![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
28 Cards in this Set
- Front
- Back
|
Describe the differences between intracellular and membrane receptor-regulated signaling pathways; what are the major types?
|
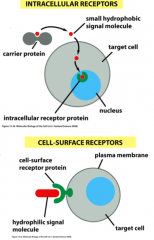
Intracellular: small, hydrophobic signal molecules that can pass through the plasma membrane to find the receptor inside the cell
*steriod hormone receptors *retinoic acid receptors *thyroid hormone receptors Membrane: hydrophilic signal molecule, binds receptor on celll surface, have first and sometimes second messengers carry signal to target *ion channel-linked receptors *G-protein coupled receptors *enzyme-coupled receptors |
|
|
Describe the Jak-STAT pathway.
|
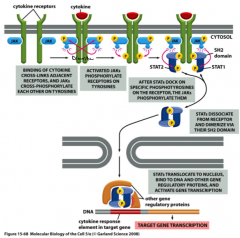
Purpose: STAT is transcription factor that turns on transcription of key cytokine response genes
Primary signal/First messenger: selected cytokines: proteins that stimulate growth and proliferation of cells in immune system Second messenger: none Response target: transcription apparatus Example: cytokine receptor with kinase activity (usually a dimer) |
|
|
Describe the cAMP-mediated signaling pathway.
|
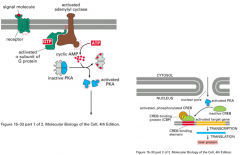
Primary signal/first messenger: selected hormones like epinephrine
Second messenger: cAMP Response target: transcription apparatus Example: Beta-adrenergic receptor (usually 7-transmembrane receptor); large heterotrimeric G-proteins |
|
|
Describe the Inositol Triphosphate (IP3) pathway.
|
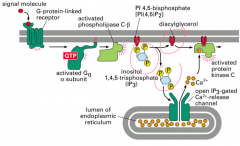
Purpose: regulates calcium release; involved in pathways for taste, manic depression and tumor promotion
Primary signal/first messenger: selected growth factors, mitogens and immune signals, like histamine Second messenger: Inositol triphosphate (IP3), diacylgylcerol (DAG) and Ca2+ Response target: endoplasmic reticulum; transcription apparatus Example: usually 7-transmembrane receptor |
|
|
Describe the phosphotidyl inositol triphosphate (PIP3) AKT kinase pathway.
|
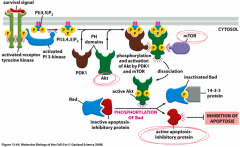
Purpose: to control cell growth and survival; inhibits apoptosis
Primary signal/first messenger: selected hormones and apoptosis control proteins Second messenger: phosphotidyl inositol bisphosphate (PIP2) and diacylgylcerol (DAG) Response target: apoptosis; transcription and translation apparatus Example: receptro with extracellular and intracellular domains and intrinsic kinase activity |
|
|
Describe the tyrosine kinase-MAP kinase pathway.
|
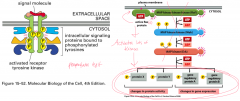
Purpose: associated with growth factor receptors, important in cancer
Primary signal/first messenger: growth factors, like EGF Second messenger: none Response target: transcription apparatus Example: tyrosine kinase dimer |
|
|
Describe the cell cycle, the four major checkpoints and what controls them.
|
G1 phase: repair DNA, prepare to synthesize DNA; cyclin D-CDK 4/6 and cyclin E-CDK 2
*G1 checkpoint: determines whether DNA is damaged before replication occurs S phase: synthesis and replication of DNA; cyclin A-CDK 2 and cyclin E-CDK 2 *intra-S checkpoint: check for replication-induced damage G2 phase: ensure preparation for mitosis; cyclin A-CDK 1 *G2 checkpoint: determines if DNA has been completely and faithfully replicated M phase: mitosis occurs; controlled by cyclin A-CDK 1 and cyclin B-CDK1 *Mitotic checkpoint: determines that chromosomes are properly aligned for cytokinesis G0: position outside cell cycle; most terminally differentiated cells are here; growth factors stimulate cells to re-enter cell cycle |
|
|
Describe the mTOR complex and its role in the regulation of cell growth.
|
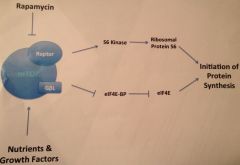
mTOR: key regulator of cell growth by regulating protein synthesis machinery
*complex formed with raptor and G-beta-L ---is can be targeted by Rapamycin to be used in tumor treatment ***translation initiation factor eIF4E is target of mTOR; factor eIF4E inhibits translation by binding, so mTOR inhibits is binding, enabling translation |
|
|
Describe the retinoblastoma (Rb) protein and its regulation of the cell cycle.
|
Rb protein: controls re-entry of cells into the cell cycle by inhibiting E2F, which suppresses transcription
***important for cell proliferation ---can be phosphorylated/inhibited by cyclin D-CDK 4/6 and cyclin E-CDK 2 (through positive feedback), allowing transcription to be activated and the cell cycle to proceed |
|
|
What are the eight things to remember about DNA replication?
|
1. Semi-conservative
2. Requires a primer 3. Bidirectional 4. Lagging strands and leading strand synthesis 5. Key reactions: priming, elongation, termination 6. Replication coupled repair: proofreading and methyl-directed mismatch repair 7. Multiple origins are regulated: must fire only once per cell cycle 8. Replication end problem: telomerase fixes ends |
|
|
Describe the photo-reversal of UV lesions DNA repair pathway.
|
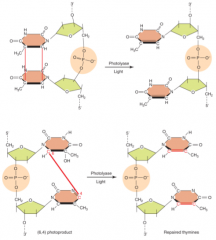
Single nucleotide damage: repairs photodimers, uses photolyase enzyme
|
|
|
Describe the base excision repair pathway (BER).
|
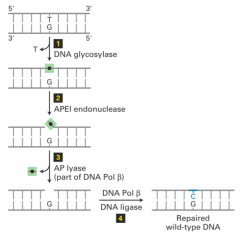
Single nucleotide damage: resects and replaces damage nucleotide bases
---Three steps: *base is removed by DNA glycosylase *phosphodiester backbone is broken and deoxyribose sugar and phosphate removed *DNA pol beta and DNA ligase repair gap ---Damage specific, non-bulky lesions (deamination, hydroxylation, oxidation); removes only one nucleotide ---can generate trinucleotide repeats |
|
|
Describe the nucleotide excision repair pathway.
|
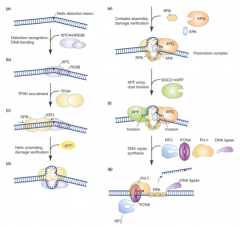
Single nucleotide damage: repairs helix distorting lesions
*can be coupled to transcription; preferential repair of transcribed strand ---repairs bulky, helix distorting lesions (UV thymidine dimers, cisplatin adducts), has broad substrate specificity (any helix distortion); removes group of nucleotides around lesion |
|
|
Describe the mismatch repair pathway.
|
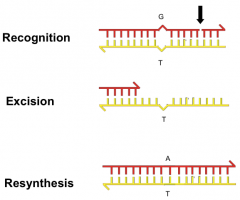
Single nucleotide damage: repairs normal nucleotides that can not base pair
*used fro inadvertant mismatched nucleotides during DNA synthesis ---Three steps: *recognition: identify non-template strand through nicks (due to primers); uses MSH complex *excision: using MLH1, PMS2 exonuclease *resynthesis: using DNA polymerase |
|
|
Describe the homologous recombination pathway.
|
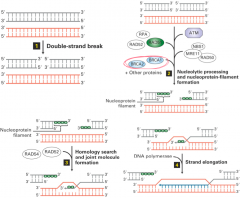
Double strand break repair: uses homologous sequence as template
*ATM and BRCA proteins are key players in break repair ***error free repair |
|
|
Describe the non-homologous end joining pathway.
|
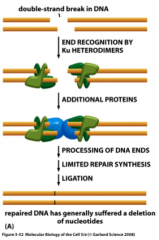
Double strand break repair: resects ends to uncover 2-3 nucleotide micro-homology
***DNA-protein kinase and Ku are used to align dsDNA breaks ***repaired DNA generally suffers deletion of nucleotides ---primarily caused by chemical mutagens or X-rays |
|
|
What are the major classes of cancer and the cell types they arise from? What are the most common cancers and those with the highest mortality rate?
|
Carcinomas: arise from epithelial cells
Sarcomas: arise from connective tissue cells Leukemias/Lymphomas: arise from leukocytes and lymphocytes Other: arise from neuronal cells (gliomas/astrocytomas), melanocytes (melanomas) Most common cancers: ---reproductive tract, digestive organs, breast, respiratory system Highest mortality: ---respiratory system, digestive organs, reproductive tract, breast |
|
|
Describe the six hallmarks of cancer and examples of cellular processes that are key to each one.
|
Sustained proliferative signaling: ligand-independent firing (no need for ligand binding to sustain growth); EGFR inhibitors
Evading growth suppressors: insensitivity to anti-growth/anti-proliferation signals (tumor suppressor genes); cyclin-dependent kinase inhibitors Activating invasion and metastasis: cells invade capillaries to metastasize through bloodstream to other organs/lymph nodes; inhibitors of HGF/c-Met Enabling replicative immortality: telomerase is up-regulated to lengthen lifespan; telomerase inhibitors Inducing angiogenesis: capillaries supply cancer stroma to provide blood, nutrients; inhibitors of VEGF signaling Resisting cell death: evasion of apoptosis; proapoptotic BH3 mimetics |
|
|
Describe the mechanism of the epidermal growth factor (EGF).
|
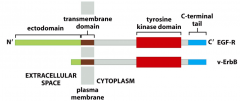
EGF: important class of oncoproteins
---mutations can lead to constitutive cell growth/proliferation |
|
|
Describe the mechanism of the Bcr-Abl protein.
|
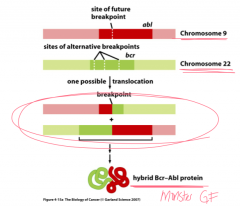
Bcr-Abl: initiated by chromosomal translocation
---chronic myelogenous leukemia (CML): occurs from translocation of chromosome 9 (Abl) and 22 (Bcr) ---has enhanced tyrosine kinase activity ---can be blocked by Gleevec: stops proliferation (leukemia) |
|
|
Describe the mechanism of the c-Myc protein.
|
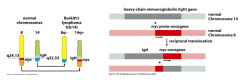

c-Myc: reciprocal translocation of chromosomes 8 and 14 results in fusion of transcription factor c-Myc with IgH enhancer results in uncontrolled of activation of c-Myc
|
|
|
Describe the mechanism of the Ras protein.
|
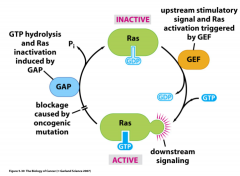
Ras: small G-protein that transduces signals from many receptors
---are activated by GTP/ inactivated by GDP ---is central regulator of many important cell signaling pathways ---can cause constitutive cell growth and proliferation; is key oncogene in many human cancers (multiple hit hypothesis) |
|
|
Describe the mechanism of the Wnt/beta-catenin signal/protein.
|
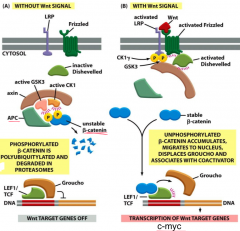
Wnt: signals inactivate the degradation complex
Beta-catenin: activates transcription by LEF1/TCF ---in absence of Wnt signal, beta-catenin degradation complex is active ---in presence, beta-catenin binds and activates transcription of Wnt target genes |
|
|
Describe the Rb pathway and how its inactivation contributes to cancer.
|
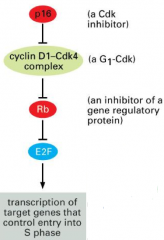
Rb: tumor suppressor - blocks passage through restriction point to inhibit E2F, which contributes to the transcription of target genes (cyclin E and A) that control entry into S phase
*important in regulating cell proliferation Cyclin D1 and Cdk4: oncogenes - inhibit Rb to allow proliferation to occur *can be inhibited in non-dividing cells by p16 - CKI; reinforces ability of Rb to maintain G1 arrest ---mutated in 25-70% of human cancers Rb can also be inactivated by viral infections (adenovirus E1A, HPV, polyoma viruses): produce E7, which binds to Rb, not allowing it to bind E2F, so transcription proceeds |
|
|
Describe p53 and how its inactivation contributes to cancer.
|
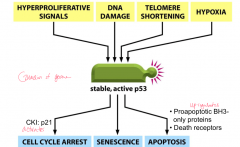
p53: tumor suppressor - important in apoptosis and cell cycle arrest
*guardian of genome: mutated in 50% of all human cancers Cell cycle arrest: activates CKI p21, which blocks cell proliferation Apoptosis: Up-regulates proapoptotic BH3-only proteins and death receptors Replication of damaged DNA in cell without p53 leads to chromosome abnormalities, gene amplification and gene loss ---can also be inactivated by viral infections (adenovirus E1A, HPV, polyoma viruses): produces E6, which binds p53 to inactivate it, not allowing p21 to be transcribed and cell proliferation to be activated |
|
|
Describe Apc and how its inactivation contributes to cancer.
|
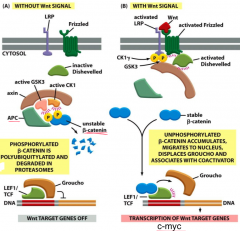
Apc: is scaffold protein; helps target beta-catenin for degradation - phosphorylates it and it becomes polyubiquitylated and degrades in proteasomes
---beta-catenin: activates transcription of Wnt target genes (cMyc) ---Wnt: signals inactivate degradation; in absence, beta-catenin degradation complex is active Mutations cause inappropriate proliferation and cells never differentiate; can lead to polyp formation (especially active in the colon) |
|
|
Describe SMAD4 and how its inactivation contributes to cancer.
|
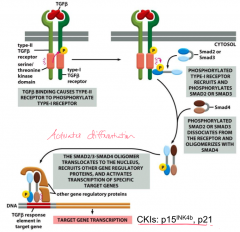
SMAD4: activates transcription of CKIs; involved in the TGF-beta pathway that inhibits proliferation by activating the transcription of CKIs (p15 and p21)
*SMAD4 and TGF-beta receptor mutations are most common - all pancreatic and 83% of colon cancers have these mutations |
|
|
Describe NF1 protein and how its inactivation contributes to cancer.
|
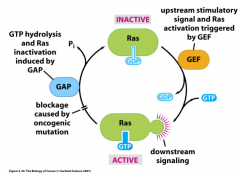
NF1: negatively regulates Ras; is a GTPas activating protein (GAP) and lack leads to Ras staying in its GTP bound activated state - constant proliferation
|

