the codon triplets.
It is very unusual for the resulting missense protein to be able to perform the functions of the wild-type one. |
|
|
Define reading frame
|
The order of the triplets of nucleotides in the codons of a gene that are translated into the amino acids of the primary sequence of a protein
|
|
|
What causes frame-shift mutations?
|

Insertions or deletions of anything other than multiples of three bases
|
|
|
What are the consequences of insertion/deletion of multiples of 3?
What determines severity? |
Same as mis/nonsense point mutations - either an amino acid is added/deleted or a stop codon is added.
Severity depends on the importance of the protein involved and the amino acids affected. |
|
|
Example of disease caused by the loss of a single amino acid
|
Cystic fibrosis - lose the Phe residue at position 508 of 1480 resulting in the trafficking failure of the CFTR protein in the epithelial membrane.
|
|
|
What are dynamic mutations? Example
|
Nucleotide repeats which expand generation by generation.
When a certain threshold of repeats is reached, the disease phenotype will become apparent e.g. Huntingtons |
|
|
Define anticipation
|
Disease phenotype becomes progressively more severe and/or manifest at an earlier age from one generation to the next
|
|
|
What are exons?
|
The coding regions of genes - code for the protein the gene encodes
Exon sequence is highly conserved between individuals |
|
|
What are introns?
What is their fate during processing? |
The non-coding regions of DNA
Spliced out during processing into mRNA Intron sequence not well conserved between individuals |
|
|
Define splicing
|
The removal of introns and joining of exons in RNA during transcription - introns are spliced out and exons spliced together
|
|
|
Where could mutations in an intron cause problems? (2)
|
Regulatory regions - repressor or promotor
Splice sites |
|
|
What can be the consequences of mutations in the regulatory elements of introns?
|
Inappropriate up- or down-regulation of gene expression
|
|
|
What can be the consequences of mutations in splice sites?
|
Interfere with correct splicing of exons together during processing of RNA leading to mistaken inclusion of introns or exclusion of exons.
Very likely this will also cause frame-shift mutations/premature stop codons |
|
|
EXPERIMENTAL
What bacterium was used to identify DNA as the hereditary material? |
Pneumococcus - two types of streptococcus pneumoniae used
|
|
|
EXPERIMENTAL
Characteristics of wild-type (smooth) streptococcus pneumoniae Why is it called smooth? |
Smooth after its appearance under a microscope
Is virulent and kills mice |
|
|
EXPERIMENTAL
Characteristics of avirulent (rough) streptococcus pneumoniae Why? |
Mutant - harmless as it lacks the appropriate surface polysaccharide needed to infect cells
|
|
|
EXPERIMENTAL
What is 'smooth avirulent' streptococcus pneumoniae? |
The wild-type rendered avirulent by heat treatment and so harmless, however retains smooth appearance.
|
|
|
EXPERIMENTAL
What was the first key experiment to determine DNA as the hereditary material? When? Who? |
Mouse injected with smooth + rough bacteria --> dead mouse
Bacteria isolated from dead mouse = solely smooth (virulent) Conclusion - something in smooth bacteria enabled mutant rough bacteria to produce the missing polysaccharide and thus become virulent. Griffith, 1928 |
|
|
EXPERIMENTAL
Details of experiment carried out to investigate what the 'something' in smooth bacteria that enabled the rough bacteria to become virulent was |
DNA extracted from wild-type (smooth) bacteria
Inject mouse with both virulent + avirulent --> dead mouse However Avirulent + virulent that has been treated with DNAase prior to injection --> live mouse BUT Avirulent + virulent that has been treated with protease trypsin --> dead mouse Avery, 1944 |
|
|
EXPERIMENTAL
Details of experiment to support findings of Griffith (1928) and Avery (1944) |
Use phage (infect bacteria) LOOK UP
Hershey-Chase, 1952 |
|
|
What are DNA and RNA?
|
Nucleic acids
|
|
|
What are nucleic acids made up of?
|
Long polymer of nucleotides
|
|
|
What are nucleotides made up of?
|
Nitrogenous base
A sugar (deoxyribose in DNA, ribose in RNA) A phosphate group |
|
|
What are the two types of nitrogenous base?
|
Purine
Pyrimidine |
|
|
Which are the two purine bases?
|
Adenine and Guanine
|
|
|
Which are the two pyrimidine bases?
|
Cytosine and Thymine (DNA)/ Uracil (RNA)
|
|
|
What direction is DNA always read?
|
5'-3' --> polarity
|
|
|
What do 5' and 3' refer to?
|
The free carbon atom from the sugar which is free at the end of the chain in question
|
|
|
Structure of the 5' end
|
Phosphate on the C5 of the sugar (deoxyribose)
|
|
|
Structure of the 3' end
|
Ends with a hydoxyl group on the C3 of the sugar
|
|
|
What is the 'anti-parallel nature of the DNA double helix'?
|
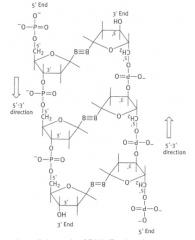
DNA strands associate into pairs and run in opposite directions (one, 5'-3' and the other 3'-5')
|
|
|
What are the base pairing rules?
|
C- G with 3 H-bonds
A-T with 2 H-bonds |
|
|
EXPERIMENTAL
How was the double helix nature of DNA first observed? |
Elucidated from X-ray diffraction images of crystallised DNA
|
|
|
How does DNA replication occur?
|
By a semi-conservative process - each daughter molecule has one DNA strand from the parent and one newly synthesised strand.
|
|
|
EXPERIMENTAL
How was semi-conservative DNA replication first show? Who? When? |
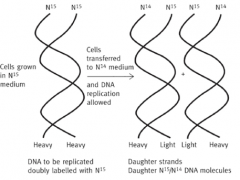
E.Coli were grown for several generations in media containing N15 - this was incorporated into their DNA
Cells suddenly switched to N14-containing media and allowed to divide. First-generation cells had DNA which was 50% N15, 50% N14 Meselson & Stahl, 1958 |
|
|
Name the enzyme that opens the DNA helix during replication
|
DNA helicase
|
|
|
Name the enzyme that prevents unwound single DNA strands from getting tangled. What kind of enzyme is this?
|
DNA gyrase
A topoisomerase |
|
|
What are topoisomerases? How do they work?
|
Enzymes which regulate the under and over-winding of DNA
e.g. During DNA replication, the DNA helix becomes overwound just before the replication fork - without topoisomerases, this overwinding would eventually halt replication. They bind to single or double stranded DNA and cut the phosphate backbone, allowing the DNA to be untangled and the backbone is then resealed. |
|
|
What stabilises the single DNA strands?
|
Single-stranded DNA binding proteins
|
|
|
What is the point where the DNA strand separate called?
|
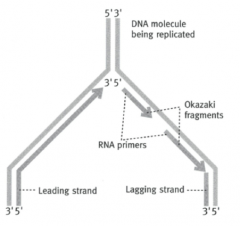
Replication fork
|
|
|
Name the enzyme the copies DNA
|
DNA polymerase (cannot initiate a new strand)
|
|
|
Which direction is DNA copied in?
|
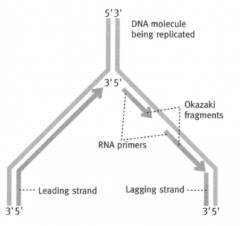
5' --> 3' direction
|
|
|
What is the 'leading strand'?
|
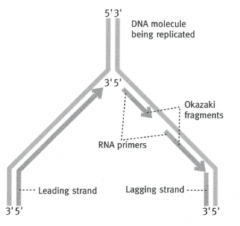
5' --> 3' strand - can be copied directly by DNA polymerase
|
|
|
What is the lagging strand?
|
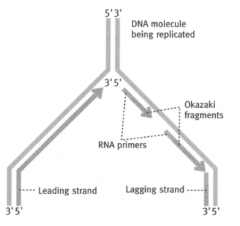
The 3' --> 5' strand
|
|
|
Name the enzyme which initiates DNA replication. What kind of enzyme is this? How does it work?
|
RNA primase - an RNA polymerase
Makes a short (10-20 bases) RNA primer - For the leading strand, DNA polymerase can extend this directly DNA polymerase must copy the lagging strand in small pieces - each piece is primed with RNA |
|
|
What are the pieces the lagging strand is copied in called?
|
Okazaki fragments
|
|
|
What happens when DNA polymerase encounters the RNA primer of a previously made fragment?
|
A 5' -->3' RNaseH removes the RNA and DNA polymerase replaces it with DNA
|
|
|
What enzyme joins adjacent Okazaki fragments together?
|
DNA ligase
|
|
|
How do DNA polymerases create the new DNA strand?
Why must the new strand be very accurate? How is this accuracy achieved? |
DNA polymerases use one strand of the original DNA as a template, sequentially adding nucleotides using the base pair rules (A&T and C&G).
These pairings must be very accurate for faithful DNA replication - to ensure maintained integrity of the genetic code. DNA polymerases have a proof-reading facility allowing them to correct mistakes and so enhance accuracy. |
|
|
What is the start codon from almost all proteins?
|
ATG
|
|
|
Name 3 stop codons
|
TAG, TGA and TAA
|
|
|
Name for genes expressed ubiquitously in cells at constant levels
|
Housekeeping genes
|
|
|
2 ways to control gene expression
Is gene expression reversible or permanent |
Internal - genetic signals
External factors - hormones Can be both |
|
|
What are transcription factors?
Do they have a long or short term effect? |
The protein products of genes that control the expression of other genes
(also known as gene regulatory factors) - bind to specific DNA sequences. Tend to have a long term effects. Include permanent changes that occur in cell differentiation. |
|
|
CHECK
Structure of gene regulatory proteins |
Have a DNA binding motif and a transcription activation motif
|
|
|
Example of a transcription protein - what is it's structure?
|
Helix-turn-helix - consists of two alpha-helices connected by a short sequence of amino acids to form the turn.
|
|
|
Explain how helix-turn-helix proteins act 'in a trans manner'
|
They affect genes far away in the genome by recruiting other regulatory proteins expressed in the cell - these recruited complexes bind to the gene and are thought to act by increasing the accessibility of the initiation site to RNA polymerase and/or exposing other regulatory sites.
|
|
|
Overall, how do enhancers act?
|
By perturbing the chromatin structure of the gene that they regulate.
|
|
|
Best example of an external factor affecting gene expression
Time frame of effect? |
Steroid hormones
Can be transient or longer term but are generally reversed when the hormone signal is removed |
|
|
Are steroids lipid soluble?
|
Yes
|
|
|
How do steroids affect gene expression?
|
Diffuse from the extracellular fluid into cells where they bind to receptor proteins - form hormone-receptor complexes.
Hormone-receptor complexes diffuse into nucleus where they bind to receptor elements in the DNA (upstream from transcription start) and either up or down-regulate gene expression. |
|
|
Aside from hormone-receptor complexes, how else can hormones regulate gene expression?
|
Can act via 2nd messenger systems to affect gene expression e.g. cAMP-sensitive response elements
|
|
|
What are transcription factors that increase the transcription of genes called?
|
Enhancers
|
|
|
What are transcription factors that inhibit the transcription of genes called?
|
Repressors/silencers
|
|
|
Examples of 3 promoter regions which enhancers bind to
How many base pairs are these elements from the transcription start site? Why are they said to be 'cis-acting'? |
The GC box
The CAAT box The TATA box Within 100bp Because they act on the gene immediately following |
|
|
How do transcription factors binding to the GC and CAAT boxes enhance transcription?
|
Increase the basal activity of the TATA box
|
|
|
How can mRNA affect gene expression?
|
mRNA processing, translation and mRNA breakdown - changes in breakdown rates relative to formation
|
|
|
EXPERIMENTAL
Studies of what have given the most information about gene expression? |
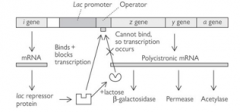
The lac operon in bacteria
|
|
|
What is an operon?
|
A group of genes with related functions which are under the control of a single promotor. Found only in prokaryotes
|
|
|
Define polycistronic
|
Describes mRNA that codes for multiple different polypeptides
|
|
|
Define monocistronic
|
Describes mRNA that codes for just one protein
|
|
|
Is the mRNA produced from an operon polycistronic or monocistronic?
|
Polycistronic
|
|
|
What does expression of the lac operon allow bacteria to do?
|
Use lactose as an energy source
|
|
|
What happens to expression of the operon in the absence of lactose?
|
The lac repressor binds to the operator and blocks expression
|
|
|
What happens to expression of the operon in the presence of lactose?
|
The metabolite of lactose, allolactose (the inducer molecule), binds to the repressor protein blocking expression and allosterically changes its shape.
The repressor protein no longer binds to the operator - RNA polymerase can transcribe the genes involved in bacterial lactose metabolism. |
|
|
How was regulation of gene expression of the lac operon shown? (2)
|
Experimentally - using the artificial substrate IPTG
The lac repressor protein has been crystallographically studies and its structre supports the proposed mechanism of action. |
|
|
In addition to mRNA, what RNAs need to be transcribed in order for a eukaryotic cell to make a new protein? What isoform of RNA polymerase transcribes each?
What are the roles of these additional RNAs? |
Ribosomal RNA (rRNA) - transcribed by RNA polymerase I
Transfer RNA (tRNA) - RNA polymerase III Small nuclear RNA (snRNA) - RNA polymerase II They are not translated into proteins, but are catalytically active in the RNA form. |
|
|
What is the first stage of gene expression? How many steps to this are there and what are they called?
|
Transcription of the gene of interest
Takes place in three steps: Initiation, elongation and termination |
|
|
What is the first step of initiation?
|
The binding of the TATA box-binding protein (TBP) and the TBP-associated factors (TAFs, collectively known as TFIID) to the TATA box
|
|
|
What is the result of the TBP and TFIID binding to the TATA box?
|
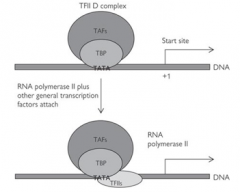
Causes the sequential recruitment of other transcription factors for RNA polymerase II (TFIIs) - TFIIA, B and F.
Followed by recruitment of RNA polymerase itself and finally TFIIE and H. This forms the initiation complex |
|
|
What occurs after assembly of the initiation complex?
|
The complex unwinds a short stretch of the DNA double helix, revealing the single-stranded DNA that it will transcribe and interacts with transcription factors that regulate the rate of transcription.
|
|
|
What is the rate of transcription initiation by the basal transcription apparatus?
|
Depends on the presence of enhancing transcription factors - in their absence, it is relatively slow
|
|
|
What is the first step of elongation?
|
RNA polymerase II selects the correct ribonuclotide triphosphate and catalyses the formation of the phosphodiester bond
|
|
|
Does the RNA molecule require a primer?
|
No
|
|
|
What direction is snRNA synthesised in?
|
5' --> 3'
|
|
|
What are the base pairing rules for RNA synthesis?
|
Normal nucleotide base pairing rules apply, except uracil (U) is paired with adenine (A) when it occurs in the template strand (the DNA strand RNA polymerase II is transcribing)
|
|
|
In what direction does the enzyme (RNA polymerase) move?
|
Unidirectionally away from the promotor region, along the DNA template.
|
|
|
What does 'the polymerase is completely processive' mean?
|
A single enzyme transcribes a complete RNA strand
|
|
|
How are transcription errors accounted for in RNA?
|
RNA polymerases do not have any proof-reading mechanism - it is less important to have such high fidelity as in DNA replication as RNA is not inherited.
Usually, many copies of RNA are made when a gene is expressed and so the mistakes are unlikely to affect overall protein synthesis. |
|
|
What indicates when RNA transcription should stop? Why are these important?
|
Termination signals
Because RNA polymerases are processive |
|
|
What is the simplest stop signal?
|
A palindromic GC region immediately followed by a T-rich region (in the RNA sequence)
|
|
|
What structure does the palindromic GC-rich region form and why?
|
Forms a hairpin structure due to base pairing
|
|
|
What does the T-rich region following the palindromic GC region cause?
|
An oligo(U) sequence immediately after the hairpin
|
|
|
Collective name for the hair pin structure and following oligo(U) sequence that form the most common stop signal?
|
The hairpin-oligo(U) complex
|
|
|
How does the hairpin-oligo(U) complex terminate transcription?
|
It is thought to destabilise the relatively weak association between the newly synthesised RNA and the DNA template strand leading to their dissociation - results in the dissociation of the RNA polymerase II complex.
|
|
|
What do the DNA strands do once transcription is complete?
|
Reform into a double helix
|
|
|
Other than coded stop signals, what else can assist in terminating transcription? Example?
|
Proteins - e.g. the rho protein, discovered in prokaryotes.
|
|
|
How does the rho protein terminate transcription?
|
Binds to the newly made RNA at C-rich, G-sparse regions and scans along the RNA towards the RNA polymerase -an ATP-dependent process.
When it reaches the RNA polymerase, the protein breaks the DNA/RNA association - halts transcription. |
|
|
Does the transcription termination signal (either protein or coded) lie in the newly formed RNA or in the DNA template?
|
The RNA
|
|
|
What is RNA as it is transcribed by DNA polymerase II called?
|
pre-mRNA
|
|
|
What is the first step in the processing of pre-mRNA?
|
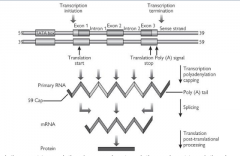
The pre-mRNA is capped:
G residue is added to the 5' end by a (very unusual) 5'-5' triphosphate bond. This (the terminal G) is then methylated on the N7 and possibly also the adjacent riboses. |
|
|
Why is the pre-mRNA capped?
|
The role of the cap is to protect the 5' terminal of the mRNA against degradation and also to improve ribosomal recognition for translation.
|
|
|
Second step of pre-mRNA processing?
|
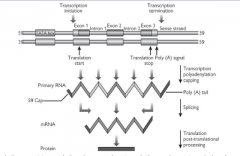
Polyadenylation - a poly(A) tail is added to the RNA
|
|
|
What is the polyadenylation consensus sequence? What does it signal? What enzyme is involved?
|
AAUAAA - signals that the pre-mRNA should be cleaved around 20 bases downstream by an endonuclease
|
|
|
What enzyme adds the poly(A) tail?
What does the tail comprise? |
Poly(A) polymerase.
Can consist of hundreds of residues |
|
|
Why is the poly(A) tail added?
|
Like the cap - thought to improve mRNA stability and enhance translation
|
|
|
Third step of pre-mRNA processing?
|
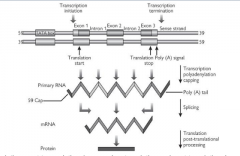
Code for introns spliced out - pre-mRNA contains sequences for introns and exons while mature mRNA contains only code for exons
|
|
|
What removes the introns? How does it know where to act?
|
Spliceosomes - splice at specific recognition sites at the ends of introns
|
|
|
What are spliceosomes?
|
Consist of a number of snRNA molecules, associated proteins called splicing factors and the pre-mRNA being processed
|
|
|
What are the snRNA and protein complexes known as alone (without pre-mRNA)?
|
Small nuclear ribonucleoproteins (snRNPs - 'snurps')
|
|
|
Can mRNA be edited?
|
Yes, post translational processing - its base sequence can be changed
|
|
|
CLINICAL
What is apolipoprotein B? Name its two forms and state where they are formed. |
A lipid and cholesterol transport protein
Apo B-100 - formed in liver Apo B-48 - formed in small intestine |
|
|
What is apo B-48 in relation to apo B-100?
How is it formed? (List ways one might expect that do not in fact occur) |
Apo B-48 is a truncated form of apo B-100
Tissue-specific mRNA editing inserts a stop codon into the apo B-100 transcript. NOT protein cleavage of the larger form, nor by alternative splicing |
|
|
Where in the cell does translation take place? How does mature RNA arrive there?
|
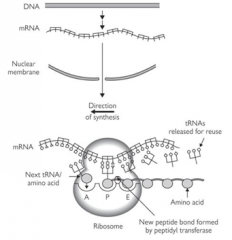
In the cytoplasm - mature RNA leaves the nucleus through pores in the nuclear membrane
|
|
|
Which organelle carries out translation?
Where are these situated in the cell? |
Ribosomes
On the cytoplasmic face of the rough endoplasmic reticulum (rER) and free in the cytoplasm |
|
|
What kind of proteins are synthesised on the rER?
|
Proteins that will be secreted, targeted to organelles or inserted into the plasma membrane
|
|
|
What kind of proteins are synthesised on free cytoplasmic ribosomes?
|
Those that will remain in the cytoplasm
|
|
|
How are ribosomes structured? (prokaryotic and eukaryotic - overall)
Hint - subunits |
Eukaryotic - 60S (large) subunit and 40S (small) subunit
Give a total 4200 KDa 80S complex Prokaryotic - 50S (large) subunit and 30S (small) subunit Total 70S |
|
|
Structures of eukaryotic ribosomes' subunits?
|
The subunits are ribonucleoproteins - composed of a large number of proteins and one or more RNA molecules.
60S - 5S (120 nucleotides), 28S (4700 nucleotides), a 5.8S (160 nucleotides) RNA subunits and 46 proteins 40S - 18S RNA subunit (1900 nucleotides) and 33 proteins. |
|
|
Structures of prokaryotic ribosomes' subunits?
|
The subunits are ribonucleoproteins - composed of a large number of proteins and one or more RNA molecules.
50S - 5S (120 nucleotides) and 23S (2900 nucleotides) RNA subunits and 31 proteins. 30S - 16S RNA subunit (1540 nucleotides) bound to 21 proteins. |
|
|
What units are ribosome subunits measured in?
|
The Svedberg unit (S) - a measure of the rate of sedimentation in centrifugation rather than size (hence why the fragment 'sizes' do not add up
|
|
|
What are the key reactive sites in ribosomes made up of?
|
Almost entirely RNA
|
|
|
Does protein synthesis begin at the amino or carboxyl terminus?
|
Protein synthesis begins at the N-terminus (amino) and runs through to the C-terminus (carboxyl)
|
|
|
In what direction is mRNA translated?
|
5' --> 3' direction
|
|
|
What is always the first amino acid in eukaryotic proteins?
|
Methionine (AUG - start codon)
|
|
|
How is the codon for methionine (the start codon) recognised by the ribosome?
|
The 40S subunit attached to the cap at the 5' end of the mRNA and scans along the strand until it finds AUG
|
|
|
Which AUG in the mRNA is usually the start codon and why?
|
The most 5' AUG as eukaryotic mRNA is monocystronic
|
|
|
What is bound to the 40S subunit?
|
initiation factor proteins, GTP and the Met-tRNAi (the specific tRNA for starting a new polypeptide chain.
|
|
|
What happens when the 40S substitute finds the initiation codon?
|
The initiation factor proteins (initiation factors) dissociate and the 60S subunit is recruited
|
|
|
How many tRNA binding sites does the 80S ribosome complex have? What are they called?
|
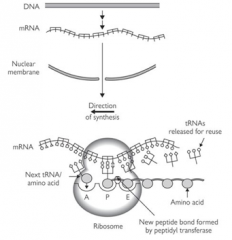
3 - P, A and E (exit) sites
|
|
|
To which site does Met-tRNAi bind?
|
P
|
|
|
Where does the next tRNA (complimentary to the next codon of mRNA) bind?
|
The A site
|
|
|
What kind of bond forms between the amino acids of these two tRNAs?
Enzyme that catalyses this reaction? |
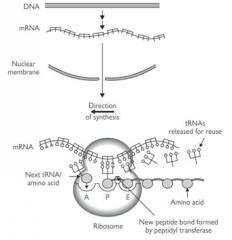
Peptide bond
Peptidyl tranferase |
|
|
What happens after the peptide bond is formed?
|
Ribosome translocates along the RNA towards the 3' end by one codon - tRNAs formerly in P and A sites, now in E and P.
Newly liberated A site can accept next tRNA. |
|
|
What drives the process of ribosome translocation?
|
GTP and elongation factors (enzymes)
|
|
|
How is the process terminated?
|
When the stop codon is reached in the mRNA template (end of the strand) - no tRNAs complimentary to stop codons, instead these are recognised by the release factor (eRF1) which causes dissociation of the ribosome from the completed polypeptide chain.
|
|
|
CLINICAL
How have the differences between prokaryotic and eukaryotic ribosomes and translation processes been therapeutically exploited? |
Antibiotics which inhibit protein synthesis only of the invading prokaryotes e.g. tetracycline (amino glycosides) and streptomycin
|
|
|
What happens to proteins synthesised on rER?
|
Either inserted into the rER membrane or enter the rER lumen.
For most secreted proteins, there is a signal sequence that allows the polypeptide to enter the lumen as it is being synthesised which is then cleaved during subsequent protein processing. |
|
|
How do signal sequences aid membrane proteins?
|
Type I membrane proteins - there is thought to be some sort of signal sequence that allows the first transmembrane region to enter the membrane
OR Type II membrane proteins - signal sequence that allows the N terminal to cross the membrane |
|
|
How many base pairs does the human genome contain? (Approx.)
|
3.2 x 10^9
|
|
|
When was the first complete sequence of the human genome released?
|
2001
|
|
|
How many genes are there thought to be?
|
30,000 genes (or open reading frames (ORFs) for proteins)
|
|
|
Exons are what percentage of total DNA?
|
1.5%
|
|
|
What is the rest of DNA? (Not exons)
|
Regulatory elements and introns
|
|
|
What is the average length of an exon?
|
145 base pairs
|
|
|
How many exons, on average, are there per gene?
|
Around 8.8
|
|
|
What is single-copy DNA and how much of the genome does it represent?
|
Mainly unique intron and regulatory sequence with a small fraction coding for proteins. Makes up almost half of the human genome.
|
|
|
What is moderately repeated DNA and how much of the genome does it represent?
|
DNA that is repeated from a few up to a thousand times, sequences varying in length from 100 to 1000s of base pairs. About 15% of the human genome.
|
|
|
What is highly repeated DNA?
|
Short (usually <20 nucleotides) sequences repeated millions of times
Also called SNPs - single nucleotide repeat sequences |
|
|
What is inverted repeat DNA?
|
Structural motif of DNA with an average length of 200 base pairs (can be up to 1000) - in humans they form hair pin structures.
Can be repeated over 1000 times per cell. |
|
|
Examples of how the size of the genome does not necessarily correlate to the number of ORFs (2)
|
Arabidopsis (Wall cress) has a genome of similar size to Drosophila (Fruit fly) but almost double the genes
The human genome is almost 10^3-fold larger than E.Coli but has less than 10-fold the number of genes |
|
|
Why is the human genome so large?
|
Large amount of introns
|
|
|
What is the function of introns?
|
No exactly known - often regarded as 'junk' DNA
However, it has been observed they must have a function as there is a large metabolic penalty to maintain them. Much of the non-coding sequence is made up of SNPs (53% of total genome) |
|
|
Why do prokaryotes have smaller genomes than eukaryotes?
|
Do not have introns to the same extent
|
|
|
What is DNA cloning?
|
The amplification of a particular DNA fragment
|
|
|
Why is DNA cloning useful?
|
Has allowed individual gene products to be studied in detail at both a structural and functional level
|
|
|
What method is used if the sequence is known?
|
The fragment is amplified using gene-specific primers by the polymerase chain reaction (PCR)
|
|
|
To what are primers made complimentary to for the PCR?
|
Made to flank the region of the gene which is to be amplified (they are approximately 25 bases long)
|
|
|
PCR Step 1 - what temperature are the template and primers heated to? Why?
|
95°C - to denature them
|
|
|
PCR Step 2 - what is the temperature changed to for this step? Why?
|
Around 60°C - temperature is reduced to one that allows the primers (which are in vast excess) to bind to their complimentary regions on the template. 60°C is the temperature the primers are designed to do this.
|
|
|
PCR Step 3: Temperature? Why?
|
72°C - allows DNA polymerase to rapidly extend the primer (rate about 1000bp/min)
|
|
|
How many times are these steps repeated? (PCR)
|
Around 30 times
|
|
|
How many copies of the sequence does the PCR produce?
|
Over 1 x 10^9
|
|
|
How is the PCR product visualised?
|
By electrophoresis on an agarose gel
|
|
|
What is agarose?
|
A carbohydrates isolated from seaweed
|
|
|
How are the DNA fragments separated by size?
|
DNA migrates through the agarose gel when an electrical current is applied - fragments move towards the anode because DNA is negatively charged (due to the phosphates). The smaller the fragment, the faster (and so further) it migrates.
|
|
|
How is the separated DNA visualised?
|
Staining with ethidium bromide - fluoresces under UV light
|
|
|
What two developments made PCR a much more feasible and routine technique?
Hint: DNA polymerase, cyclers |
Isolation and purification of thermostable DNA polymerase from bacteria that have evolved to live in hot springs - this polymerase is able to withstand repeated exposure to 95°C and is most active at 72°C
Manufacture of thermal cyclers - programmable heating blocks which can rapidly change temperature. |
|
|
What method of DNA cloning is used if part of the required sequence is known?
|
A labelled DNA probe is attached to this fragment and can be used to identify the gene
|
|
|
Define transfection
|
Deliberately introducing nucleic acids into cells
|
|
|
How is the fragment identified if the function of the gene is known?
|
By testing cells transfected with the gene for that particular response
|
|
|
How is a cDNA library created and what is it used for?
|
Isolate all mRNA from a particular cell that is known to express the protein of interest.
mRNA converted into cDNA using enzyme reverse transcriptase (RT) cDNA then clone into a vector using transcription enzymes and amplified in E. coli The clones can be screened to identify which contains the gene of interest To represent all the mRNA present in the cell, the cDNA library must have many 1000s of clones. |
|
|
What is the primer for the RT reaction?
|
Often an 15-18mer of poly(T) that hybridises to the poly(A) tail of the mRNA
|
|
|
From what are restriction enzymes isolated?
How do they act? |
Bacteria
Cut DNA at specific sequence of 4 or more bases |
|
|
What are the two types of 'ends' that REs can produce?
|
Sticky ends - either 3' or 5' overhangs which can be joined by DNA ligase to a compatible DNA fragment that has a complimentary overhang
Blunt ends - no overhang and can be ligated to any other blunt end |
|
|
What results from the joining of two unrelated DNA fragments?
|
A recombination DNA molecule
|
|
|
What are vectors (or replicons)?
|
Carrier DNA that can independently replicate in a host organism to produce multiple copies of itself
|
|
|
Most commonly used vectors?
Where do they replicate? |
Plasmids
E. coli |
|
|
What features does a plasmid need to have?
|
An origin of replication, gene coding for antibiotic resistance for selection of transformed cells, multiple cloning site (MCS - a number of unique RE sites close together that allow the insertion of the gene of interest), promoters flanking the MCS to allow the cloned gene to be expressed
|
|
|
Types of vector other than plasmids
|
Bacteriophages, cosmids, bacterial artificial chromosomes (BACs) and yeast artificial chromosomes (YACs)
|
|
|
Why are BACs, YACs and cosmids useful?
|
Have the advantage of being able to take larger inserts than plasmids and bacteriophages
YACs have been useful in mapping eukaryotic genes which can be up to several million bps in length |
|
|
Why is PCR often used in tandem with creating cDNA?
What is this technique called? |
To look for gene expression in a particular tissue
RT-PCR |
|
|
CLINICAL
How can RT-PCR be used? |
In combination with sequencing, as a diagnostic tool for diseases involving known point mutations
|
|
|
Techniques that allow recognition of either nucleic acid or protein from samples (3)
|
Northern, southern and western blotting
|
|
|
Details of northern blotting
|
Uses labelled antisense DNA probe to look for copies of mRNA which have been separated by size on a gel and blotted onto a filter.
The probe will hybridize to its complimentary message - allows determination of the size and quantity of the mRNA transcript in the sample |
|
|
Southern blotting
Useful in? |
Similar to northern except that the template is restriction enzyme-treated DNA rather than mRNA
Genotyping e.g. gene knock-out animals |
|
|
Western blotting
|
Protein samples are run on a denaturing polyacrylamide gel and then transferred onto a filter.
The filter is then probed using an antibody to the protein in question. The antibody biding can be quantified using an enzyme-conjugated 2° antibody (e.g. anti IgG-horseradish peroxidase) that can be assayed colourimetrically |
|
|
What are the two main techniques of DNA sequencing?
When were they developed? |
The Maxam-Gilbert method - chemical cleavage
The Sanger method - interrupted enzymatic cleavage Late 1970s |
|
|
First step of the Maxam-Gilbert method
|
Double-stranded DNA to be sequenced is cut into fragments and the end of one is labelled by attaching a radioactive phosphorus (32P) group (32P-dATP) using part of (the Klenow fragment) E. coli DNA polymerase I
|
|
|
M-G Step 2
|
Labelled DNA is chemically treated in 4 separate reactions - each induces it to break (chemical cleavage) into fragments specifically at positions occupied by one of the four possible bases. This results in a set of radioactive fragments extending from the 32P label to each of the positions occupied by each base in the fragment
|
|
|
M-G Step 3
|
Fragments then undergo electrophoresis on a polyacrylamide gel and so become separated by size. The fragment's sequence can then be read from an autoradiograph - photo on X-ray film.
|
|
|
What is the overall principle of the Sanger method?
|
Based on the random interruption of the synthesis of a DNA strand in a reaction analogous to the PCR
|
|
|
How many reactions are in the Sanger method?
|
4, each with dNTPs and a single ddNTP species
|
|
|
What is a ddNTP?
|
A dideoxynucleoside triphosphate - has an H on C3 of the sugar group rather than the OH in a dNTP
|
|
|
How are ddNTPs incorporated into DNA?
Why can they not be extended? |
By a polymerase
Due to the lack of a 3'OH for the next nucleotide |
|
|
What is significant about the concentrations of d and ddNTPs in the Sanger method reactions?
|
Concentrations are such that dNTP is incorporated randomly at every position
|
|
|
How and why is ddNTP labelled?
|
Used to use 32P-ddNTPs but now more common to use ddNTPs labelled with fluorescent molecules (different colour for each base)
Labelled so that the differently sized fragments can be visualised |
|
|
How are the fragments separated?
How is the sequence visualised? |
Electrophoresis on polyacrylamide gel
For 32P-ddNTPs, an autoradiograph is taken of the gel For fluorescent, the gel can be read with a laser for the fluorescent ddNTP method. (coloured wave graph) |
|
|
CLINICAL
Why is direct sequencing of PCR produces from patient tissue useful? |
Allows relatively easy diagnosis of genetic diseases e.g. haemophilia and CF - one or more small regions of DNA can be amplified by PCR and then sequenced directly to determine if a mutation in the region(s) of the gene amplified.
|
|
|
Define a 'genetic disease'
|
The disease is always manifested in 'standard' environmental conditions if the genetic abnormality is present - some have complete expression no matter what the environment e.g. Down's syndrome whilst others are only revealed in environments which are now considered to be normal but were not in the recent evolutionary past e.g. certain types of hyperlipidaemia which manifest as ischemic heart disease in populations with high-calorie, high-fat diets.
|
|
|
What is a single gene abnormality?
|
A mutation in a gene which codes for a structural protein which leads to either no production of the protein or a protein with no function
|
|
|
Clinical example of a single gene abnormality
|
Duchenne muscular dystrophy - a deletion of the dystrophin gene leads to absence of this protein which is essential for the maintenance of the myocyte membrane during muscle contraction.
|
|
|
How are single gene abnormalities transmitted? (3)
|
Modes of inheritance:
Autosomal dominant Autosomal recessive X-linked |
|
|
Result of autosomal dominant inheritance of the gene anomaly
|
The abnormal gene has an effect i.e. it is penetrant, even when a normal gene is present on the other allele of the homologous chromosome
|
|
|
Define penetrance
|
The regularity with which an allele is expressed in a person who carries it.
|
|
|
Result of autosomal recessive inheritance of the gene anomaly
|
Abnormal gene only has an effect if the allele on the homologous chromosome has the same abnormality or has been deleted/silenced by hypermethylation
|
|
|
Result of X-linked inheritance of the gene anomaly
|
There is incomplete homology between X and Y chromosomes with X having a substantial amount of additional genomic material - if an abnormal gene occurs in this region, then there is no corresponding normal allele on the Y chromosome to counteract it, even if the abnormal gene would otherwise have a recessive pattern of inheritance. Males tend therefore to be more affected by X-linked diseases than females
|
|
|
Mechanisms of single gene abnormalities
|
Deletion or insertion of bases within a coding region of a gene.
Point mutation (non or missense) of a base within a coding region of a gene. Fusion of two genes Deletion, insertion or point mutation in a non-coding region of a gene |
|
|
Results of deletion and insertion of bases within a coding region
CLINICAL examples |
Leads to misreading of all the bases within that gene following the mutation, causing either no production of the gene protein product or an abnormally functioning protein.
E.g. Duchenne muscular dystrophy - deletion The factor VIII gene in some types of haemophilia A - insertion |
|
|
Results of a point mutation (non or missense) of a base within a coding region of a gene.
CLINICAL example |
Substitution of one base for a different base (aside from silent) leads to the coding of an abnormal protein
e.g. Sickle cell anaemia (autosomal recessive blood disorder) |
|
|
Ethnic difference in disease frequencies
|
e.g. Sickle cell anaemia
|
|
|
Results of the fusion of two genes
CLINICAL example |
Produce an abnormal protein product
e.g. Lepore haemoglobins |
|
|
Results of deletion, insertion or point mutation within a coding region of a gene.
CLINICAL example |
Can interfere with the binding of transcription factors to this region and cause the loss or reduced transcription of the coding region of the gene
e.g. some hereditary haemolytic anaemias |
|
|
Types of chromosome
|
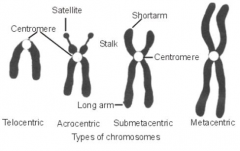
Classified by position of centromere:
Metacentric - central (median), arms of equal length Sub-metacentric - near centre (sub-median), two unequal arms Acrocentric - near one end (sub-terminal), one arm very short, the other long Telocentric - terminal |
|
|
Why do chromosomal abnormalities normally have a more complex phenotype?
|
Because much larger amounts of genetic material are involved
|
|
|
Define aneuploidy
|
The state of having a chromosome number that is not a multiple of the haploid number.
|
|
|
How does trisomy occur?
examples (3) |
When a pair of homologous chromosomes fail to separate at meiosis.
Causes over expression of the genes on this chromosome e.g. trisomy 13 - Patau syndrome trisomy 18 - Edward's syndrome trisomy 21 - Down's syndrome |
|
|
Incidence and symptoms of Down's syndrome?
|
1 in 650-1,000 - one of the most common genetic disorders
Learning difficulties, premature cataract development, susceptibility to certain bacterial infections, congenital heart defects and a predisposition to leukaemia |
|
|
Incidence, symptoms and life expectancy of Edward's syndrome?
|
1 in 6000
Severe learning disabilities Small, abnormally shaped head Long fingers that overlap and clenched fists low-set ears Smooth 'rocker bottom' feet (with a rounded base) heart and kidney problems feeding problems in infancy, leading to poor growth breathing problems bone abnormalities, such as a curved spine A third of babies born alive will die within a month of birth because of life-threatening medical problems. |
|
|
Incidence, symptoms and life expectancy of Patau syndrome?
|
1 out of every 10,000 newborns
Heart defects Brain or spinal cord abnormalities Microphthalmia (very small or poorly developed eyes) Cleft palate Hypotonia (weak muscle tone). Due to the presence of several life-threatening medical problems, many infants with trisomy 13 die within their first days or weeks of life. |
|
|
How does monosomy occur?
CLINICAL example |
Occurs as the other daughter cell of a meiotic division which produced a trisomic cell - usually results in a non-viable foetus which spontaneously aborts during the first trimester
Turner's syndrome - only one X chromsome |
|
|
Explain deletion as a mechanism for chromosomal abnormality
CLINICAL examples |
Loss of a portion of a chromosome - deleted portion will be lost at the next mitotic division. May not be much phenotypic effect if the homologous chromosome remains in tact (still one copy of the genes)
e.g. DiGeorge syndrome, William's syndrome |
|
|
Details of DiGeorge syndrome
|
In 22q11 deletion, a tiny part of the long arm of one of the two copies of chromosome 22 is missing at position 11.
1 in every 4000 of the population |
|
|
Details of William's syndrome
|
1 in 10,000 people worldwide
|
|
|
How do ring chromosomes occur?
|
Specific case of deletion in which material is lost from each end of a chromosome and the new ends join together to form a ring - causes many problems at mitosis.
|
|
|
How do inversions occur?
|
Two breakages occur within a chromosome and the central segment rotates before rejoining
|
|
|
When do isochromosomes occur?
|
Occur when one arm of a chromosome is lost (e.g. the short arm) and the other arm duplicates to replace it (e.g. the long arm) - produces genetic monosomy of the lost arm and trisomy of the duplicated arm
|
|
|
How do translocations occur?
What affect will it have? |
A breakage occurs and material is attached to a different chromosome.
If material is exchanged between two non-homologous chromosomes, a normal phenotype may be produced however major abnormalities will occur in gametes if only one of the translocated chromosomes segregates into an individual cell. |
|
|
Reciprocal translocation
Balanced and unbalanced |
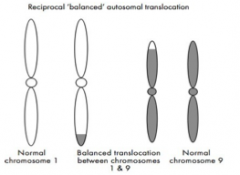
Can occur between any of the chromosomes and involve pieces of any size.
Arises when an exchange of chromosome material takes place between two different chromosomes. Since in these translocations there does not appear to have been any loss or gain of chromosome material, the translocation is described as `balanced’. |
|
|
Robertsonian translocation
|
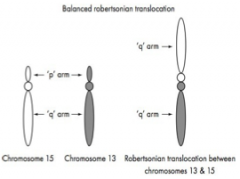
Only involve exchanges between chromosomes 13, 14, 15, 21 and 22 as these are acrocentric chromosomes.
The exchange involves loss of the short arms of two chromosomes and fusion of the remaining two long arms at their centromeres The result is one long chromosome that consists of two long arms. |
|
|
What are trinucleotide repeat diseases?
|
Amplification of a specific trinucleotide sequence in a gene (could be classed as insertions into coding or non-coding regions of the affected gene but they have specific properties that set them apart).
|
|
|
What occurs in a trinucleotide repeat disease?
Example? |
A specific trinucleotide sequence which normally has several repeats in a gene is amplified to have many hundreds or thousands of repeats
e.g. CGG in fragile X syndrom |
|
|
When does this amplification occur?
|
May be during spermatogenesis as in Huntington's or oocytogenesis as in fragile X
|
|
|
How does the amplification have an effect in non-coding regions?
|
The amplified repeat sequence interferes with transcription and can induce methylation (and therefore non-function) of the whole promotor region
|
|
|
What effect does the amplification have in coding regions?
|
Produces proteins with abnormal functions e.g. a form of the huntington disease gene (huntingtin) that abnormally inactivates proteins normally associated with it.
Diseases present earlier and more severely with successive generations due to the amplification which occurs during gametogenesis - anticipation. |
|
|
What is investigated in evolutionary medicine?
|
The potential imbalance between environmental development and evolution of the human genome. The genome has been subject to selections by evolutionary pressure over millions of years however in the last 1000 the environment has been changing much more rapidly than any genetic changes and this may lead to disease.
|
|
|
What is a polymorphism?
|
Variations in DNA sequence between individuals
(that doesn't appear to produce a specific deleterious (damaging) phenotype) |
|
|
What are restriction fragment length polymorphisms?
|
Difference in homologous DNA sequences between individuals that can be detected by the presence of fragments of different lengths after digestion by restriction enzymes which cleave the DNA specific sites.
|
|
|
What are variable number tandem repeat (VNTR) sequences?
Practical use? |
A tandem repeat from a single genetic locus in which the number of repeats varies from individual to individual.
These sequences are used for identification purposes (as in DNA fingerprinting) |
|
|
What are short tandem repeat sequences?
Also called? |
A type of variable number tandem repeat, STRs are repeating sequences of 2-6 base pairs of DNA.
Also called microsatellites |
|
|
What are short tandem repeat polymorphisms?
Practical use? |
Different number of copies of the repeat element (STR) that can occur between individuals in a population.
Basis of genetic fingerprinting using in criminal investigations and paternity suits |
|
|
Single nucleotide polymorphisms (SNPs)
How are these detected? |
Restriction fragment length polymorphisms are just surrogate measures of single nucleotide polymorphisms - the most common type of genetic variation among people. Each SNP represents a difference in a single nucleotide. e.g. a SNP may replace C with T in a particular stretch of DNA.
Can be detected directly through high through-put sequencing or oligonucleotide microarrays (rather than analysing restriction fragment length polymorphisms) |
|
|
Why is it difficult to find the polymorphisms with are important in the pathogenesis of a specific disease?
|
Because there are so many - two individuals will have around 2.5 million single nucleotide polymorphisms between them (1 in 1300 nucleotides)
|
|
|
What is linkage?
|
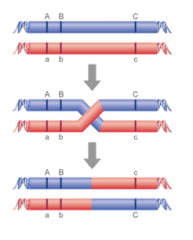
The phenomenon whereby alleles at loci close together on the same chromosome will tend to be inherited together, because it will be rare for a crossover to occur between the loci at meiosis.
|
|
|
Association studies (linkage analysis)
|
Investigate statistical association between any genetic marker and a particular phenotype (usually a specific gene causing disease)
|
|
|
How is the strength of association between a marker and disease determined?
|
Statistical analyses can be made when genetic markers (e.g. SNPs) are examined in a large population of individuals in which it is known which members have or do not have the disease
|
|
|
What does very strong association suggest?
|
That the marker is very close to/actually in the region of DNA that codes for proteins involved in the disease.
|
|
|
How are association studies made easier?
|
Use populations with a relatively restricted gene pool and long-known pedigrees. Found in geographically or socially insular communities.
|
|
|
Linkage studies are useful to identify single gene defects - 2 examples
Can studies be used to define polygenic diseases? |
Cystic fibrosis (CFTR) and Huntington's disease
Studies are much more difficult |
|
|
Linkage disequilibrium
|
A non-random association of two genes on the same chromosome - the occurrence of some combinations of alleles/genetic markers in a population more or less frequently than would be expected from a random formation of haplotypes from alleles based on their frequencies.
|
|
|
Define haplotype
|
A combination of alleles that are located closely together on the same chromosome and tend to be inherited together.
|
|
|
Pharmacogenetics and pharmacogenomics
|
The study of how medical genetics could be used to improve the therapeutic benefits of administered drugs
|
|
|
Pharmacogenetics
|
Relates to the study of how an individual gene might affect the body's response to a drug.
Historically, term has been applied specifically to describe the study of polymorphisms, that effect enzymes involved in drug metabolism, with a bearing on the potential toxicity of a given drug |
|
|
CLINICAL
How are pharmacogenetics clinically applicable? Example |
identifying patients with particular polymorphisms, prior to drug administration, could identify those at risk of toxic side-effects and would help physicians modify the treatment regime accordingly.
Much work has centred on polymorphisms in P450 enzymes which are involved in the metabolism of many drugs in the liver. |
|
|
Pharmacogenomics
|
Relates to how the entire genome affects the body's response to a drug.
Encompasses both the impact of a specific gene e.g. codes for a target protein such as an enzyme or receptor, and also the array of genes that might determine and individual's ability to absorb and distribute a drug and its subsequent clearance through metabolism and excretion. |
|
|
CLINICAL
Implications of pharmacogenomics |
It is hoped that knowledge of a patient's genome will facilitate 'individualised medicines' tailored to a patient's genetic make-up - help doctors to choose the best drug at the optimal dose for a patient whilst avoiding adverse reactions determined by genetic predisposition.
Also, will help in identifying new gene product targets for drug intervention. |
|
|
Assortative mating
Positive and negative |
The mating of individuals with more traits in common (similar phenotypes) than likely in random mating
Positive assortative mating occurs when phenotypically similar individuals tend to mate, and negative assortative mating when unlike individuals tend to mate. |
|
|
Genetic drift
|
The process of change in the genetic composition of a population due to chance or random events rather than by natural selection, resulting in changes in allele frequencies over time.
Usually most noticeable in small populations |
|
|
Heterozygote advantage
|
The greater fitness of an organism that is heterozygous at a given locus compared to either homozygote - dominant or recessive.
NB: Since loss-of-function mutations tend to be recessive (because dominant mutations of this type generally prevent the organism from reproducing and thereby passing the gene on to the next generation), the result of any cross between the two populations will be fitter than the parent. |
|
|
Genetic recombination
|
The transmission-genetic process by which the combinations of alleles observed at different loci in two parents become shuffled in offspring.
|
|
|
LOD scores, Hardy-Weinburg, GWAS, sib-pairs, CANCER GENETICS,
prenatal diagnosis (clinical), FISH |
.
|

